Introduction
Materials and Methods
Plant Materials and DNA Extraction
SSR Analysis
Genetic Diversity and Population Structure Analysis
Results
SSR Analysis
Genetic Diversity and Discrimination of Korean Ever-Bearing Strawberry
Population Structure Analysis
Discussion
Introduction
Strawberries are one of the world's most widely consumed fruits and are divided into June-bearing type and Ever-bearing type according to the flowering pattern. Flower buds of the June-bearing type differentiate at low temperatures and under short-day condition, and flower buds of the Ever-bearing type differentiate at high temperatures and under long-day conditions. In addition, the day-neutral type, which can differentiate regardless of the day length, is included in the Ever-bearing type. Breeding of these cultivated strawberries is accomplished through the selection of excellent seedlings by a single cross; the use of various species or genetic resources during breeding is insufficient, because most genes that derive from several genetic sources are genetically vulnerable (Sjulin and Dale, 1987; Daubeny, 1990; Dale and Sjulin, 1990). Particularly, it is not easy to collect genetic resources for Ever-bearing strawberry, and it is not easy to distinguish it from similar varieties because there are few differences among varieties when cultivating new ones.
It is essential to have information on genetic diversity and population structure of the target crops for breeding. Therefore, many researchers develop molecular markers for genetic diversity and population structure analysis for various plants, such as rice (Chung and Park, 2010; Cui et al., 2010), millet (Cho et al., 2010a), Cymbidium (Moe et al., 2010), garlic (Zhao et al., 2010), and strawberry (Davis et al., 2006; Shimomura and Hirashima, 2006; Govan et al., 2008; Gil-Ariza et al., 2009), and use them to characterize the germplasm. Random amplified polymorphic DNA (RAPD), amplified fragment length polymorphism (AFLP), cleaved amplified polymorphic sequence (CAPS), and simple sequence repeat (SSR) markers are widely used for the identification of crop varieties and genetic diversity (Nehra et al., 1990; Congju et al., 2000; Tyrka et al., 2002; Bernet et al., 2003; Kunihisa et al., 2005; Cho et al., 2010a, 2010b; Honjo et al., 2011). In complex genomic species, SSR markers are used in genetic and breeding researches because SSR markers are highly polymorphic and highly reproducible and have a high number of alleles and co-dominant inheritance (Zorrilla-Fontanesi et al., 2011).
SSR markers are more effective in evaluating genetic diversity and flexibility than previously developed DNA markers and are therefore widely used for the identification of unisexual propagated crops, such as grapes, apples, pears, potatoes, carnations, turf grass species, Ramie, and strawberries (Bowers et al., 1996; Guilford et al., 1997; Ashkenazi et al., 2001; Dangl et al., 2001, 2005; Kimura et al., 2002, 2009; Smulders et al., 2003; Yamamoto et al., 2006; Wang et al., 2010; Kim et al., 2016). Shimomura and Hirashima (2006) and Govan et al. (2008) selected 4 and 10 SSR suitable markers, respectively, for strawberry genetic mapping. Brunings et al. (2010) selected EMFv104 and EMFvi136 markers, and Hong et al. (2014) selected strawberry SSR markers in Korea. However, knowledge of the genetic diversity of Ever-bearing strawberry cultivars has been limited by the use of SSR markers, and most of them are June-bearing strawberry cultivars.
Thus, this study evaluated the genetic diversity and population structure of 22 Ever-bearing strawberry cultivars and 6 June-bearing strawberry cultivars using 10 selected SSR markers.
Materials and Methods
Plant Materials and DNA Extraction
We used 28 strawberry genotypes preserved at the Highland Agricultural Research Institute of RDA, 22 cultivars of Ever-bearing strawberry, and six cultivars of June-bearing strawberry (Table 1). Genomic DNA was extracted from young leaves of each cultivar using the NucleoSpin®Plant II Kit (Macherey-Nagel, Cat. 740770.250, Germany). The extracted DNA was quantitated with a spectrophotometer (DS-11 + Spectrophotometer, DeNovix, USA) at 10 ng·uL-1 for analysis.
Table 1. The Ever-bearing and June-bearing strawberry cultivars used in this study and their origins 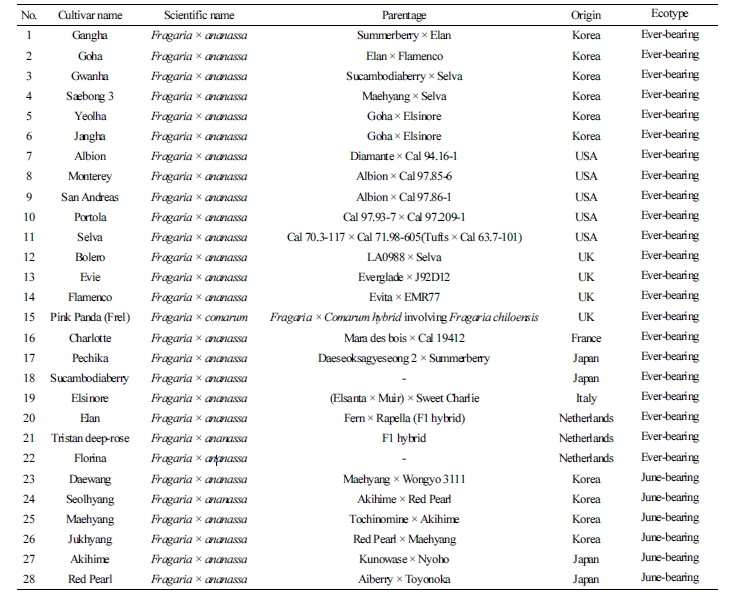 |
SSR Analysis
In this experiment, 37 markers (related to strawberry) were selected (Sargent et al., 2006; Lewers et al., 2005; Bassil et al., 2006; Govan et al., 2008; Hosseini et al., 2013; Hong et al., 2014); then, primers were constructed and polymorphisms were tested using these primers among 28 cultivars. PCR products were confirmed by polymorphism using an automated capillary electrophoresis system (Advanced Analytical Technologies, Inc., USA). In the SSR primer selection, 10 markers with high polymorphism and reproducibility were selected, and fluorescent markers were labeled with FAM (Bioneer, Korea) in the forward primers. SSR analysis was performed on 28 cultivars (Table 1).
The PCR reaction was performed by adding 20 ng of genomic DNA, 10 µL of 2X TOPsimple™ PreMix-nTaq (Enzynomics), and 1 µL of each forward and reverse primer (10 µM·µL-1) and then adjusting the total volume to 20 µL by adding distilled water. The PCR conditions depended on the SSR marker type. Conditions of PCR amplification for markers 1-4 were as follows: 95°C (15 min), 25 cycles at 95°C (30 s), 65°C (30 s, 1°C decrease per repeat), and 72°C (30 s); 30 cycles at 95°C (30 s), 55°C (30 s), and 72°C (30 s); and a final extension at 72°C for 5 min. Conditions of PCR amplification for markers 5 and 6 were as follows: 94°C (15 min), 10 cycles at 94°C (30 s), 65°C (40 s, 0.5°C decrease per repeat), and 72°C (60 s); 25 cycles at 94°C (30 s), 55°C (45 s), and 72°C (60 s); and a final extension at 72°C for 5 min. Conditions of PCR amplification for markers 7-10 were as follows: 94°C (15 min), 30 cycles at 95°C (40 s), the appropriate annealing temperature (52°C-58°C, 40 s), 72°C (40 s), and a final extension at 72°C for 10 min (Table 2). After PCR completion, 5 µL of the amplified product was electrophoresed on 1.5%(w/v) agarose gel to confirm amplification. Then, 0.5 µL (50 ng·µL-1) of the PCR product was mixed with 0.5 µL of GeneScan 500 LIZ dye Size Standard (Applied Biosystems, USA) and 9 µL of Hi-Di formamide (Applied Biosystems, USA). Then, it was denatured at 95°C for 3 min and stabilized at 4°C for 1 min. The denatured PCR product was electrophoresed using an automated sequencer (DNA Analyzer 3730xl, Applied Biosystems, USA) and analyzed by a marker using the Gene Mapper program (Applied Biosystems, USA).
Table 2. The primer sequences, repeat motifs, and annealing temperatures of 10 SSR markers used in the analysis of 28 strawberry cultivars 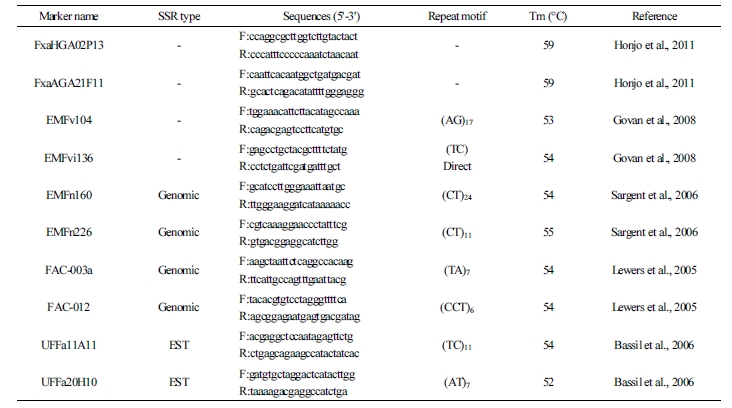 |
Genetic Diversity and Population Structure Analysis
The data from multiple allelic SSRs were used for population statistical analysis with the R package Poppr 2.4.1 (Kamvar et al., 2014) to identify the number of observed alleles (NA), 1-D (Simpson index) (Simpson, 1949), Hexp (Nei’s genetic diversity) (Nei, 1978), and indices of evenness (Grünwald et al., 2003). The pair-wise genetic distances were calculated by Nei’s formula (Nei and Le, 1979) as binary representation of the presence (1) or absence (0) of alleles. The phylogenetic tree of the results was prepared using the phylogenetic software package PAUP 4.0 based on the genetic matrix with neighbor-joining (Saitou and Nei, 1978).
STRUCTURE V.2.3.3 was used to infer population structure with the Bayesian clustering method (Pritchard et al., 2000). The admixture model was applied with 20 runs for each K value from 2 to 5, and each run was performed with a burn-in period of 250000 generations and 500000 Markov Chain Monte Carlo (MCMC) replications. The optimal statistic K was determined by and calculated with STRUCTURE HARVESTER (http://taylor0.biology.ucla.edu/struct_ harvest/) based on the method of Evanno et al. (2005).
Results
SSR Analysis
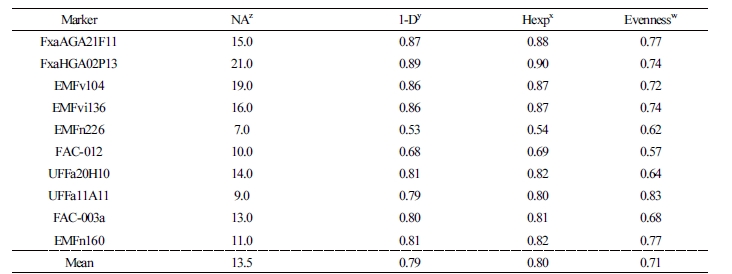
Using SSR analysis, a minimum of 7 (EMFn226) alleles and up to 21 (FxaHGA02P13) were identified by a marker, and the total number of alleles was 135 (average 13.5) (Table 3). The values of indicators of genetic diversity 1-D and Hexp were the lowest in EMFn226, 0.53 and 0.54, and the highest in FxaHGA02P13; the mean values were 0.79 and 0.80, respectively. In this study, 1-D and Hexp values were high either because the genetic diversity of the analytical material was high or the marker with high breed identification was selected first. However, the number of alleles was 7.0, 1-D was 0.53, and Hexp was 0.54 in EMFn226. EMFn226 markers were found to be less effective than other markers for comparing the genetic diversity of strawberries. Indices of evenness indicate a single genotype as the value approaches zero (Grünwald et al., 2003). Indices of evenness ranged from 0.57 to 0.83, and the average value in this study was 0.71. Among the 10 SSR markers, 9 showed a high evenness value of ≥0.60. Therefore, the SSR markers selected in this study could be used to identify strawberry cultivars.
Genetic Diversity and Discrimination of Korean Ever-Bearing Strawberry
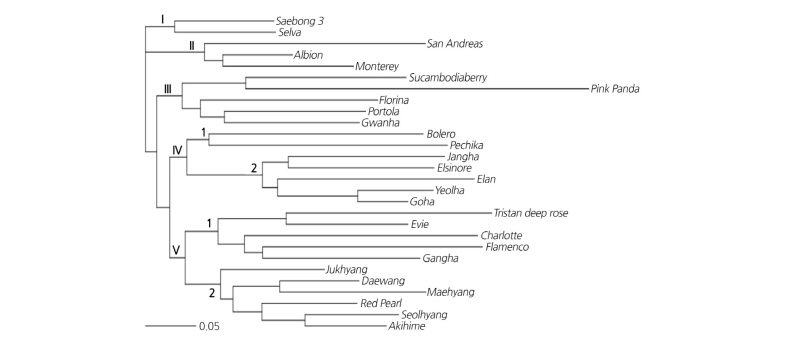
Based on the 135 alleles generated using the selected 10 SSR markers, the genetic relationships of 28 cultivars were examined, and the results of the dendrogram are shown in Fig. 1. The 28 strawberry cultivars are divided into five groups. Groups I to IV are all Ever-bearing strawberry cultivars, and Group V includes both Ever-bearing and June-bearing cultivars. Group I includes ‘Saebong 3’ and ‘Selva’; there is little genetic distance between the two cultivars because ‘Saeabong 3’ is bred as a crossbreed from ‘Selva’ (Table 1). Group II includes ‘San Andreas’, ‘Albion’, and ‘Monterey’. All three cultivars were bred at Strawberry Breeding & Research of UC Davis (University of California, Davis). The genetic distance among the three cultivars is small since ‘San Andreas’ and ‘Monterey’ are bred as a crossbreed from ‘Albion’ (Table 1). Group III contains ‘Sucambodiaberry’, ‘Pink Panda’, ‘Florina’, ‘Portola’, and ‘Gwanha’; among them, ‘Sucambodiaberry’, ‘Pink Panda’, and ‘Gwanha’ are cultivars that bear pink flowers. Because ‘Pink Panda’ of Group III is a cultivar of the species F. comarum, it was distinguished and was the most genetically distant from the remaining 27 cultivars (F. × ananassa) based on the genetic relationship analysis. Group IV includes ‘Bolero’, ‘Pechika’, ‘Jangha’, ‘Elsinore’, ‘Elan’, ‘Yeolha’, and ‘Goha’; the genetic distances of Group IV-2 are close to each other because ‘Jangha’ and ‘Yeolha’ were bred as a crossbreed of ‘Goha’ × ‘Elsinore’ (Table 1). Group V was divided into two groups: Group V-1 included only Ever-bearing cultivars, whereas Group V-2 included June-bearing cultivars.
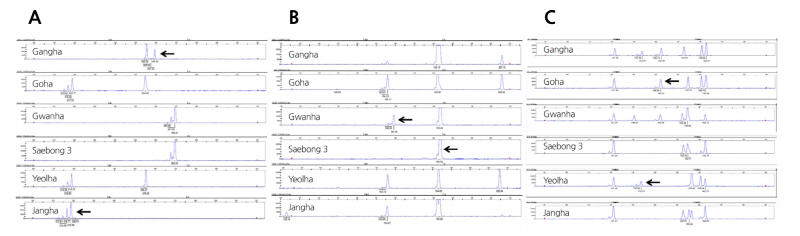
Discrimination of domestic Ever-bearing strawberry cultivars using the three selected SSR markers (UFFa20H10, FAC-012, and EMFvi136) is shown in Fig. 2. Using UFFa20H10, we distinguished between ‘Gangha’ and ‘Jangha’ (Fig. 2A). The amplified fragments of ‘Gangha’ measured 255 bp and 259 bp, while those of ‘Jangha’ measured 215, 217, and 219 bp. Using FAC-012, we distinguished between ‘Gwanha’ and ‘Saebong 3’ (Fig. 2B). The amplified fragments of ‘Gwanha’ measured 167 and 184 bp, while that of ‘Saebong 3’ measured only 184 bp. Using EMFvi136, we distinguished between ‘Goha’ and ‘Yeolha’ (Fig. 2C). The amplified fragments of ‘Goha’ measured 121, 142, 155, 161, and 163 bp, while those of ‘Yeolha’ measured 121, 134, 156, 161, and 163 bp. Using these three SSR markers, we were able to identify six Ever-bearing strawberry cultivars bred in Korea.
Population Structure Analysis
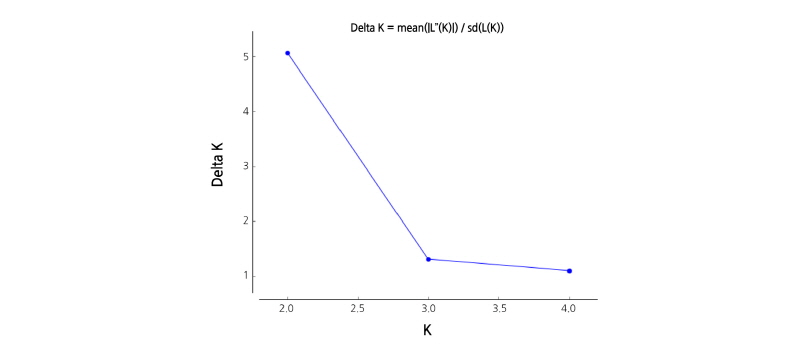
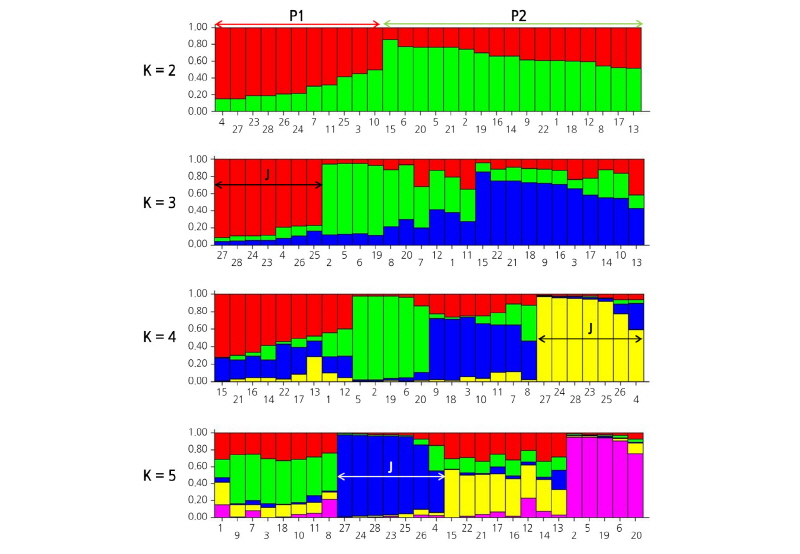
Population structure analysis of 28 strawberry cultivars was performed using a model-based approach (Pritchard et al., 2000) using STRUCTURE V.2.3.3 software. Because it is difficult to deduce the exact value of K, Pritchard and Wen (2003) suggested the selection of the lowest value representing the major structure of the data. By analyzing the distribution of 28 strawberry genetic groups with different K values, it was confirmed with K = 2 that the 28 strawberry genetic groups were divided into two (Fig. 3).
In case of ΔK = 2, population was divided into two groups of P1 (June-bearing cultivars that comprised a part of Ever-bearing cultivars) and P2 (only Ever-bearing cultivars), based on ‘Pink Panda’ (Fig. 4, K = 2). P1 contained June-bearing cultivars, such as ‘Daewang’, ‘Seolhyang’, ‘Maehyang, Jukhyang’, ‘Akihime’, and ‘Red Pear’, and some of the Ever-bearing cultivars, such as ‘Saebong 3’, ‘Albion’, ‘Selva’, ‘Gwanha’, and ‘Portola’. ‘Saebong 3’ (unpublished) was bred as a crossbreed from ‘Maehyang’ (June-bearing); ‘Albion’ (Shaw and Larson, 2006) was bred as a crossbreed from Cal 94.16-1(June-bearing); Cal 71.98-605 (father of ‘Selva’) (Bringhurst and Voth, 1984) was bred as a crossbreed from June-bearing resources; and ‘Gwanha’ (Lee et al., 2012) was bred as a crossbreed from ‘Selva’. All four cultivars are of the Ever-bearing type, but they are derived from June-bearing ancestors and are classified as P1. Although ‘Portola’ (Shaw and Larson, 2009) is a day-neutral cultivar, it first fruited around winters. Thus, ‘Portola’ has characteristics similar to those of June-bearing cultivars and is considered to belong to the P1 group (Table 1, Fig. 4). When analyzing ΔK value from 3 to 5, June-bearing cultivars (‘Daewang’, ‘Seolhyang’, ‘Maehyang’, ‘Jukhyang’, ‘Akihime’, and ‘Red Pearl’) and ‘Saebong 3’ showed a tendency to be grouped together (Group V-2 in Fig. 1).
Discussion
Strawberry plants are a vegetative propagation crop and are difficult to distinguish because of their morphological characteristics (Bassil et al., 2006). Cultivated strawberries are genetically complex octoploids (2n = 8x = 56) and have different parentages; however, most of the alleles are shared among cultivated strawberries. Because the polyploidy of strawberries has been a significant obstacle in revealing the genetic characteristics of the cultivated strawberry, there is limited information about the genome structure.
SSR markers have been used for the identification of varieties and genetic diversity in various crops, such as cucumber (Kwon and Choi, 2013), corn (Wang et al., 2011), tomato (Bredemeijer et al., 2002), wheat (Röder et al., 2002), and potato (Reid et al., 2011). Seventy-five genotypes and cultivars of strawberry have been identified using four SSR markers in Japan (Honjo et al., 2011). In the United States, Chambers et al. (2013) identified 219 genetic resources using eight SSR markers. In addition, Yoon et al. (2012) have identified 59 cultivated strawberries and Hong et al. (2014) identified 100 strawberry genetic resources and cultivars in Korea. However, most studies have focused on June-bearing cultivars and have rarely identified and contributed to the database on Ever-bearing strawberry genotypes and cultivars. Strawberries, such as June-bearing and Ever-bearing cultivars, are ecologically diverse, and recently, the importance of Ever-bearing strawberry breeding has been growing due to their increased cultivation area. This study was conducted to investigate the genetic differences between Ever-bearing and June-bearing strawberries using 10 SSR markers (Figs. 1 and 4). Therefore, it is meaningful that 10 SSR markers were selected to identify the Ever-bearing type. Using these markers, it is possible to identify the species of Korean Ever-bearing strawberries (Fig. 2). Regarding the breeding of Ever-bearing strawberries, it is possible to distinguish between June-bearing and Ever-bearing strawberries using 10 selected markers and to increase the efficiency of breeding.
The results of distance- and model-based analyses are not clearly distinguished and are slightly different for Ever-bearing and June-bearing types (Figs. 1 and 4). This is because most strawberry cultivars used in this study were mixed with the mother group and the cultivated strawberries have high ploidy. Evanno et al. (2005) reported that the K value in most cases depends on the type of marker, the number of samples, and the number of non-identifiers entered in each sample. In this study, the number of SSR markers and the number of populations in the sample were small and could not be clearly distinguished. Therefore, it will be necessary to carry out the experiment again by increasing the number of markers and the number of populations.
Here, we report significant information pertaining to genetic diversity and population structure of Ever-bearing and June-bearing strawberry cultivars using SSR markers. Through genetic diversity and population structure analyses, we found that Ever-bearing and June-bearing strawberries are structurally different, which will help efforts to breed a new cultivar.