Introduction
Materials and Methods
Plant materials
Genomic DNA extraction
PCR amplification and DNA sequencing
High-resolution melting (HRM) analysis
Results and Discussion
Evaluation of 32 commercial tomato cultivars for TSWV resistance using the Sw-5b gene-based marker
Detection of a functional SNP to distinguish resistant and susceptible Sw-5b alleles
Development of the SNP marker for TSWV-resistance
Introduction
Tomato spotted wilt virus (TSWV) causes a very destructive viral disease that threatens worldwide tomato production (Solanum lycopersicum L.syn Lycopersicon esculentum Mill.) (Saidi and Warade, 2008). TSWV is a virus belonging to the genus of negative RNA virus Tospovirus within the family Bunyaviridae. Disease symptoms caused by TSWV include bronzing, curling, necrotic spots and streaks, and stunting of stems and leaves (German et al., 1992). This virus is effectively transmitted by thrips; adult thrips can transmit viruses to a new, healthy plant only 30 minutes after feeding on viruses from infected plants. Prevention of TSWV in crop plants is difficult due to the wide range of plant hosts. Tomato chlorotic spot virus (TCSV) and Groundnut ringspot virus (GRSV), which both belong to Tospovirus and are closely related to TSWV, also cause a decrease in the total production of tomato crops (Brommonschenkel et al., 2000).
The use of resistance genes is a very efficient way to reduce disease. Resistance sources to Tospoviruses have been found in diverse cultivars of Solanum lycopersicum (Saidi, 2008). Some resistance genes for TSWV have also been identified in accessions of S. habrochaites (previously L. hirsutum PI127826 and LA1353) and in S. habrochaites var. glabratum (PI134417 and LA1223). To date, eight resistance genes for TSWV, Sw1a, Sw1b, sw2, sw3, sw4, Sw-5b, Sw-6, and Sw-7, have been reported (Finlay, 1953; Price et al., 2007; Saidi, 2008). Among them, Sw-5b has been widely deployed in tomato breeding because of its durable and reliable resistance to multiple Tospoviruses (Stevens et al., 1991). The genetic location of Sw-5b gene has been mapped between restriction fragment length polymorphism (RFLP) markers CT71 and CT220 near a telomeric region of the long arm of chromosome 9 in tomato (Stevens et al., 1995). This gene was cloned, and the protein encoded by this gene consists of the coiled-coil (CC) domain, the nucleotide binding-adapter shared by APAF-1, R proteins and a CED-4 (NB-ARC) and a leucine- rich repeat (LRR) domain (Spassova et al., 2001). This gene is similar to two known resistance genes in tomato: the nematode resistance gene Mi (51% identity at the amino acid level) and the Prf gene resistant to Pseudomonas syringae (43% identity at the amino acid level) (Spassova et al., 2001).
Maker-assisted selection (MAS) has been widely applied for the breeding of disease- resistant cultivars in many crops, including tomato, by utilizing genetic markers linked to corresponding resistance genes (Foolad, 2007; Foolad and Sharma, 2004). Among diverse cultivars, single nucleotide polymorphisms (SNPs) in certain genes responsible for important phenotypes, such as disease resistance, have been widely used in genome mapping, association studies, diversity analysis, and tagging of economic important genes in plant genomics (Giancola et al., 2006; Rafalski, 2002). Most SNPs have been discovered and verified in tomato (Labate and Baldo, 2005; Yang et al., 2004) and successfully used in selection of various bacterial disease resistances (Yang et al., 2005). Molecular markers, such as RFLP and cleaved amplified polymorphic sequence (CAPS) markers, are based on the presence of SNPs or insertion/deletion (INDEL) mutations (Nasu et al., 2002). SNPs can also be used directly for developing SNP markers. High-resolution melting (HRM) analysis or SNP chips like Fluidigm have been recently developed and are currently being widely used (Wu et al., 2008).
Molecular markers linked to TSWV resistance have also been developed and reported in tomato (Foolad and Panthee, 2012). In particular, for TSWV resistance conferred by the Sw-5b gene, diverse molecular markers such as RAPD (random amplified polymorphic DNA), RFLP, SCAR (sequence charac-terized amplified regions), CAPS, and INDEL markers have been reported (Dianese et al., 2010; Garland et al., 2005; Langella et al., 2004; Roselló et al., 2001; Śmiech et al., 2000; Stevens et al., 1995). Since the successful cloning of the Sw-5b gene, a gene-based marker and two SNPs specific to the Sw-5b allele for TSWV resistance have been reported (Shi et al., 2011). However, SNP analysis between resistant and susceptible alleles of the Sw-5b gene has not yet been intensively examined.
In this study, we first detected SNPs between resistant and susceptible alleles of Sw-5b genes in 32 commercial tomato cultivars. Then, SNP markers for HRM using this SNP information were developed.
Materials and Methods
Plant materials
Thirty-two commercial tomato cultivars were used in this study (Table 1): thirteen cultivars from Nongwoo Bio (Suwon, Korea), eight cultivars from Syngenta (Basel, Switzerland), five cultivars from Monsanto Korea (St. Louis, Missouri, United States), four cultivars from Asia Seed (Seoul, Korea), and one cultivar each from Takii (Kyoto, Japan) and Sakata (Sakata, Japan). Moneymaker and M82 were included as TSWV-susceptible cultivars. Only two cultivars, ‘TY sense Q’ and ‘Titichal’ among the 32 commercial cultivars used in this study were described by the company as being resistant to TSWV. These tomato cultivars were grown in a growth chamber (14 h light/ 10 h dark, 26-28°C, 63.03 µmol・m-2・s-1 light intensity) for 4-5 weeks for genomic DNA isolation.
Genomic DNA extraction
Genomic DNA (gDNA) was extracted from young leaves using the cetyl-trimethyl-ammonium bromide (CTAB) method with minor modifications (Murray and Thompson, 1980). Briefly, fresh, young leaves (0.2 g) were ground in a 2 mL microfuge tube with 300 µL extraction buffer [(2% w/v) CTAB, 1.4 M NaCl, 20 mM EDTA, 100 mM Tris–HCl pH 8.0, 1% PVP, 0.1% sodium bisulfite, and 0.2% (v/v) 2- mercaptoethanol] for gDNA extraction. Extracted gDNA was quantified using a Nanodrop 2000/UV-Vis spectrophotometer (Thermo Scientific, Wilmington, DE, USA) and diluted to 10 ng・µL-1 for polymerase chain reaction (PCR).
PCR amplification and DNA sequencing
For the primer pairs Sw5b-f1/r1 and Sw5b-f2/r2 (Table 2), PCR amplification was conducted using procedures described by Shi et al. (2011). PCR fragments were separated by gel electrophoresis with 1% agarose gel in 0.5 X TAE buffer and visualized under UV light using Dual UV transilluminator after staining with EcoDyeTM DNA Staining Solution (Biofact Co., Daejeon, Korea). The PCR fragments amplified from the primer pairs Sw5-f1/r1 were sequenced in Macrogen Inc. (Seoul, Korea). DNA sequences of the PCR products amplified with the primer pair Sw5-f1/r1 were aligned with ClustalW2 database from EMBL-EBI (https://www. ebi.ac.uk/Tools/msa/clustalw2/).
High-resolution melting (HRM) analysis
For HRM analysis, the Sw5b-SNP primer pair and the SNP1 and SNP2 primer pairs reported in Shi et al. (2011) (Table 2) were designed to amplify the DNA region carrying the target SNP. PCR was carried out in 20 μL reaction mixtures containing 10X h-Taq Reaction Buffer (25 mM MgCl2 mix), 10 mM of each dNTP Mix, 10 pmol of each primer, 2.5 U/µL DNA polymerase, 20X EvaGreenTM (Solgent, Daejeon, Korea), 5X Band DoctorTM and 10ng of gDNA, using a CFX ConnectTM Real-Time PCR Detection System (BIO-RAD, Hercules, USA). Cycling conditions were 95°C for 15 min, followed by 40 cycles of 95°C for 20s, 60°C for 15 s, and 72°C for 30 s, and HRM was analyzed for every increment of 0.2°C between 65 and 95°C.
Results and Discussion
Evaluation of 32 commercial tomato cultivars for TSWV resistance using the Sw-5b gene-based marker
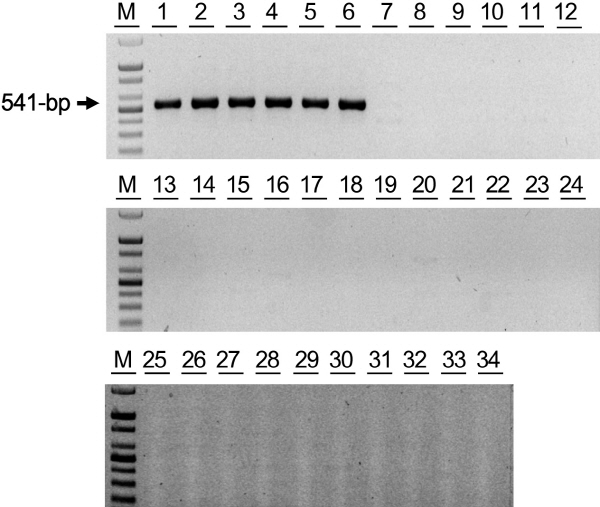
To examine whether commercial tomato cultivars currently available in Korea carry resistance to TSWV, 32 cultivars were collected from the seed companies listed in Table 1. The dominant PCR marker specific to Sw-5b with the primer pair Sw5b-f2/Sw5b-r2 (Table 2) was previously reported (Shi et al., 2011). Collected tomato cultivars were screened by PCR with this primer pair, including the TSWV- susceptible cultivars, Moneymaker and M82. PCR amplified 541-bp DNA products from only six cultivars (‘TY sense Q’, ‘Titichal’, ‘Savera’, ‘TY tiny’, ‘Daylose’ and ‘Shinheuksu’) of 32 cultivars (Fig. 1), indicating that only these six cultivars carry the Sw-5b allele for TSWV-resistance. Except for ‘TY sense Q’ and ‘Titichal’ that were already described by the provider to carry resistance to TSWV, ‘Savera’, ‘TY tiny’, ‘Daylose’ and ‘Shinheuksu’ were newly identified as cultivars carrying resistance to TSWV in this study.
Previously, Shi et al. (2011) also screened 14 tomato accessions and 10 commercial tomato cultivars for TSWV- resistance and found that only three commercial cultivars (‘Talladega’, ‘Sophya’, and ‘BHN-444’) and one accession (LA3667) carried the Sw-5b allele for resistance to TSWV as shown in the Stevens cultivar, the original source of the Sw-5b allele for resistance to TSWV. In this study, completely different commercial cultivars from what Shi et al. (2011) used were screened, and only six of them were determined to be TSWV-resistant. Both studies indicate that not many commercial cultivars in the market carry resistance to TSWV, a situation that needs to be remedied to reduce the large amount of agricultural loss.
Detection of a functional SNP to distinguish resistant and susceptible Sw-5b alleles
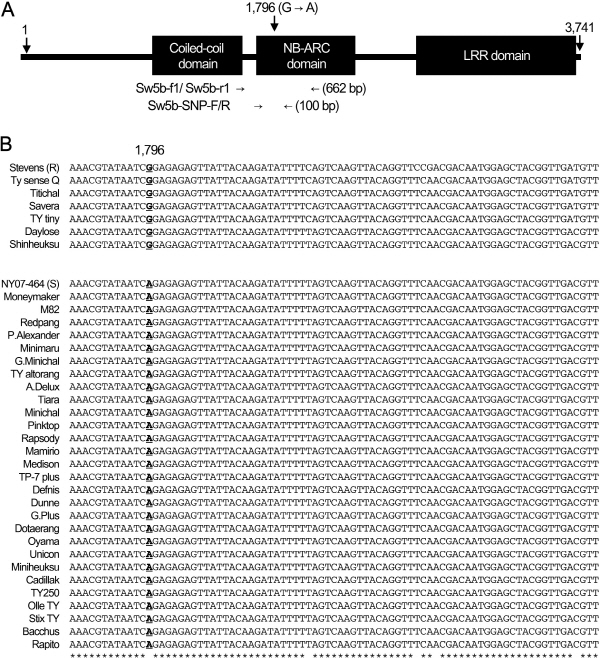
To find SNPs for distinguishing between resistant and susceptible Sw-5b alleles, another primer pair Sw5b-f1/ Sw5b-r1 (Table 2, Fig. 2A) was designed based on a previous report (Shi et al., 2011). PCR with this primer pair amplified 662-bp DNA products from both resistant and susceptible Sw-5b alleles in all 32 cultivars (data not shown). All PCR products were sequenced, and their DNA sequences were aligned using CluswalW2. In addition, the alleles from the resistant cultivar ‘Stevens’ (GenBank Accession No. AY007366) and the susceptible cultivar ‘NY07-464’ (GenBank accession No. FJ686042.1) were retrieved from the GenBank database and included in this alignment. Sequence alignment revealed several SNPs in this region, including G/A at 1,796-bp from the translational start site of Sw-5b allele (Fig. 2C). Among these SNPs, the G/A SNP at 1,796-bp was 100% correlated with genotypes for TSWV resistance. All six resistant cultivars (‘TY sence Q’, ‘Titichal’, ‘Savera’, ‘TY tiny’, ‘Daylose’ and ‘Shinheuksu’) have a G base like the AY007366 allele, while all 22 susceptible cultivars have an A base like the NY07-464 allele. This base change was a non-synonymous substitution, i.e., G to A base substitution results in change of the 599th amino acid, glutamine (Gln599) to arginine (Arg599). Among the three domains of the Sw-5b protein, the 599th amino acid is located in the NB-ARC domain (Fig. 2A). It is likely that, due to different features between these two amino acids, the tertiary structure of the Sw-5b protein changes rendering the protein non-functional.
To date, only one paper has reported SNPs specific to TSWV resistance in tomato (Shi et al., 2011). In that report, six SNPs were found in the Sw-5b allele, and all of them are located in the DNA region, encoding the NB-ARC domain of the Sw-5b protein. Of these six, only two SNPs [A/T at 31,279 position and A/G at 31,392 position in Sw-5b (AY007366) of ‘Stevens’ cultivar] were confirmed to dis-tinguish the TSWV-resistant cultivars from the susceptible cultivars. It has not been determined if the other four SNPs (C/T at 31,415 position, G/A at 31,317 position, G/A at 31,434 position, and C/T at 31,459 position in AY007366) distinguish the TSWV-resistant cultivars from the susceptible cultivars in SNP genotyping. Interestingly, the C/T SNP at position 31,459 is exactly the same SNP confirmed in this study, both of which are the reverse-complement bases of the G/A SNP shown in Fig. 2. In our study, no other SNPs that can distinguish the TSWV-resistant cultivars from the susceptible cultivars were found. This implies that only the G/A SNP can be used for determining TSWV resistance in commercial tomato cultivars available in Korea.
Development of the SNP marker for TSWV-resistance
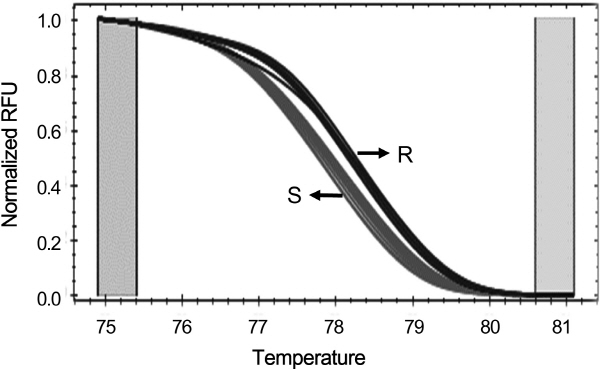
To examine whether the G/A SNP can be used as a SNP marker for selecting TSWV-resistance, the primer pair Sw5b- SNP-F/R was designed. The expected size of the PCR product was 100 bp (Table 2, Fig. 2A) and HRM analysis was carried out. In general, HRM polymorphism is determined by changes in the melting curve shape and location. HRM analysis with the primer pair Sw5b-SNP-F/R clearly distinguished two genotypes at temperatures between 75°C and 80°C (Fig. 3). As summarized in Table 1, only six cultivars (‘TY sense Q’, ‘Titichal’, ‘Savera’, ‘TY tiny’, ‘Daylose’ and ‘Shiheuksu’) were categorized as the resistant group, while all others were the susceptible group. This grouping exactly matched the results from the gene-specific PCR marker shown in Fig. 1. These results indicate that the SNP primer set can be used as a SNP marker for TSWV resistance in commercial tomato cultivars available in Korea.
To compare the accuracy between the SNP marker developed in this study and two SNP markers reported in Shi et al. (2011), HRM analysis was conducted with two SNP markers, SNP1 and SNP2, in the 32 commercial tomato cultivars listed in Table 1. As summarized in Table 1, both SNP1 and SNP2 markers were less accurate than the Sw-5b SNP marker developed in this study. Based on these results, G/A SNP at the 1,796-bp site from the trans-lational start site of Sw-5b allele (corresponding to 31,459 position in ‘Stevens’ cultivar allele AY007366) is more accurate for determining TSWV resistance in commercial tomato cultivars.