Introduction
Materials and Methods
Plant Materials
DNA Extraction, PCR Amplification, and Sequencing
Sequence Editing and Alignment
Phylogenetic Analysis
Statistical Analysis
Results and Discussion
Sequence Analysis
Genetic Variation
Phylogenetic Relationship
Introduction
The genus Brassica, which belongs to the family Brassicaceae, is grown for oil, condiments, vegetables, and fodder around the world. This genus comprises approximately 100 species, including cabbage (B. oleracea L.), Chinese cabbage (B. campestris L.), rapeseed (B. napus L.), mustard (B. juncea L.), turnip rape (B.rapa L.), black mustard (B. nigra Koch.), and Ethiopian mustard (B. carinata Braun) (Ashraf and McNeilly, 2004). Brassica species have a long history of cultivation and are widely distributed in Asia, Europe, the Northern and Eastern Mediterranean region, and Northeast Africa (Warwick, 1993). Their wide distribution has resulted in some confusion regarding their nomenclature and classification. Until the 1950s, researchers distinguished three groups within the Brassica genus: mustard type rape (B. juncea L.), turnip type rape (B. campestris L.), and swede type rape (B. napus L.). These groups were identified based on their biological and agronomic characteristics, utilization features, and genetic relationship. However, the differentiation of the species and the genetic relationships among the members of these three groups were unclear, as the genus Brassica comprised not only cultivated types, but also many weedy and hybrid types that arose in breeding programs and via natural evolution. Even among different sub-species of one cultivated type, extensive diversity in morphological characteristics and genetic diversity can be observed.
Roast meat, including beef, pork, chicken, and seafood, is a central component of Korean cuisine, and is popular all over the world. In Korea, roast meat is often served with lettuce and perilla leaves, because vegetables complement the flavor of the meat. With the development of the Korean food culture and consideration for health and nutrition, the variation of the accompanying vegetables has increased and now includes cabbage, Chinese cabbage, some edible wild herbs, pepper, and garlic. Thus, plant breeding programs now focus on these vegetables. In Korea, there are many cabbage varieties, some of which were developed in plant breeding programs, and some of which occurred by natural hybridization. Therefore, efficient methods for species identification and determination of phylogenetic relationships among Chinese cabbages and cabbages are required.
Much research has focused on deciphering the phylogenetic relationships among Brassica species. Xu et al. (2004) collected 89 Chinese cabbage plants from different provinces of China, and evaluated their morphological traits. They found that the Chinese cabbage germplasm could be divided into two groups according to their geographical origins and that there was large genetic diversity among the local varieties. Meng et al. (2005) analyzed the morphological traits of 65 Chinese cabbage (B. campestris L.) varieties, which exhibited different extents of diversity. Chen et al. (2004) analyzed the genetic diversity of 55 B. campestris accessions from the Hunan province of China using random amplified polymorphic DNA (RAPD) markers, and found a total of 220 polymorphic sites. In addition, Chen et al. (2006) and Huang et al. (2009) evaluated the polymorphism of B. campestris accessions from the Jiangsu and Guizhou provinces of China, respectively, using RAPD markers. These studies reported an abundance of polymorphic RAPD marker sites among the different cultivars. Liu and Meng (2006) evaluated 7 B. napus and 22 B. rapa (B. campestris) accessions from Europe, North America, and China using restriction fragment length polymorphism (RFLP) and amplified fragment length polymorphism (AFLP) markers, and the results suggest that the B. campestris accessions were more polymorphic than the B. napus accessions. Xin et al. (2006) developed a new expressed sequence tag (EST) and simple sequence repeats (SSRs) and successfully identified 15 EST-SSR primer sets to analyze the genetic diversity of 29 Chinese cabbage accessions. Studies based on chloroplast DNA markers suggest that B. campestris and B. oleracea are close relatives of each other (Yanagino et al., 1987; Warwick et al., 1992). Despite numerous analyses of the genetic diversity of B. campestris L. and B. oleracea L. accessions, the phylogenetic relationships among these accessions are still unclear.
Recently, PCR-based molecular identification methods have become routine, efficient means for accurate authentication among plant species (Joshi et al., 2004). DNA-based molecular markers can provide useful information that could be used to study genetic diversity and relationships among the cultivated, weedy, or hybrid Brassica varieties. Among the various DNA markers, the 5S rRNA genes, which exist as multiple copies in the eukaryotic genome, are the most widely used gene family for determining phylogenetic relations among plant species. In higher eukaryotes, 5S rRNA genes exist in tandem repeats and the number of repeats varies from less than 1,000 to more than 75,000 (Campell et al., 1992). The 5S rRNA gene consists of a coding and non-transcribed spacer (NTS) region. The coding region is highly conserved and commonly 120 bp in length, whereas the NTS region varies in length depending on the species, and is highly variable in length (Dvořák et al., 1996; Baum and Johnson, 1996). Thus, the 5S rRNA region is ideal for studying the organization and evolution of multigene families in various plant species (Scoles et al., 1988; Gottlob McHugh et al., 1990). Another 18S rRNA gene can also be used to discriminate between plant species (Soltis et al., 2000; Bleidorn et al., 2003), but due to the resolution of the 18S gene, is usually used for genus discrimination.
In this study, we selected 18 Chinese cabbage (B. campestris L.) and cabbage (B. oleracea L.) accessions from Korea, and evaluated the genetic variation and phylogenetic relationships based on the 5S and 18S rRNA gene sequences. This work identifies novel sequences that can be used to distinguish between the accessionss and to determine the phylogenetic relationships among Brassica species, specifically B. campestris L. species. These findings can be used to help plant breeding programs and in efforts to conserve genetically diverse cabbage materials.
Materials and Methods
Plant Materials
Eighteen common Chinese cabbage (B. campestris L.) and cabbage (B. oleracea L.) accessions from Korea were selected for this study (Table 1). Three of these accessions (B2, B16, and S7) were cabbage, and the remaining 15 (B3-B15 and B17-B18) were Chinese cabbage. One was a hybrid variety, Brassicoraphanus (B4), that was the result of hybridization of a Raphanus sativus (male parent) and a Brassica species (female parent), three were Brassica oleracea L. varieties (B2, B16, and S7), and the remaining 14 materials were B. campestris L. varieties. B. oleracea, B. campestris, and Brassicoraphanus are common crops that are widely used as edible vegetables (Table 1). Mature seeds were sown in plates containing clay/ vermiculite (3:1, v:v). After germination, the seedlings were watered with 1/2 Hoagland nutrient solution (Hoagland and Arnon, 1950) on alternate days. When six mature leaves were present on a plant, fresh leaf tissue was sampled and immediately stored in liquid nitrogen for DNA extraction. The voucher specimens, common name, variety or isolate, and other details of these 18 plants are listed in Table 1.
Table 1. Species, common name, and varieties of Chinese cabbages and cabbages investigated in this study.
| |
No. means number; spp. means species; var. means variety; - means not detected. | |
DNA Extraction, PCR Amplification, and Sequencing
DNA was extracted using the modified cetyltrimethylammonium bromide (CTAB) method described by Doyle and Doyle (1987). The 5S rRNA gene was amplified using the P1 (5’-GATCCCATCAGAACTCC-3’) and P2 (5’-GGTGCTTTAGTGCTGGTAT-3’) primer set in a 20 μL PCR reaction (Park et al., 2000). The 18S rRNA gene was amplified using the universal primers 18SF (5’-AACCTGGTTGATCCTGCCAGT-3’) and 18SR (5’-TGATCCTTCTGC AGGTTCACCTAC-3’, Sogin, 1990). The PCR reactions contained 1 μL of template DNA (~1-100 ng), 10 μL 2 × PCR Dye Master Mix (containing 2 × Taq DNA polymerase, 2 × PCR buffer, 2 × dNTP, and moderate loading dye, QIAGEN, Korea), and 0.1 μmol·L-1 of each primer (including the forward and reverse primer). PCR amplification conditions were as follows: 35 cycles of denaturation at 95°C for 1 min, annealing at 53-57°C for 1 min, and a final extension step at 72°C for 1 min. The amplification products were analyzed by electrophoresis using a 1.0% agarose gel, and then purified for DNA sequence analysis using a QIAquick PCR Purification Kit (QIAGEN, Korea) or Gel Purification Kit (QIAGEN, Korea), according to the manufacturer’s instructions. Purified PCR products were subsequently sequenced at BGI in Beijing, China (http://www.genomics.cn/index).
Sequence Editing and Alignment
For editing and assembly of complementary strands, DNAMAN version 6.0 (Lynnon Biosoft Corporation, USA, http:// www.lynon.com/) was used. Analogues of the sequences and nucleotide sequence comparisons were detected with Basic Local Alignment Search Tool (BLAST) network services against databases (http://www.ncbi.nlm.nih.gov/). Multiple sequence alignment of the 5S rRNA genes was performed using DNAMAN version 6.0, to reveal single nucleotide polymorphisms.
Phylogenetic Analysis
Intraspecial genetic divergences were assessed using pairwise distance calculations (Meyer and Paulay, 2005). Jaccard coefficients, which were used to represent identity among the ecotypes, were calculated based on the similarity coefficient [Sj = a/(a + u)]. In the 5S and 18S rRNA gene sequences, ‘1’ was used for base variation and ‘0’ for no variation; ‘a’ represents the number of the same bases and ‘u’ represents the number of different bases between two accessions. The phylogenetic relationship among 18 Brassica materials was estimated after the construction of a phylogram based on the multiple sequence alignment of various DNA sequences using DNAMAN version 6.0. Based on the typical variable nucleotide sites in the 5S and 18S rRNA gene sequences, the phylogenetic relationship among the studied materials was estimated again. Genetic distance (GD) was determined using MEGA software and the mean GD of the intraspecies and interspecies distance was calculated by adding the individual GD values and dividing this by the number of samples.
Statistical Analysis
Statistically significant differences between the means were determined based on two-way analysis of variance (ANOVA) using Duncan’s multiple-range test (Duncan, 1955). A p value of less than 0.05 was considered significant.
Results and Discussion
Sequence Analysis
To amplify the 5S and 18S rRNA genes from the Brassica samples, the universal P1/P2 and 18SF/18SR primer sets were used. The 17 rRNA 5S and 15 rRNA 18S sequences were successfully amplified in all samples and submitted to the NCBI GenBank database (with accession numbers KT074414-KT074429, KT074436, and KT225359-KT225373, and KT225359-KT225373). BLAST analysis of the B2 5S gene sequence (representing B2, B15, and B18 materials in this study) revealed 97% sequence identity with the existing B. oleracea 5S rRNA gene (NCBI GenBank accession: AJ621401) and 90% sequence identity with the existing B. rapa subsp. chinensis 5S rRNA gene (NCBI GenBank accession: KP099057). The BLAST results of the B3 5S gene sequence (representing most Brassica species investigated in our study) showed 88% sequence similarity to the B. rapa subsp. chinensis 5S rRNA gene sequence (NCBI GenBank accession: LC009528), and 87% sequence similarity with the B. campestris 5S rRNA gene (NCBI GenBank accession: X60723). The finding that the sequence similarity between our sequences and the existing sequence resources was not very high might be due to the limited amount of 5S rRNA gene resources in the NCBI GenBank database. Our results have been submitted to the NCBI GenBank database, and these sequences largely enrich the 5S rRNA gene resource of Brassica species, and facilitate a more comprehensive genetic diversity and phylogenetic relationship analysis. The BLAST results of the S7 5S rRNA gene sequence showed 98% sequence identity with the existing 5S rRNA genes from Atractylodes lancea subsp. luotianensis (NCBI GenBank accession: GQ995231) and Jiangsu (NCBI GenBank accession: GQ995229). There was no existing Brassica sequence resource similar to the S7 5S rRNA gene sequence. This result indicates the importance and genetic diversity of S7, which could be used for functional plant breeding. Our sequence alignment results demonstrate the validity of our amplification approach. Furthermore, our sequencing results enrich the Brassica 5S rRNA gene sequence resources.
The BLAST result for the B2 18S rRNA gene sequence (representing B2, B15, and B18 voucher specimens) showed 99% sequence identity with the existing B. rapa subsp. chinensis small subunit rRNA gene (NCBI GenBank accession: KP099064), Camelina sativa 18S rRNA (NCBI GenBank accession: FN599860), Sisymbrium orientale 18S rRNA gene (NCBI GenBank accession: AB856329), and even with a Arabidopsis thaliana chromosome 3 DNA sequence (NCBI GenBank accession: CP002686). BLAST analysis revealed sequences from various species that belonged to different genera, suggesting that the 18S rRNA gene is well-conserved among plant species. We could use this characteristic to prevent sequence slipping during DNA alignment. Nevertheless, the analogue results validate our amplification. A BLAST analysis conducted on the NCBI server yielded the same results for the B4, B6, and B2 sequences, suggesting that our 18 Brassica accessions had low levels of genetic variation in the 18S rRNA gene region.
Genetic Variation
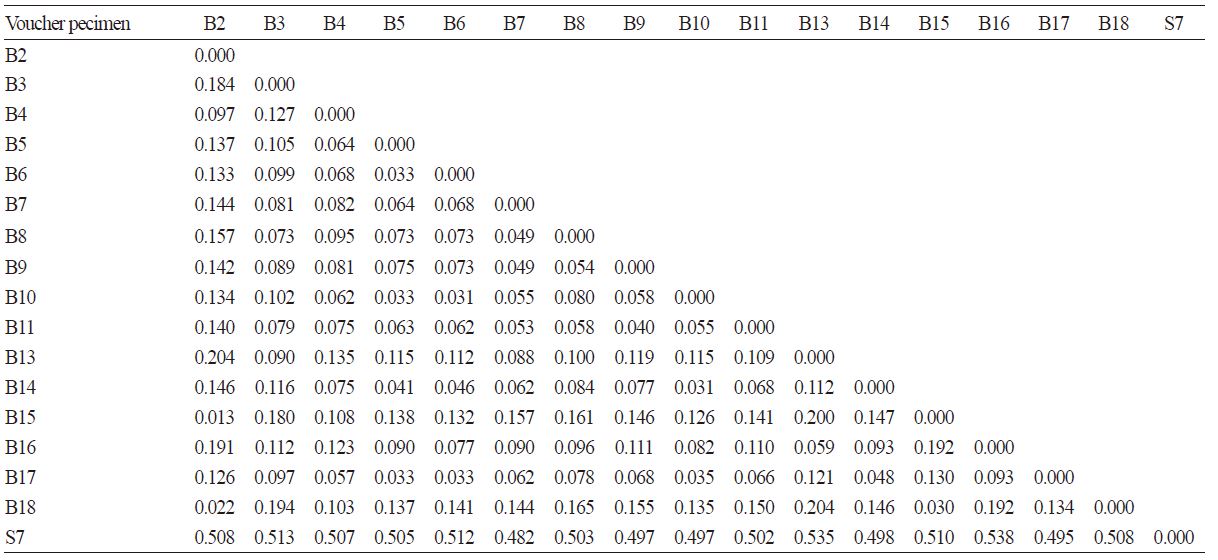
Although B. campestris and B. oleracea are considered to be close relatives, we found that there was considerable genetic variation between these two species, which may be due to self-incompatibility (Nasrallah and Nasrallah, 1989). We deteceted many variable nucleotide sites in the 5S rRNA gene among the samples from the 17 cabbages. S7, belonging to B. oleracea L. variety gem, showed the most genetic variation: the 5S rRNA gene amplified with the same primer set was 160 bp shorter in length than the 5S rRNA gene from the other Brassica species. S7 had a longer genetic distance (GD) than other Brassica materials, rather than having abundant nucleotide variations (Table 2). The other two B. oleracea L. materials investigated in this study, B2 and B16, were expected to show a closer phylogenetic relationship with each other than with B. campestris L.; however, this was not the case. B2 and B16 had a 5S rRNA gene of approximately 470 bp in length, and had a higher GD value with S7 than with other Brassica samples, respectively (Table 2). Interestingly, hybrid cabbage B2, leaf Chinese cabbage B15, and hybrid Chinese cabbage B18 shared nearly three different leaf profiles, but showed the same nucleotide variation when compared with other Brassica material. There were 51 nucleotide sites that were specifically variable between these three Brassica species and others. For instance, in the 68-71 bp, 76 bp, 82-83 bp, and 88-90 bp regions (calculated according to the 5S rRNA gene sequence alignment), B2, B15, and B18 were specifically ‘ATTA’, ‘A’, ‘AT’, and ‘CGG’, respectively. However, others were 4 bp-indel, ‘A’ substitution, 2 bp-indel, and ‘TA(A/C/G)’ at the relative nucleotide location. At the 359-364 bp and 429-431 bp regions, there was also a 6 bp-nucleotide indel/substitution [--AATC/ ACGG(A/C)A] and a 3 bp-nucleotide substitution [GCG/(A/G)AA], respectively. Among B2, B15, and B18, the GD values were reduced to below 0.030 (Table 2). In addition, B16 and B13 shared a relatively low GD value (0.059), which might indicate that, compared to spp. pekinensis, spp. oleifera was more closely related to B. oleracea var. capitata. Other B. campestris L. samples showed similar GD levels with each other, suggesting that there is still genetic diversity within spp. pekinensis, but that they have a relatively close phylogenetic relationship.
Table 2. Genetic distance (GD) of 18 Chinese cabbages and cabbages investigated in this study according to the 5S rRNA sequences
| |
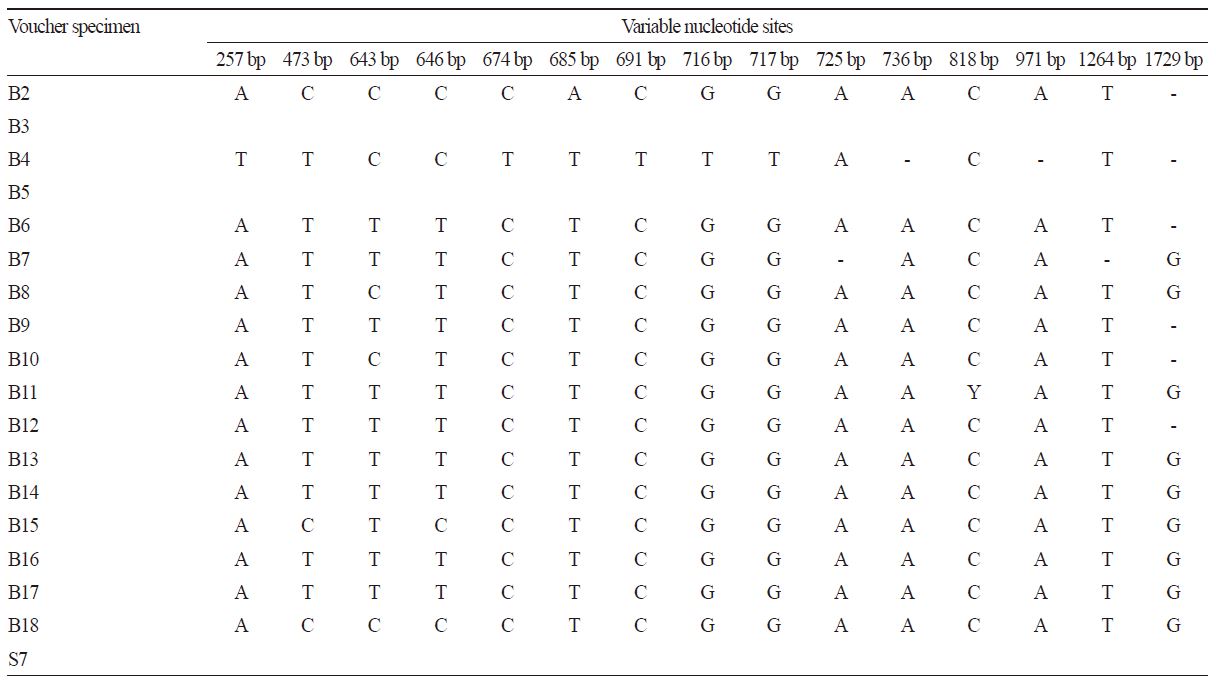
The 18S rRNA gene was relatively conserved with respect to sequence length. However, the nucleotide variation that was present in this region was specific and characteristic. Among the 15 Brassica materials investigated in this study, there were 15 specific variable nucleotide sites (Table 3). Similar to the nucleotide variation pattern of the 5S rRNA gene, B2, B15, and B18 also showed the same substitution at 473 bp (C/T, B2, B15, and B18/others, the same below) and 646 bp (C/T, calculated according to the 18S gene sequence alignment). The hybrid species B4 resulting from Raphanus sativus (male parent) and Brassica species (female parent) showed its own specific nucleotide variations at 257 bp (T/A, B4/others, the same below), 674 bp (T/C), 691 bp (T/C), 716-717 bp (TT/GG), 736 bp (-/A, - means nucleotide indel), and 971 bp (-/A). These specific nucleotide variations would affect the grouping in the phylogenetic tree.
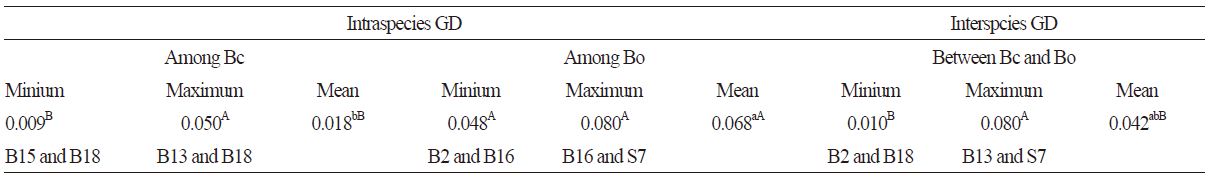
Combining the 5S rRNA and 18S rRNA sequence data could improve our understanding of the genetic diversity and phylogenetic relationship of the Brassica species. The GD values of S7 with other cabbage accessions were all significantly higher than the GD values between any combination of accessions (Table 4). Particularly, S7 and B14, and S7 and B16 shared the highest GD values of 0.073 and 0.080, respectively. B3 and B12 had the lowest GD values (Table 4), and their morphological characteristics were also very similar to each other. There were no significant differences between the intraspecies and interspecies GD values. However, the mean intraspecies GD value within B. campestris L. was significantly higher than the mean GD value within B. oleracea L. (Table 5). Among the 15 B. campestris L. accessions investigated in this study (including the hybrid species B4), there were significant differences in GD values between the different accession pairs (Table 5). However, there was no significant difference between the GD values of the B. oleraceaL. accessions. The interspecies GD values also showed significant differences between B. campestris L. and B. oleracea L. Statistical analysis suggested that higher genetic diversity appeared in the B. campestris L. accessions than in the B. oleraceaL. accessions.
Table 4. Genetic distance (GD) of the 18 Chinese cabbages and cabbages investigated in this study according to the 18S rRNA sequences.
| |
Genetic variation is not correlated with variation in morphological characteristics and geographical distribution between cultivars of B. campestris and B. oleracea and within species (Simonsen and Heneen, Ta1995), largely due to selfincompatibility. However, Nasrallah and Nasrallah (1989) found that self-compatible strains of B. campestris do exist and were generated by self-incompatible strains. Thus, genetic variation could not be used to distinguish between B. campestris L. and B. oleracea L. accessions. Similar results using isozyme and RFLP markers were obtained from the work of McGrath and Quiros (1991). They detected the inheritance of RFLP markers in B. campestris in comparison with B. oleracea, and suggested that both species had extensive gene duplication, conserved gene synteny, and even gene copy number.
Phylogenetic Relationship
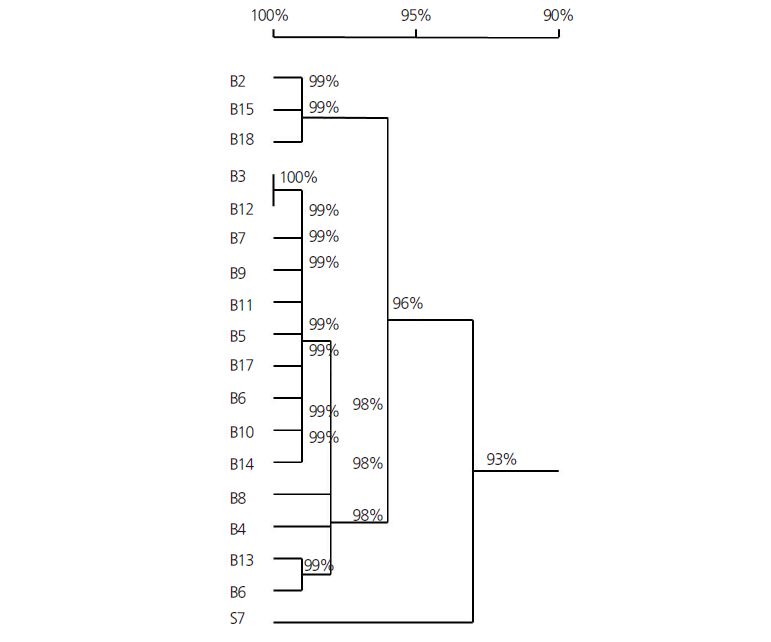
Phylogenetic trees of Brassica species and related genera have been constructed (Song et al., 1990; Attia et al., 1987; Yanagino et al., 1987; Warwick et al., 1992) and suggest that B. campestris and B. oleracea are more closely related to each other than to other Brassica species. To visually express the genetic variation between these Brassica species, we constructed a phylogenetic tree combined with the 5S and 18S rRNA sequences (Fig. 1). Surprisingly, S7 has 93% sequence similarity with other Brassica species. At 96% broader criterion, the other 17 Brassica species were divided into two groups, with B2, B15, and B18 comprising one group, and the remaining species comprising the other. Within this large Brassica group, B13 and B16 formed one subgroup, B4 and B8 formed the next subgroup, and the remaining species formed the last subgroup (Fig. 1). This grouping result was found to closely correlate with genetic variation. While sharing three different types of leaf profiles, B2, B15, and B18 were grouped together due to having the same nucleotide variations in both the 5S and 18S genes (Table 2 and Fig. 1). Another B. oleracea L. species, B16, showed a close phylogenetic relationship with B. campestris L. var. oleifera B13, mainly caused by the genetic variation of the 5S gene (Table 2 and Fig. 1). However, the grouping results were relative to the morphological characteristics. Common Chinese cabbages have curly leaves and a layered yellow and white core (like B5, B6, B7, B10, B11, and B18) or unconvoluted leaves with long or short stems (like B3, B9, B12, and B15), while cabbages have curly leaves and also a layered core, but harder leaves (like B2, and B16). The leaf profile of B13 differs from that of common Chinese cabbage, and was more closely related to that of red cabbage B. oleracea L. (B16); the hybrid species B4 shows a characteristic leaf profile and has a relatively distant phylogenetic relationship with common Chinese cabbage or cabbage; and B8 shows a prominent root characteristic, and although B8 belongs to B. campestris L., it had a parallel phylogenetic relationship with B4, which is another B. campestris L. species. Our phylogenetic tree of B. campestris and B. oleracea was more refined than that obtained in previous studies (Song et al., 1990; Simonsen and Heneen, 1995), which might be due to the number of samples and sample cultivars.
In conclusion, we selected distinct, popular edible cabbage accessions in Korea and evaluated their genetic variation and phylogenetic relationships. Some materials were very rare, which limited the investigations. Our results provide a valuable reference for future Brassica phylogenetic studies. The phylogenetic tree shows clear phylogenetic relationships among Brassica species, and provides a theoretical basis for plant breeding programs aimed at generating improved varieties. Particularly, S7 and B8 could be selected as individuals for hybridization. This study provides additional information that facilitates the evaluation of the genetic relationships among different Brassica cabbage vegetables. Our results can be used to develop crops with improved characteristics and direct plant breeding programs of Brassica species.


.jpg)
.jpg)