Introduction
Materials and Methods
Plant materials for polyploid induction
Plant materials for doubled haploid induction
Shed-microspore culture for haploid induction
Soil acclimatization and cutting propagation of haploid plants for doubled haploid induction
Oryzalin treatments
Plant cultivation and seed harvesting
Ploidy analyses
Stomatal observations
Evaluation of shoot and root growth characteristics
Statistical analyses
Results
Shed-microspore culture technique to induce doubled haploids
Chromosomal effects of the oryzalin concentration and treatment duration
Induction of doubled haploids by means of an oryzalin treatment
Comparison of the leaf and stomatal characteristics of diploid and tetraploid P. grandiflorum plants
Evaluation of doubled haploid characteristics and selection of superior lines
Discussion
Introduction
There is growing interest in herbs, spices, nutraceuticals, and medicinal plants worldwide (Ferrie 2007). Platycodon grandiflorum, a medicinal crop, is a perennial plant of the Campanulaceae family, native to Korea, northeastern China, Japan, the Russian Far East, and southeastern Siberia (Ji et al. 2020). Additionally, the root of P. grandiflorum is among the most widely used traditional herbal medicines in Korea and is a functional resource for managing hypertension and diabetes (Han et al. 2000; Xu et al. 2004; Zhang et al. 2015; Zhang et al. 2022). Domestic cultivars of P. grandiflorum include ‘Jangbaek’ (PVP No. 2003-127, Korea Seed & Variety Service), developed by the Rural Development Administration (RDA); ‘Eutteum’ (PVP No. 2005-18, Korea Seed & Variety Service), developed by the Chungcheongbuk-do Agricultural Research and Extension Services; and ‘Sancheon’, marketed by Kyungshin Seed Co., Korea. However, most farmers are unaware of the exact cultivar they are growing and typically cultivate landrace varieties propagated from seeds collected from wild populations or imported cultivars. P. grandiflorum is a cross-pollinated crop with greater genetic variation relative to that exhibited by self-pollinated crops (Burgarella et al. 2024). Consequently, maintaining cultivar uniformity is challenging and requires considerable time and effort for cultivar breeding.
The improvement of plant genetic resources through induced or natural polyploidy has been a major target of plant breeding programs over the past century (Udall and Wendel 2006; Sattler et al. 2016; Sadat Noori et al. 2017). Polyploids are crucial for plant breeding because they enhance genetic diversity (Sattler et al. 2016). Generally, polyploid plants have larger cells compared to the numbers observed in diploid plants, leading to larger organs and stems (Del Pozo and Ramirez-Parra 2015; Zhang et al. 2019). The primary advantages of polyploids include improvements in various horticultural traits, such as increased height; larger tubers, rhizomes, or roots; greater biomass; enhanced photosynthetic capacity; and larger, thicker leaves than in diploids (Ranney 2006; Caruso et al. 2011). Polyploids can be induced in plants to promote desirable characteristics, although they can also occur naturally (Sattler et al. 2016). For example, tetraploid watermelons have been developed as parent lines for producing seedless fruits, leading to the production of triploid seedless watermelons (Compton et al. 1996; Wehner 2008).
Plant breeding has been conducted for centuries to increase crop yields and enhance cultivar traits. The development of doubled haploid (DH) technology in the late 20th century significantly reduced the time required to produce pure homozygous lines (Hooghvorst and Nogués 2021). Haploids consist of cells derived from a single immature gametic cell and contain only one set of chromosomes (n instead of 2n) (Germana 2011). Doubled haploids are produced by duplicating the chromosomes of haploid plants, enabling the rapid development of fully homozygous lines (Kasha and Maluszynski 2003). This approach significantly reduces the time required compared with traditional breeding methods that depend on multiple generations of self-pollination. Therefore, converting sterile haploids into fertile doubled haploids is essential for reducing the breeding period, and developing an efficient chromosome doubling protocol is crucial to achieving this goal.
Various antimitotic agents have been employed for chromosome doubling in polyploid and haploid plants (Blakeslee and Avery 1937). Colchicine, widely used since its first report in 1937, is effective but limited by its low affinity for plant tubulin and high toxicity to human cells, often causing sterility, abnormal growth, and chromosomal aberrations (Blakeslee and Avery 1937; Morejohn et al. 1984; Luckett 1989; Touchell et al. 2020). In contrast, oryzalin, a dinitroaniline herbicide, specifically targets plant tubulin with minimal vertebrate toxicity, offering a safer and more efficient alternative (Dhooghe et al. 2011; Beranová et al. 2022). Previous studies (Ascough et al. 2008; Sakhanokho et al. 2009; Rêgo et al. 2011) have demonstrated the superior efficiency of oryzalin at lower concentrations, as demonstrated in Rhododendron (Väinölä 2000), Solanum (Greplova et al. 2009), and cucumber (optimal at 75 ppm) (Ebrahimzadeh et al. 2018).
Given the high genetic variability and heterozygosity of P. grandiflorum, a cross-pollinated medicinal crop, achieving cultivar uniformity through conventional breeding is challenging (Burgarella et al. 2024). Therefore, this study aimed to develop an optimized oryzalin-based protocol for inducing polyploidy and doubled haploids (DHs) in P. grandiflorum using microspore-culture-derived haploids, facilitating the rapid development of homozygous lines and reducing the breeding time.
Materials and Methods
Plant materials for polyploid induction
In this study, P. grandiflorum (Jacq.) A. DC., developed by the Department of Herbal Crop Research at the National Institute of Horticultural and Herbal Science (NIHHS), Rural Development Administration (RDA), Eumseong, Korea, and its commercially available cultivar ‘Sancheon’ were used. Baroker horticultural soil (Seoul Bioscience, Eumseong, Korea) was used to fill 72-cell seedling trays, into which seeds were subsequently sown. Approximately three weeks after sowing, when the first true leaves emerged, oryzalin was applied to the apical meristem of the leaves. Polyploids identified through a ploidy analysis at the stage when three to four true leaves appeared were transplanted to a greenhouse, which comprised four 120 cm-wide ridges, each covered with black plastic mulch. The plants in the greenhouse were irrigated using a drip irrigation system.
Plant materials for doubled haploid induction
To induce DH formation, we used five lines of P. grandiflorum (Jacq.) A. DC., developed by the Department of Herbal Crop Research at NIHHS, RDA, Eumseong, Korea. Baroker horticultural soil (Seoul Bioscience, Eumseong, Korea) was used to fill 72-cell seedling trays, into which seeds were subsequently sown. When the third or fourth true leaf emerged around the end of April, 30 plants per line were transplanted to a greenhouse, which consisted of four ridges, each 120 cm wide and covered with black plastic mulch. Flower buds began to form in mid-June and continued blooming until mid-September. Suitable buds were then collected and cultured (Ahn et al. 2024).
Shed-microspore culture for haploid induction
To generate DHs, haploid plants were induced through the shed-microspore culture technique in a double-layer medium consisting of solid and liquid phases (Ahn et al. 2024). Flower buds smaller than 12 mm were used for the shed-microspore cultures. The flower buds underwent surface sterilization by sequential treatments with 70% ethanol for 30 s, rinsing with sterilized distilled water, and disinfection by shaking in a 2% NaOCl solution for 15 min. Finally, they were rinsed five or more times with sterile distilled water. Before the shed-microspore culture step, moisture on the sterilized buds was removed using sterilized filter paper. The sterilized flower buds were dissected to isolate the anthers, which were then incubated on a solid bottom layer of media. Subsequently, 1.5 mL of a top layer of a liquid medium was added, after which the cultures were incubated. The solid bottom layer consisted of the Schenk and Hildebrandt medium supplemented with 2% sucrose and 1% activated charcoal and solidified with 0.4% Gelrite. The composition of the liquid top layer was identical to that of the solid bottom layer but with only 2% sucrose. Both layers were adjusted to pH 5.8 and then sterilized by autoclaving. P. grandiflorum cultures incubated on the solid-liquid double-layer medium were cold-treated at 9°C for seven days, followed by dark incubation at 26 ± 2°C. After two weeks of incubation, 1.5 mL of liquid medium was added to the culture. Embryos formed through the shed-microspore culture technique were transferred to a MS medium containing 2% sucrose and 0.4% Gelrite. The cultures were maintained under warm-white LED light (3000 K, 50 µmol·m-2·s-1) with a 16 h photoperiod at 26 ± 2°C for regeneration.
Soil acclimatization and cutting propagation of haploid plants for doubled haploid induction
When the shoots reached a length of 5–6 cm, they were acclimatized to soil by gently rinsing the roots under running water to remove as much adhering medium as possible while also preventing root detachment. Following the preparation of a 72-cell seedling plug tray with moistened soil, the plantlets were transplanted to facilitate acclimatization. To acclimate the plants to the greenhouse conditions at 25 ± 2°C, the air vents of the plastic lids were opened during the first week and water was provided from the bottom of the tray via a capillary mat system. After the plants were acclimatized to the soil, the lateral shoots were cut at an angle of approximately 45°. The cut surfaces were treated with the rooting hormone Rooton (ISK Bioscience, Japan) before cutting propagation, which was performed in 200-cell nursery plug trays with moisture supplied through bottom irrigation and lids used to maintain the humidity.
Oryzalin treatments
To induce polyploidy in P. grandiflorum, oryzalin was applied to the shoot apical meristems of seedlings following cotyledon expansion and the emergence of the first true leaf. The treatments were performed using four concentrations of oryzalin (12.5 mg·L-1, 25 mg·L-1, 50 mg·L-1, and 100 mg·L-1) for durations of one, two, and three days. Each day, 5.0 µL of the oryzalin solution was applied once to the shoot apex. The oryzalin stock solution was prepared by dissolving 100 mg of oryzalin (Duchefa, Netherlands) in 1 mL of ethanol and diluting it with distilled water to a final volume of 50 mL. Working solutions were prepared by supplementing the stock with 5% ethanol and 5% Tween 20 and diluting with distilled water to the desired treatment concentrations. Following treatment, the seedlings were placed under shade screens in a greenhouse to prevent rapid evaporation, and the relative humidity was maintained above 95% by covering the trays with transparent lids.
For the induction of doubled haploids, oryzalin was applied to the shoot apical meristems of haploid plants obtained through cutting propagation. Only plants confirmed to be haploids via a ploidy analysis using flow cytometry were subjected to the oryzalin treatment. A 50 mg·L-1 oryzalin solution (5.0 µL) was applied once daily for three consecutive days. The treated haploid plants were maintained under shaded and high-humidity conditions identical to those used in the polyploidy induction experiments.
Plant cultivation and seed harvesting
After the oryzalin treatment, P. grandiflorum plantlets were transferred to the greenhouse when three to five true leaves had emerged. The greenhouse comprised four ridges, each 120 cm wide and covered with black plastic mulch. The plants in the greenhouse were irrigated using a drip irrigation system. When P. grandiflorum plants began to bloom, transparent bags were placed over the flowers to prevent pollen mixing, and self-pollination was performed. The seeds of P. grandiflorum plants were harvested before the pods dried completely and were further dried under sunlight.
Ploidy analyses
The ploidy analysis was conducted using young leaves collected from each plant. A Petri dish was prepared with leaf tissue, followed by the addition of 0.25 mL of CystainTM UV Precise P Nuclei Extraction Buffer (Sysmex Partec GmbH, Arndtstraße 11 a-b, 02826 Goerlitz, Germany). A sharp razor blade was used to cut the tissue, after which the extraction buffer containing the released nuclei was filtered through a 30 µm CellTricsTM nylon filter (Sysmex Partec GmbH, Arndtstraße 11 a-b, 02826 Goerlitz, Germany) attached to a 3.5 mL sample tube. After staining the filtered extracts with 1 mL of CystainTM UV Precise P Staining Buffer, flow cytometry was conducted for the analysis. DNA peaks, measured relative to the standard peak of diploid P. grandiflorum, were used to determine the ploidy levels (tetraploid or DH) by comparing them with standard levels. The ploidy analysis was performed using a CyFlow ploidy analyzer, which was equipped with UV (365 nm) and green (532 nm) lasers (Sysmex Partec GmbH, Arndtstraße 11 a-b, 02826 Goerlitz, Germany). Nuclei peak positions were determined and analyzed using CyView software (version 1.6), a flow cytometry standard file analysis tool running on the WindowsTM platform.
Stomatal observations
The nail polish impression method, which was applied to compare the size and density of the stomata, was used for the diploid control and tetraploid plants (Grant and Vatnick 2004). Leaf samples were collected from the third to fourth nodes below the shoot apex before the onset of flowering. Only healthy leaves, free from any signs of disease or discoloration, were selected for this analysis. For sample preparation, transparent nail polish was applied to the back of the leaves and left to dry for approximately ten minutes, at which time it was removed from the leaves using transparent tape. The extracted imprint was then affixed to a glass slide for stomatal observation. Stomata characteristics were observed under an optical microscope at ×400. Five stomata per plant were measured in each treatment group, with ten plants assigned to each group. The stomatal and anatomical characteristics of the leaves were analyzed using ImageJ software (https://imagej.nih.gov/ij).
Evaluation of shoot and root growth characteristics
Plant height was measured from the soil surface to the stem tips as an indicator of the shoot growth characteristics. Leaf shape was classified into six categories: broad, linear, oblong, elliptical, lanceolate, oblanceolate, and ovate. Leaf size was categorized as large (≥ 6cm), medium (4–6 cm), or small (≤4 cm), and the number of lateral branches was classified as high (≥10), medium (5–10), or low (≤5). Flower shape was classified into five categories: pouch-, bell-, broad bell-, cup-, and T-shaped. Petal color was categorized into three types: purple, white, and pink. Roots were carefully washed to remove the soil without damaging the roots, followed by complete moisture removal before proceeding with the measurements to determine the root growth characteristics. Root length was measured as the length of the longest primary root. The extent of lateral root development was classified as high (≥10), medium (5–10), or low (≤5). Root diameter was measured at the thickest part of the primary root using a Vernier caliper (Mitutoyo, Japan).
Statistical analyses
Statistical analyses were conducted using the Statistical Analysis System (SAS) Enterprise Guide 7.1 (SAS, 2009; SAS Institute Inc., Cary, NC, USA) to calculate the mean and standard deviation and to perform an analysis of variance. The statistical significance of the analysis results was assessed using Duncan’s multiple range test, conducted at a 5% significance level (p < 0.05).
Results
Shed-microspore culture technique to induce doubled haploids
Shed-microspore cultures were used to produce haploid P. grandiflorum plants. First, the anthers of heterozygous P. grandiflorum plants were cultured on a double-layer medium with a liquid top layer and a solid bottom layer, with this followed by incubation in the dark (Fig. 1A). Embryogenesis was identified at approximately 70 days after the initiation of the culture (Fig. 1B). The developed embryos were cultured on ½ Murashige and Skoog (MS) medium for approximately four weeks to induce shoot and root formation (Fig. 1C). Plantlets were then sub-cultured on the same medium to regenerate complete plants (Fig. 1D). Subsequently, the medium was carefully removed to avoid root damage, and the plants were transplanted in 105-cell seedling trays for soil acclimatization (Fig. 1E) for approximately four weeks to obtain putative haploid plants. Lateral branches from soil-acclimatized putative haploid plants were used as cuttings to induce DHs (Fig. 1F).
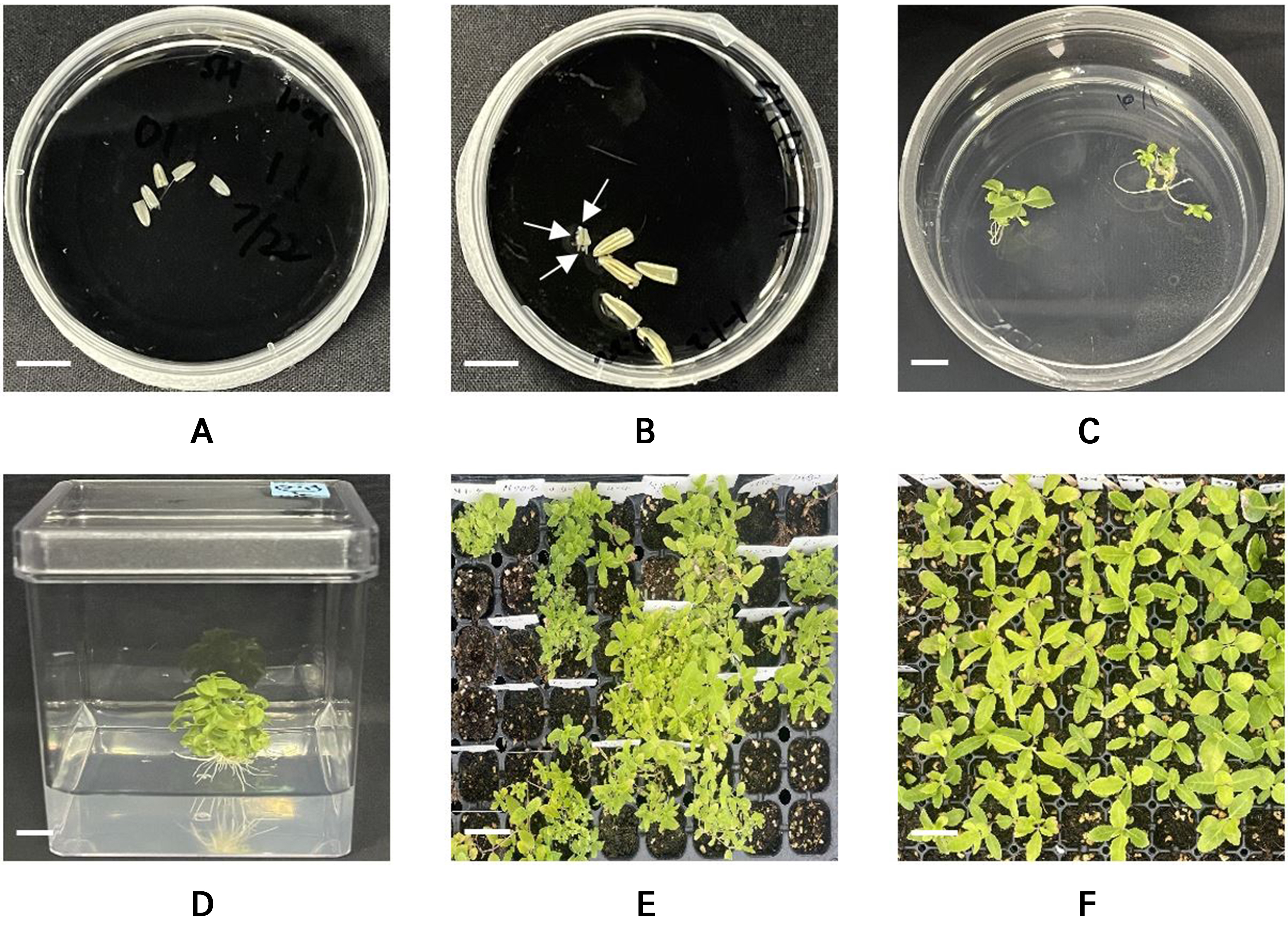

Fig. 1.
Images depicting different stages in the process of the shed-microspore culture of P. grandiflorum: (A) initiation of shed-microspore culture in double layer medium; (B) embryogenesis after approximately 70 days of shed-microspore culture in double layer medium; (C) shoot and root induction on ½ Murashige and Skoog (MS) medium; (D) plant regeneration on ½ MS medium; (E) soil acclimation of regenerated plants; and (F) planted lateral branches. Arrows indicate embryos generated through shed-microspore culture. Scale bars represent 1 cm (A–D) and 3 cm (E, F).
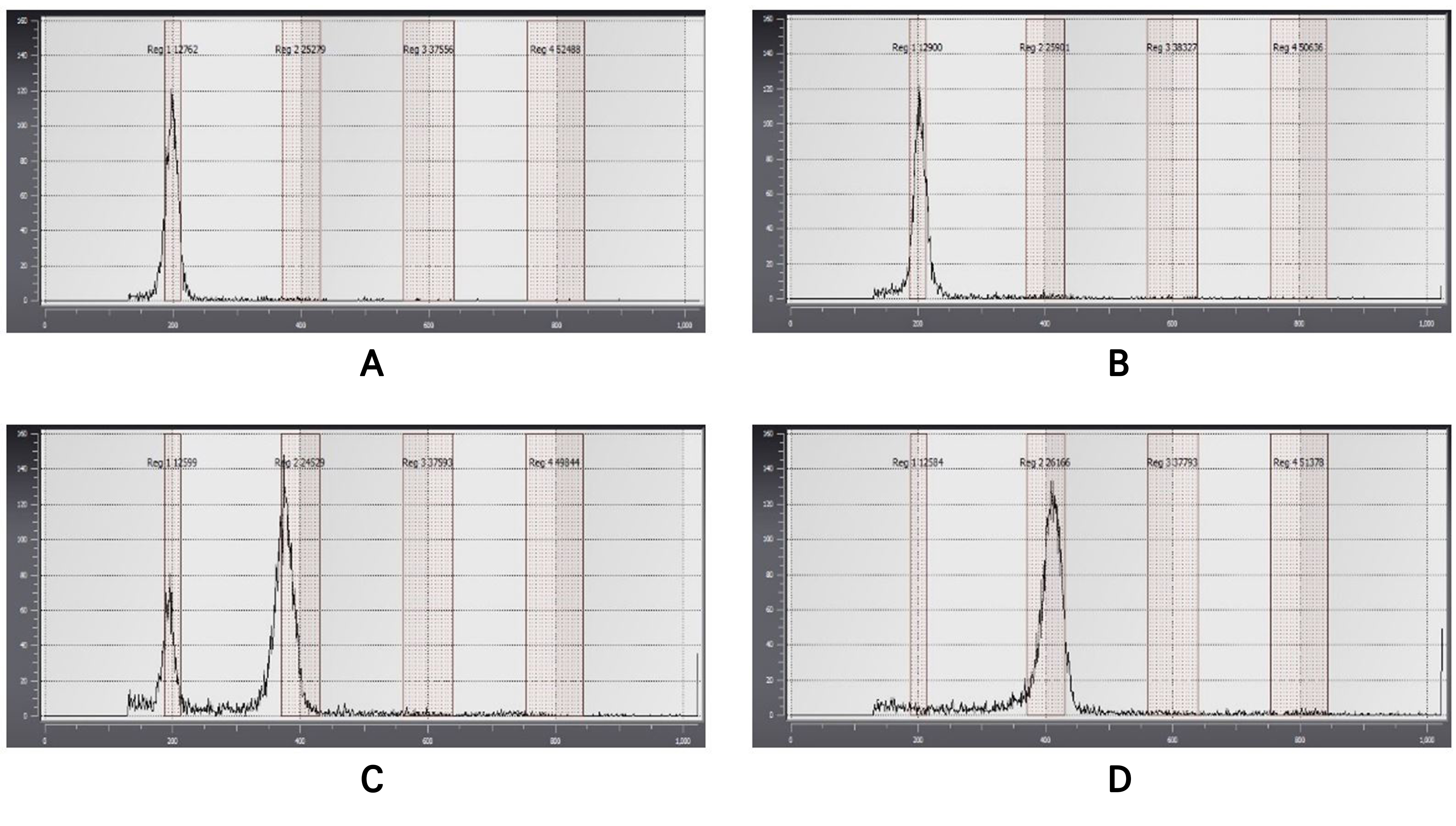
Chromosomal effects of the oryzalin concentration and treatment duration
The effects of different oryzalin concentrations and treatment durations on polyploid formation were investigated. After the emergence of the first true leaves post-sowing, oryzalin was applied to the shoot apical meristem of 864 plantlets, of which 336 survived. A flow cytometry analysis of the surviving plants was then conducted to assess their ploidy levels relative to the ploidy levels exhibited by the control plants (Fig. 2A). The results of the flow cytometry analysis revealed that 151 plants were diploids (Fig. 2B), 122 were chimeras (Fig. 2C), and 63 were tetraploids (Fig. 2D). The number of surviving plants and the induction ratio of tetraploids varied depending on the treatment concentration and period. When treated with 50 mg·L-1 of oryzalin for three days, the highest tetraploid induction rate of 27.78% was observed (Table 1 and Fig. 3). Similarly, the highest number of 118 surviving plants was observed at the oryzalin concentration of 50 mg·L-1. We also confirmed that the number of surviving plants decreased when the oryzalin concentration exceeded 50 mg·L-1 (Table 1). In summary, these results indicate that an oryzalin concentration of 50 mg·L-1 with a treatment period of three days is the most effective condition for inducing polyploidy in P. grandiflorum.
Table 1.
Results of the polyploid induction treatments based on the oryzalin treatment period and concentration
Values represent the number of individuals for each treatment, conducted with 24 individuals per treatment in triplicate. The distribution ratio represents the proportion of the number of surviving individuals to the total number of treated individuals for each replicate. Tetraploid distribution ratios were expressed as means ± standard errors (SE).
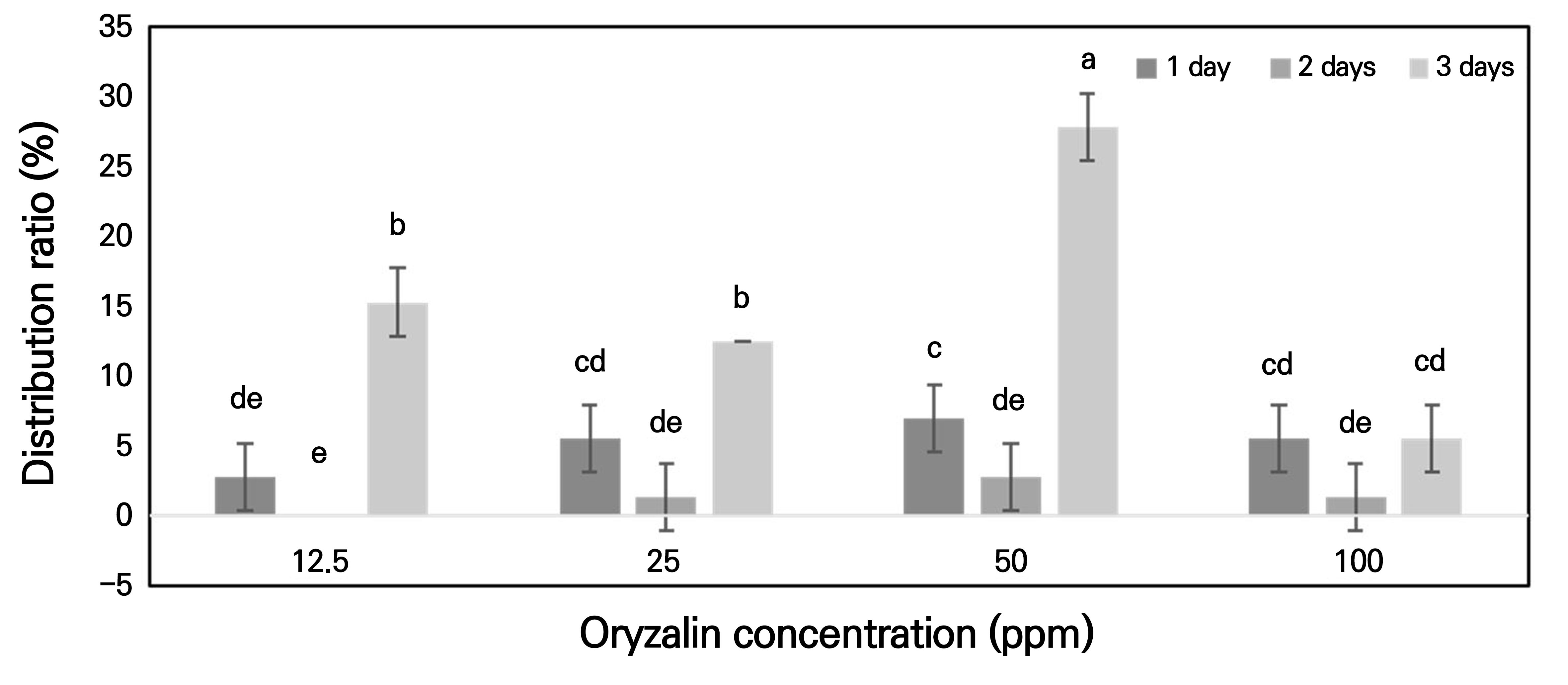
Fig. 3.
Distribution ratios of polyploids based on the oryzalin treatment concentration and treatment period. Mean values sharing the same letter(s) were not significantly different in Duncan’s multiple comparison range test (p < 0.05). Each treatment was repeated thrice, with at least 20 replicates per treatment. Error bars represent standard deviation.
Induction of doubled haploids by means of an oryzalin treatment
DHs were induced by treating haploids derived from a shed-microspore culture with oryzalin. Only plants confirmed to be haploids via a ploidy analysis using flow cytometry were subjected to the oryzalin treatment. For the induction of DHs, oryzalin was applied at a concentration of 50 mg·L-1, the most effective concentration identified in the previous polyploid induction experiment, with 5 µL of the oryzalin treatment solution applied daily for three days. The results of the flow cytometry analysis of the oryzalin treatment groups compared to those for the diploid control group (Fig. 4A) revealed that 238 DHs (Fig. 4B) were obtained from 564 haploid plants. In addition, haploids without chromosomal doubling (Fig. 4C) and chimeras (Fig. 4D) with diploid and haploid characteristics in the same plant were obtained.
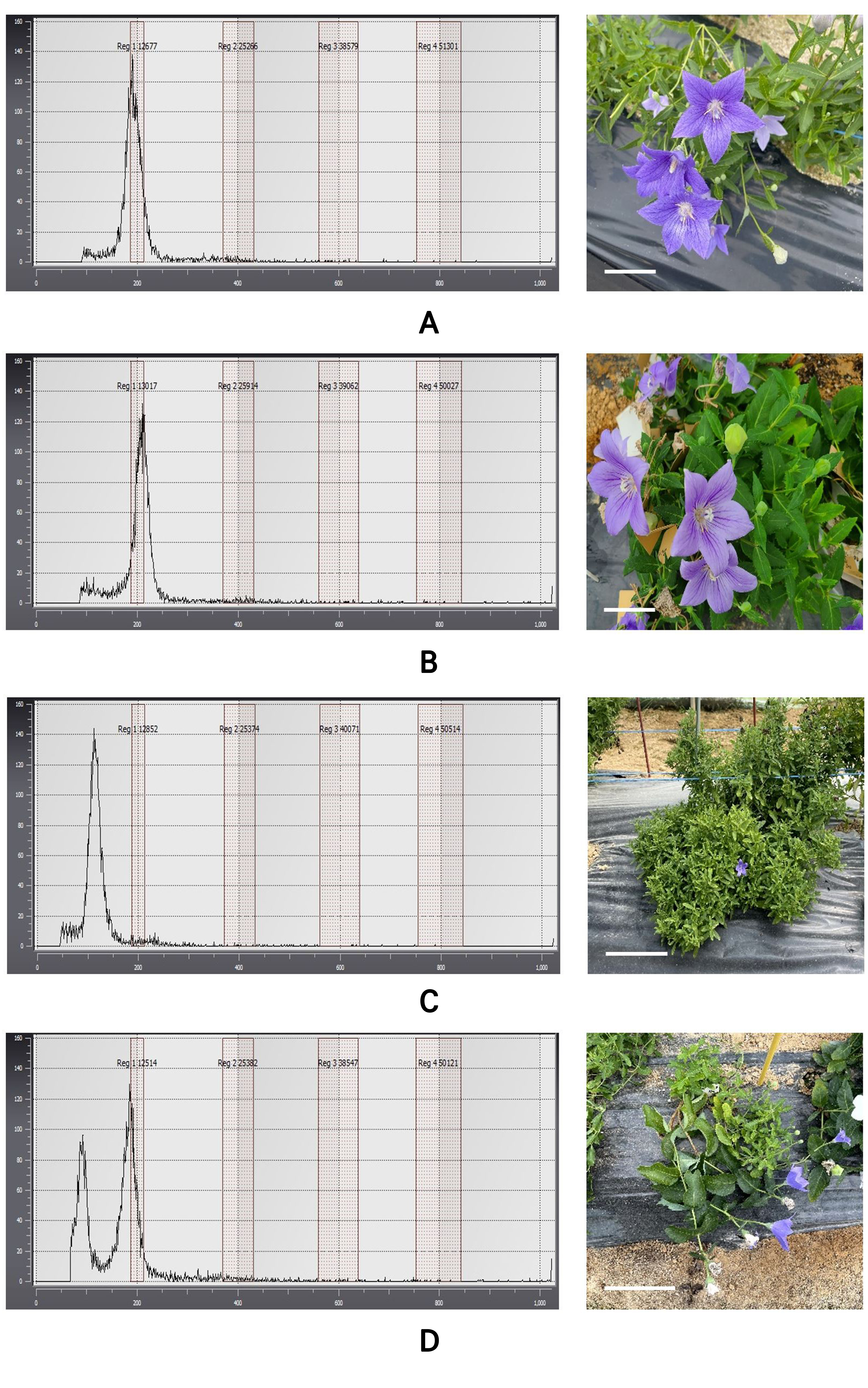
Fig. 4.
Ploidy analysis histogram and plant characteristics of diploids (control), doubled haploids and chimeras in P. grandiflorum: (A) diploid (control); (B) doubled haploid; (C) haploid without chromosome doubling after oryzalin treatment; (D) chimera exhibiting the characteristics of both haploid and diploid. Scale bars represent 3 cm (A, B) and 10 cm (C, D).
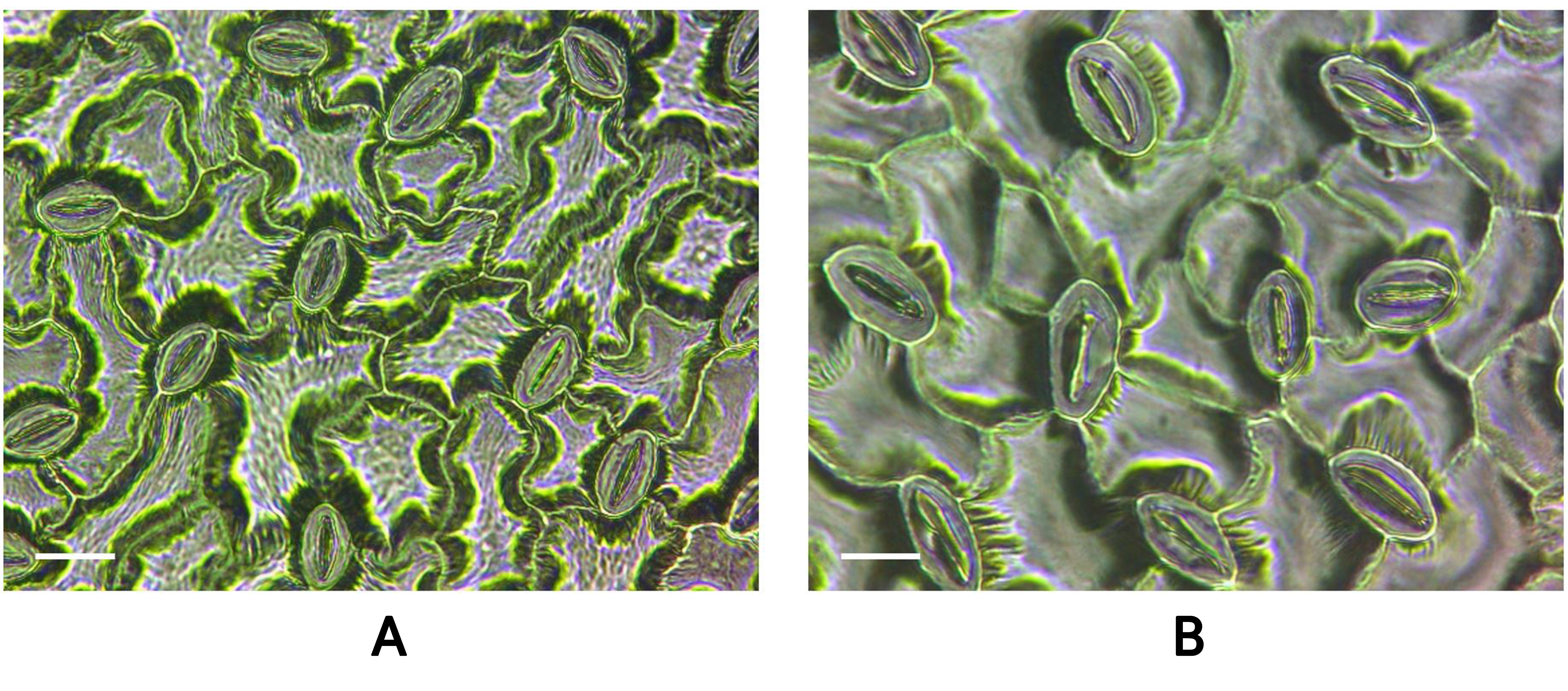
Comparison of the leaf and stomatal characteristics of diploid and tetraploid P. grandiflorum plants
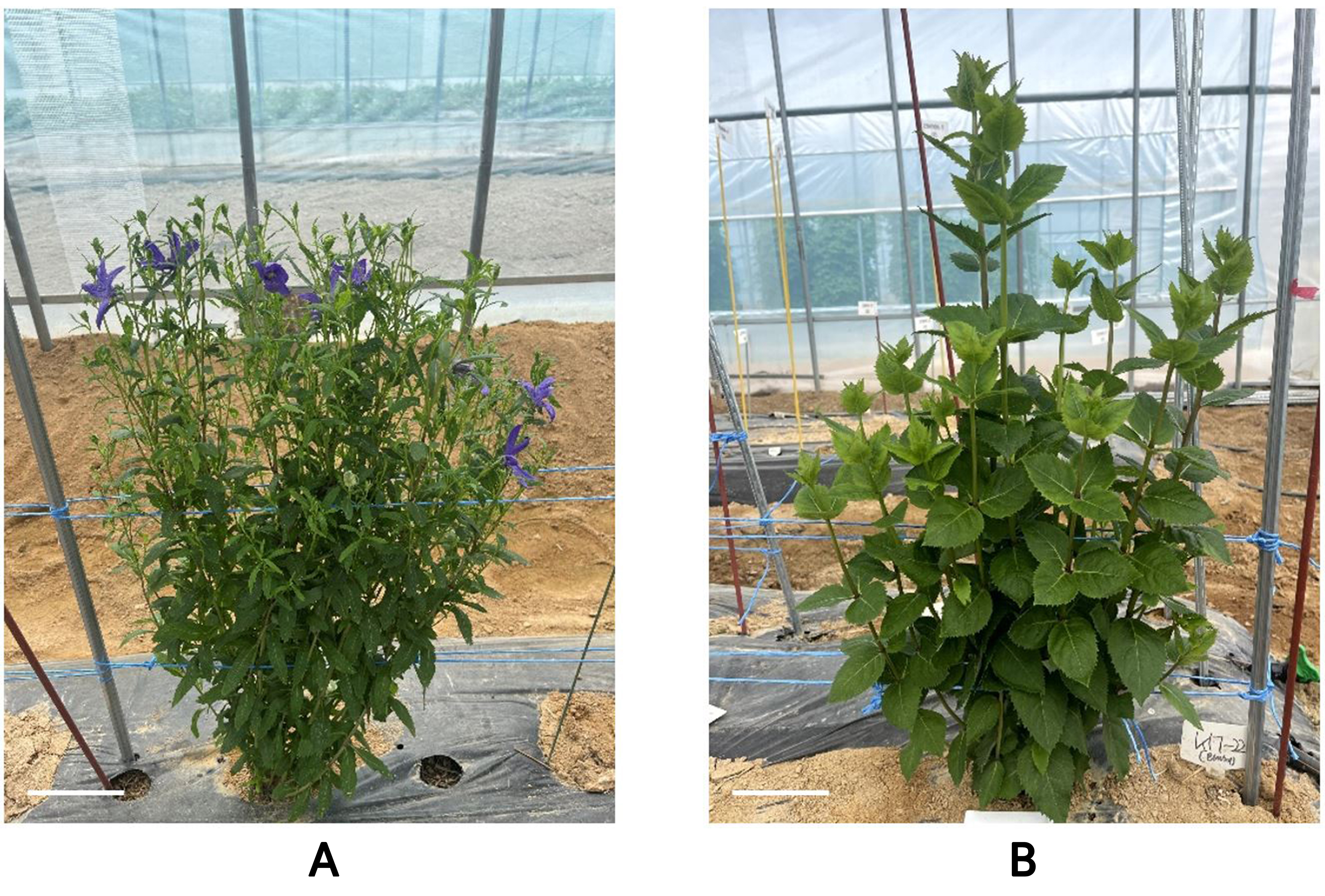
The control diploid plants (Fig. 5A) and tetraploid plants obtained after the oryzalin treatments (Fig. 5B) differed in terms of their leaf characteristics. The leaf length of the diploid plants was 3.79 ± 0.14 cm, whereas that of the tetraploid plants was 6.51 ± 0.16 cm, indicating a significantly greater leaf length in tetraploids compared to diploids (Table 2). The leaf area of the diploid plants was 3.55 ± 0.49 cm2, whereas that of the tetraploid plants was 21.08 ± 1.25 cm2, indicating a significantly larger leaf area in tetraploids than in diploids (Table 2). Additionally, differences in stomatal characteristics were observed between the diploid controls (Fig. 6A), and tetraploid P. grandiflorum plants (Fig. 6B). The stomatal length of the diploid control averaged 25.65 ± 0.18 µm, while that of the tetraploid plants averaged 36.72 ± 0.34 µm, indicating significantly larger stomata in the tetraploids compared to the diploids (Table 3). In contrast, the stomatal density per unit area was 11.13 ± 0.44 in the diploids and 7.5 ± 0.42 in the tetraploids, indicating that the diploids had approximately 1.48 times more stomata per unit area than the tetraploids (Table 3).
Table 2.
Leaf length and area of diploid (control) and tetraploid P. grandiflorum
| Ploidy Level | Leaf Length (cm) | Leaf Area (cm2) |
| Diploid (control) | 3.79 ± 0.14 a | 3.55 ± 0.49 a |
| Tetraploid | 6.51 ± 0.16 b | 21.08 ± 1.25 b |
Table 3.
Stomata length and number of stomata in diploid and tetraploid P. grandiflorum
| Ploidy Level | Stomata Length (µm) | Number of Stomata |
| Diploid (control) | 25.65 ± 0.18 b | 11.13 ± 0.44 a |
| Tetraploid | 36.72 ± 0.34 a | 7.5 ± 0.42 b |
Evaluation of doubled haploid characteristics and selection of superior lines
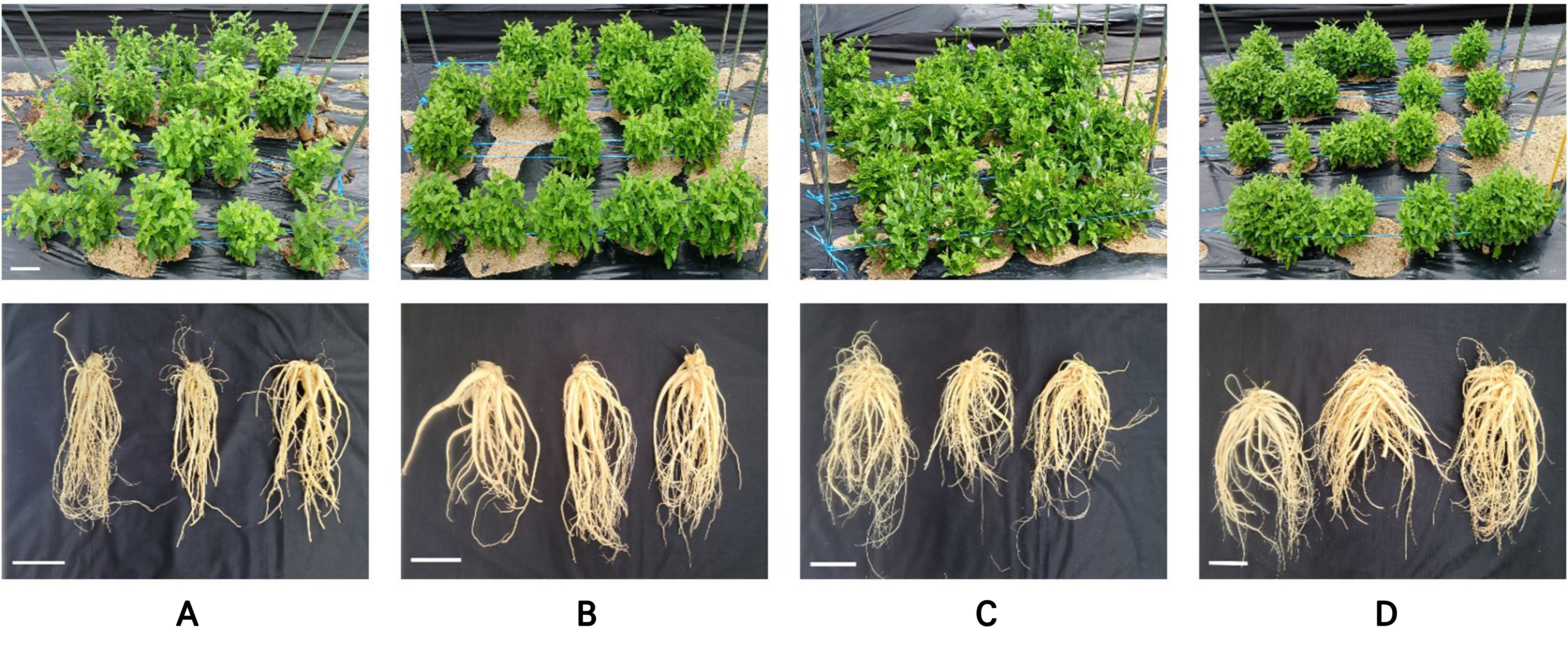
The 238 individuals identified as DHs in the ploidy analysis were transplanted into the greenhouse, and seeds were harvested from 116 plants. The seeds harvested from the shed-microspore-derived DHs were sown and transplanted to evaluate the plant growth characteristics of these DHs (Table 4). Plant characteristics were evaluated by comparing the induced lines with the diploid cultivar ‘Jangbaek’, developed by the Rural Development Administration (RDA). Aboveground traits, in this case the plant height, leaf shape, and leaf size, were assessed. Given that P. grandiflorum is primarily cultivated for its roots, eleven superior lines were selected based on the root weight, root dry weight, and root length compared to the diploid control cultivar ‘Jangbaek’: 22H63-1, 22H1131-5, 22HM471-5, 20H12-4, 20H93-5, 20H94-7, 20H95-15, 20H97-1, 20H103-1, 20H213-2, and 20H113-4. Additionally, five superior lines, 22H711-2, 20H2-1, 20H29-1, 20H76-2, and 20H212-3, were selected based on the root dry weight and root length. The selected lines exhibited uniform characteristics without segregation, indicating that they could be classified as fixed (pure) lines (Fig. 7).
Table 4.
Characteristics of major doubled haploid lines of P. grandiflorum
Discussion
This study developed an optimized oryzalin-based protocol that can be used to induce polyploids and doubled haploids (DHs) in P. grandiflorum, achieving the highest induction efficiency at 50 mg·L-1 applied for three days (Table 1). Of 216 treated plants, 118 survived at this concentration, compared to 47 at 100 mg·L-1, indicating a concentration-dependent trade-off between induction efficiency and plant viability, consistent with findings in cucumber at 75 mg·L-1 (Ebrahimzadeh et al. 2018). Compared to colchicine, which often causes cytotoxicity and abnormal development (Lim and Earle 2008; Lim and Earle 2009), oryzalin minimized adverse effects, highlighting its suitability for medicinal crops and its potential applicability to other cross-pollinated species. These results provide evidence that the proposed method is an efficient means of rapidly developing homozygous lines in P. grandiflorum, a cross-pollinated medicinal crop with high heterozygosity, significantly reducing the breeding time.
In this study, the polyploid induction efficiency was 27.78%, and out of 564 haploid plants treated, 238 were successfully doubled to produce DH lines. This indicates that nearly half of the haploid plants could be recovered as stable DHs, demonstrating the practical applicability of this approach. For example, induction efficiency rates reported in different crops vary considerably. In cucumber, an oryzalin treatment achieved chromosome doubling efficiency rates as high as 92.31% (Ebrahimzadeh et al. 2018), whereas in watermelon the efficiency was around 25% (Bae et al. 2020), while in Brassica napus the efficiency after an oryzalin treatment averaged 66.53% (KlíMa et al. 2008), indicating substantial variation among species. Thus, the efficiency obtained in P. grandiflorum is within the range observed in other species, underscoring the significance of our findings in the broader context of polyploid and DH induction.
Flow cytometry is a reliable method for confirming chromosome doubling and has been used to demonstrate genetic stability in plant tissue culture studies (Ochatt et al. 2011). It has been successfully applied to confirm polyploid induction in various medicinal plants, including ginger bush (Tetradenia riparia) (Hannweg et al. 2016), Brazilian ginseng (Pfaffia glomerata [Spreng.]) (Gomes et al. 2014), Cannabis sativa L. (Bagheri and Mansouri 2015), and Thymus persicus (Tavan et al. 2015). In this study, flow cytometry confirmed ploidy levels, with diploid controls peaking at channel 200, haploids at channel 100, and oryzalin-induced tetraploids and DHs at channels 400 and 200, respectively (Figs. 2 and 4). Chimeric individuals were also detected (Figs. 2C and 4D), providing baseline data for morphological analyses. Tetraploids exhibited significantly larger leaf sizes and areas compared to diploids (Table 2), aligning with reports of enhanced organ sizes in polyploids (Lin et al. 2010). Stomatal morphology further validated ploidy differences, with polyploids showing larger stomata (36.72 ± 0.34 µm) and lower density levels than diploids (25.65 ± 0.18 µm; Table 3), serving as reliable supplementary indicators (Majdi et al. 2010; Gantait et al. 2011; Gomes et al. 2014; Xu et al. 2016).
DH lines from microspore cultures displayed notable phenotypic variation of the leaf morphology, flower shape, root length, and biomass (Table 4). This diversity, likely due to genetic recombination during meiosis (Ferrie and Möllers 2011; Mercier et al. 2015), enables the selection of superior lines with desirable traits. Critically, several DH lines produced viable seeds via self-pollination, confirming their reproductive capacity and value as breeding materials. This is particularly significant for P. grandiflorum, where high heterozygosity hinders trait fixation through conventional breeding.
Doubled haploid lines are expected to show high levels of genetic stability and uniformity, as has been reported in cotton (Mahill et al. 1984) and wheat (Zhang et al. 2025). Such stability ensures that desirable traits are reliably inherited across generations. However, initial vigor levels can vary depending on the crop, with barley DH lines, for example, occasionally indicating an incomplete morphology or reduced vigor during early development but recovering and showing normal growth after regeneration and field establishment (Forster and Thomas 2005). DH lines have consistently been proven to accelerate cultivar development by eliminating segregation in subsequent generations (Chaikam et al. 2019; Zargar et al. 2022). The DH lines obtained in this study thus represent valuable resources for breeding programs in P. grandiflorum.
Superior doubled haploid (DH) lines were selected primarily based on mean value differences in root-related traits. Although DHs possess complete homozygosity and are therefore genetically stable, it is often observed that their growth and fertility gradually recover or stabilize through successive generations (Szadkowski et al. 2010; Rauf et al. 2021; Cui et al. 2023). Accordingly, in this study, superior lines were identified not only by comparing mean trait values but also by evaluating uniformity within each line. The selected DH lines are currently being advanced for an additional two to three generations to ensure further stabilization, ultimately to be compared with commercial cultivars to confirm whether statistically significant differences exist.
Due to their complete homozygosity, they can be directly utilized as parental lines in hybrid breeding or advanced as stable candidates for new cultivar development. Furthermore, once molecular markers associated with important traits such as the root yield, bioactive compound accumulation, and stress tolerance are identified, these DH lines can serve as reference materials for marker-assisted selection (MAS). The combination of DH technology and MAS will enable more efficient and precise breeding, ultimately accelerating the development of elite cultivars in this medicinal crop.