Introduction
Materials and Methods
Plant material and DNA extraction
Phenotype analysis
Evaluation of SlMYB12 gene-based SCARs
Cloning of SlMYB12
SNP array analysis and CAPS marker development
Genetic association analysis
Results
Inheritance of fruit peel color
SCAR marker evaluation and cloning of SlMYB12
Genetic mapping of fruit peel color using SolCAP SNPs
Phenotyping of 64 SolCAP tomato germplasm
Discussion
Introduction
Domesticated tomato (Solanum lycopersicum L.) belongs to the Solanum section Lycopersicum, which is composed of four taxonomic groups. These groups include 13 wild species and their relatives that have diversified in Western South America approximately 12 million years ago (Caicedo and Peralta, 2013). Cultivated tomatoes are among the most widely consumed vegetables in the world owing their potential health benefits and dietary value. The net global production of tomatoes was 164 million tons in 2013 (FAO, 2012). Due to its short generation time and easy growth, tomato also serves as a model for genetic and biological investigations of fruit development and physiology in plants (Kimura and Sinha, 2008).
The color of tomato fruit has received a great deal of interest from a breeding point of view because of its aesthetic effects, as well as its antioxidant characteristics of pigmentation. Different fruit colors are preferred depending on regional consumer culture, with red tomatoes being popular in Europe and Western countries and pink tomatoes preferred in Asia (Ballester et al., 2010). Fruit color is mainly determined by flesh and peel color, which are composed of various flavonoids, carotenoids, and chlorophylls.
Flavonoids are a group of polyphenolic compounds con-tain-ing aglycone that can be classified into chalcones, aurones, flavonols, and anthocyanins (Ballester et al., 2010). Flavonoids lead to yellow, red or blue pigmentation in petals to attract pollinator animals, and may influence disease resistance (Treutter, 2005) or UV filtration (Harborne and Williams, 2000) in plants. Flavonoids predominantly exist in the pericarps (fruit peel) of tomatoes rather than the flesh (Ballester et al., 2010). Unlike flavonoids, carotenoids mainly exist in flesh in the form of all-trans-lycopene, showing reddish color in tomato during ripening (Ballester et al., 2010). As a precursor for abscisic acid in plants (Grotewold, 2006), carotenoids exert photoprotective effects in chloroplasts to prevent auto-oxidation during photosynthesis (Choudhury and Behera, 2001). It has been reported that the external color of red tomato is determined by yellow-colored narin-genin chalcone (NGC) of flavonoids in red tomato peels (Ballester et al., 2010). NGC, which is one of the most abundant flavonoids in tomato fruit peel, accumulates naturally in the cuticle of red tomato skin upon ripening and is responsible for the yellow-colored peel (Hunt and Baker, 1980). Con-versely, pink tomatoes are caused by the presence of a transparent epidermis lacking the yellow pigment NGC (Lindstrom, 1925).
Genetic research has shown that pink tomatoes result from the monogenic, recessive y (yellow) locus, while red tomatoes have the dominant Y allele (Rick and Butler, 1956). SlMYB12, the gene encoding the transcription factor, confers a high level of flavonoids in tomato peel. A genetic association study showed that SlMYB12 is located on chro-mosome 1 and segregates perfectly with the characteristic pink fruit color in a wild tomato species (Ballester et al., 2010). Virus-induced gene silencing of SlMYB12 resulted in a decrease in the accumulation of NGS in pink-colored tomato fruit and a complementation test indicating that SlMYB12 is the gene for the Y locus controlling external fruit color in tomato (Ballester et al., 2010).
DNA marker technology has led to ground-breaking changes in plant genetics and breeding. There are various methods to apply DNA markers to plant breeding for development of new varieties, most notably marker-assisted selection (MAS). MAS is a selection strategy for individual plants that contain molecular markers closely linked to a target gene (s) or quantitative trait loci (QTL). Although a putative gene for fruit peel color has been cloned (Ballester et al., 2010), molecular markers publicly available for MAS of this trait have not yet been reported. Recent advances in genome sequencing technologies such as next generation sequencing (NGS) have resulted in development of a large scale SNP genotyping array for tomatoes (SolCAP Tomato SNP array on Illumina Infinium platform) based on NGS- derived transcriptome sequences (Hamilton and Buell, 2012; Sim et al., 2012; Thomson et al., 2012). This high-throughput genotyping platform enables the rapid genotyping of 7,720 SNPs in parallel. High-density linkage map construction and discovery of DNA markers for MAS can be facilitated by using this SNP array (Sim et al., 2012).
In the present study, we attempted to evaluate SlMYB12 as a gene controlling the traits of fruit peel in domesticated tomato, and to develop molecular markers that can be practically applied to select fruit peel color. To accomplish this, we investigated the genetic association of SlMYB12 with fruit peel color by genetic mapping and characterization of a SlMYB12-harboring genomic region via SNP information revealed by the SolCAP Tomato SNP array.
Materials and Methods
Plant material and DNA extraction
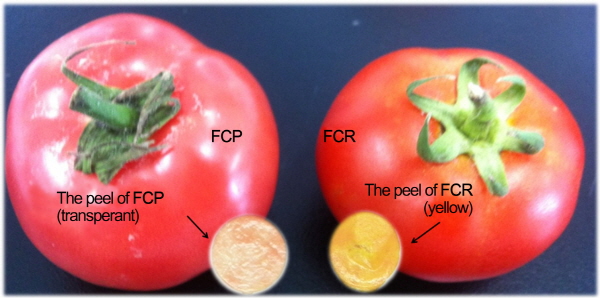
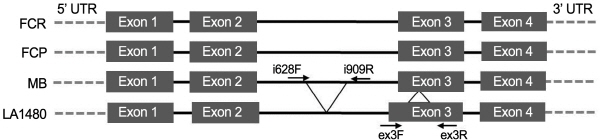
An F2 population was generated for genetic mapping of fruit peel color in domesticated tomato (S. lycoprsicum). The F1 progeny was produced by hand pollination using ‘FCR’ (red-fruited inbred line with yellowish fruit peel) as the maternal parent and ‘FCP’ (pinked-fruited inbred with transparent fruit peel) as the paternal parent (Fig. 1). Subsequently, the F2 generation was produced by controlled self-pollination of a random F1 plant. As a reference array of accessions for marker validation, 64 SolCAP germplasms for which genome-wide SNP genotypes are available from DB were collected and evaluated for their fruit peel color. Genomic DNA extraction and purification was conducted using true leaves according to the method described by Park et al. (2013).
Phenotype analysis
For phenotyping of the F2 progeny, seeds for 272 F2 plants including ‘FCR’, ‘FCP’ and F1 were sown in a 50-cell tray and germinated. Seedlings were grown for four weeks and then transplanted to plastic pots (6 L) filled with soil mixture (Biosengseng growing mix, Seoul Agro-materials, Seoul, Korea). Plants were grown to maturity in a greenhouse under natural light and temperature conditions at Pusan National University (Miryang, Korea) from March to August, 2014. For phenotyping of the SolCAP tomato germplasm, seeds for 64 accessions (four replications per accession) were sown in a 30-cell tray and seedlings were transplanted into soil mixture beds for nutrient solution culture. Plants were grown under natural light and temperature conditions in a plastic film house located at Muju, Korea from April to October, 2014. The mature tomato fruits were harvested at 45-50 days after anthesis. To determine the color of the fruit epidermis, the fruit peel was carefully separated from the flesh using a scalpel and then placed on white paper. After the flesh residue attached to the peel was scratched and removed, the color of the fruit peel was scored as Y for yellowish color or T for transparent (Fig. 1).
Evaluation of SlMYB12 gene-based SCARs
Allelic variations in SlMYB12 (35 bp indel in the second intron and 72 bp indel in the third exon) have been reported for the red-fruit cultivar ‘Moneyberg’ (S. lycopersicum) and the pink-fruit wild species LA1480 (S. chmielewski) (Ballester et al., 2010). A sequence-characterized amplified regions (SCAR) marker (i628F/i909R) has been developed for detec-tion of the 35 bp indel of the second intron by Ballester et al. (2010). To develop an additional SCAR based on the 72 bp indel in the third exon (Table 1, Fig. 2), the genomic DNA sequences of SlMYB12 from ‘Moneyberg’ and LA1480 were retrieved from the Sol Genome Network (SGN) database (http://sol genomics.net/) and aligned using Multialign (http://multalin.toulouse.inra.fr/multalin/). A PCR primer pair (ex3-F/ex3-R) flanking the 72 bp indel was designed using Primer3 (http://bioinfo.ut.ee/primer3-0.4.0/). PCR of this SCAR was conducted in a total volume of 20 μl consisting of 20 ng of genomic DNA, 0.3 μM of each forward and reverse primer, 1X PCR buffer, 0.2 mM dNTPS, and 0.6 U of Taq polymerase (Solgent, Daejeon, Korea). The PCR conditions were as follow: 1 cycle of 5 min at 95°C followed by 35 cycles of 30 s at 94°C, 30s at AT (annealing temperature) (Table 1), 1 min at 72°C, and 7 min at 72°C. Gel electrophoresis was performed using 1.5% agarose gel containing Tris-acetate EDTA (TAE) at 160 V for 1 h 20 min and then visualized under ultraviolet light after ethidium bromide staining.
Cloning of SlMYB12
Genomic DNA sequencing of SlMYB12 from ‘FCR’ and ‘FCP’ was performed to identify DNA sequence polymorphisms between these parental lines and to develop a gene-based marker (Table 1, Fig. 2). Four PCR primer pairs were designed to amplify different genomic regions encompassing SlMYB12, including the 5’ and 3’-untranscribed region (UTR) and possible promoter sequence (700 bp upstream of 5’-UTR and 800 bp downstream of 3’-UTR) (Table 1). After PCR following the protocol described above, the amplified DNA fragments were eluted from agarose gel using ExpinTM Gel SV (GeneAll Biotechnology, Seoul, Korea), after which the purified PCR amplicons were cloned using the pGEM-T- Easy Vector (Promega, Madison, WI, USA) and Hit-DH5α competent cells (RBC, Banqiao, Taiwan). Plasmid DNA was then harvested using the GeneAll Plasmid kit (GeneAll Biotechnology, Seoul, Korea), after which the insert DNA was sequenced using the dye-termination method by Genotech (Daejeon, Korea).
SNP array analysis and CAPS marker development
Genomic DNA of ‘FCR’ and ‘FCP’ was genotyped using the SolCAP Tomato SNP array on the Infinium platform (Illumina Inc., San Diego, CA, USA) to screen for SNPs in a genomic region carrying SlMYB12. Genotyping of the SolCAP array was carried out by SNP Genetics Inc. (Seoul, Korea). The annotation information matrix for SolCAP SNPs was downloaded from the SGN database. Four SNPs (solcap_snp_sl_13762, 456, 38123 and 531) located proximal to SlMYB12 were selected and converted to cleaved am-plified polymorphic sequences (CAPS) markers (CAPS-13762, 456, 38123 and 531) using the CAPS designer (http:// solgenomics.net/) (Table 1). PCR amplification was per-formed as described above. MnlI and PvuII restriction enzyme digestion of the PCR products was conducted according to the manufacturer’s instructions (NEB, Ipswich, Suffolk, England). Gel electrophoresis was performed using 2.5% agarose gel containing Tris-acetate EDTA (TAE) at 160 V for 1 h 20 min, then visualized under ultraviolet light after ethidium bromide staining.
Genetic association analysis
Genetic linkage analysis was carried out to identify the locus for fruit peel color (y) and its association with the CAPS using JoinMap version 4.0 at an LOD value of 3.0, after total of 272 F2 plants were genotyped by four SlMYB12- linked CAPS markers (CAPS-13762, 456, 38123, and 531). Genetic distances between loci were calculated using the Kosambi mapping function. In addition, the genetic relatedness of eight pink fruit accessions and 56 red fruit accessions of the SolCAP tomato germplasm with ‘FCR’ and ‘FCP’ was evaluated by drawing dendrogram based on the genotypes for seven SolCAP SNPs surrounding the SlMYB12. Nucleotide sequences of 64 SolCAP germplasm on these seven SNPs were obtained from SolCAP germplasm SNP data (http:// solcap/msu/edu/) to identify applicability of the markers for other germplasms. Pairwise similarity coefficients among the accessions were calculated according to the method described by Nei and Li (1979). Cluster analysis was per-formed based on marker data using the unweighted pair group method on arithmetic average (UPGMA). All statistical analyses and phenetic tree construction were conducted using the NTSYS_PC version 2.02k software (Rohlf, 2002).
Results
Inheritance of fruit peel color
Fruit peels of the matured red tomato line ‘FCR’ were yellowish, while those of the matured pink tomato line ‘FCP’ were transparent (Fig. 1), indicating that the pink fruit appearance of ‘FCP’ may be due to the absence of yellow-colored flavonoid NGC in epidermal cells, which might be caused by malfunction of SlMYB12. Upon phenotypic analysis of 272 F2 progenies from the cross ‘FCR’ × ‘FCP’, 208 F2 plants showed the fruit peel trait for ‘FCR’ (yellowish, Y), while 64 showed the ‘FCP’ (transparent, T) trait (Table S1). The segregation ratio in this F2 population followed a 3:1 Mendelian ratio (χ2 = 0.31, p <0.05) for single dominant gene model, suggesting that the yellow tomato peel trait of ‘FCR’ is controlled by a single dominant allele, Y.
SCAR marker evaluation and cloning of SlMYB12
In a previous study (Ballester et al., 2010), a mutant allele (y) of SlMYB12 possessing indels in the second intron and third exon was reported for a wild species, LA1480 (S. chmielewski), producing pink fruits and a SCAR marker amplifying a 35 bp indel in the second intron was developed (Fig. 2). In our study, we developed a second SCAR that amplifies a 72 bp indel in the third exon (Table 1, Fig. 2). These two SCARs were genotyped for the prenatal lines ‘FCR’ and ‘FCP’ and both lines showed markers for the red wild-type allele (Y) and no polymorphisms were detected, indicating that the transparent fruit peel of the domesticated pink tomato lines was not determined by the indel mutations of SlMYB12 (Fig. 2). Thus, in the search for other SlMYB12 mutations in domesticated lines (S. lycopersicum), we sequenced the SlMYB12 alleles including the genomic sequence from 700 bp upstream of the 5’-UTR to 800 bp downstream of the 3’- UTR of the ‘FCR’ and ‘FCP’ genes. However, no sequence variations were detected and the whole length of 2.5 Kb was completely identical. Consequently, this hampered the development of a gene-based marker that could be useful for assessing direct the genetic association between fruit peel color and SlMYB12 in the elite Korean breeding lines (‘FCR’ and ‘FCP’, S. lycopersicum). Therefore, we screened for sequence variations within other genes neighboring the SlMYB12 using the SolCAP Tomato SNP array, then used them as anchored markers for mapping the fruit peel color.
Genetic mapping of fruit peel color using SolCAP SNPs
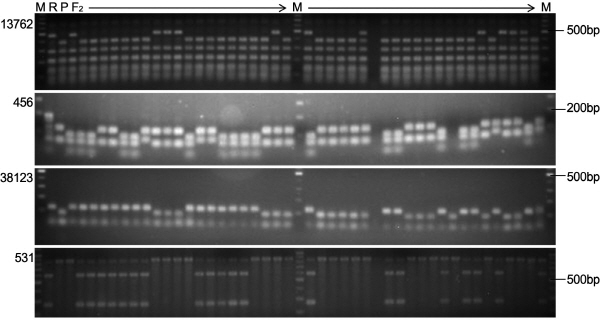
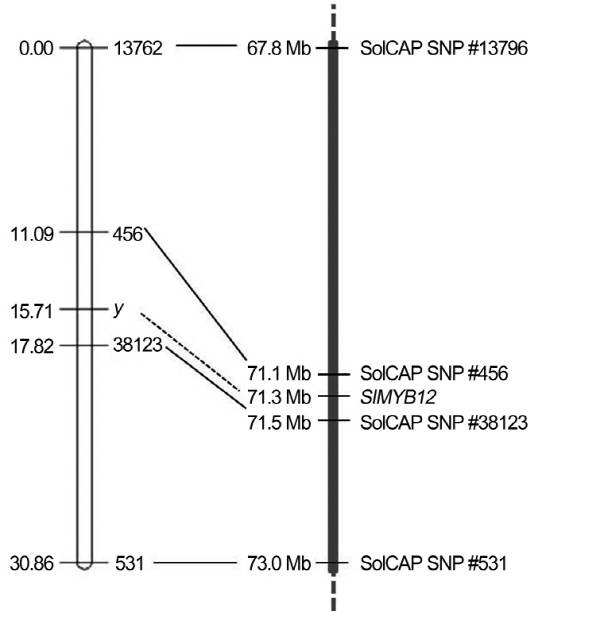
To assess the genetic association between SlMYB12 and fruit peel color in the F2 progeny, we conducted a genome- wide high-throughput SNP discovery from ‘FCR’ and ‘FCP’ using the SolCAP Tomato SNP array. The array-based geno-typing platforms permitted rapid scoring of 7,720 SNPs in parallel (Sim et al., 2012) and resulted in the detection of 1,631 polymorphic SNPs between ‘FCR’ and ‘FCP’. Of those SNPs, 55 were detected from chromosome 1, and 11 of those that were physically anchored in a 10 Kb region harboring SlMYB12 were selected for development of CAPS markers. Among the 11 SNPs, four SNPs proximal to SlMYB12 [solcap_snp_sl_13762 at 67,855,513 bp, 456 at 71,090,567 bp, 38123 at 71,476,848 bp, and 531 at 73,023,201 bp on the pseudochromosome 1 of the reference genome (http://solgenomics.net/, ITAG2.3 release) based on genomic annotation retrieved from the SolCAP Tomato Infinium SNP annotation table (http://solcap/msu/edu/)] were located inside the restriction enzyme recognition sites and successfully converted to CAPS (Table 1). These CAPS, 13762, 456, 38123, and 531 were genotyped on the F2 population (‘FCR × ‘FCP’) and showed the rate of concordance with phenotype, 89.7, 94.1, 97.4, and 89.2%, respectively (Table S1, Fig. 3). A genetic linkage map composed of the four CAPS and fruit color locus Y was constructed (Fig. 4). In this partial genetic map spanning 30.86 cM, the order of four CAPS markers was determined to be 13762-456-38123-531. The Y locus was mapped in the middle of two proximal CAPS-456 (genetic distance from Y = 4.62 cM,) and CAPS-38123 (2.11 cM). We then anchored the physical location of SlMYB12 (SGN, 40ch01: 71,255,600-71,258,882) on this genetic map and found that SlMYB12 was also physically located between solcap_snp_sl_456 (165,033 bp from the start codon) and 38123 (217,966 bp from the stop codon) (Fig. 4). These findings imply that, despite no sequence mutations being detected in SlMYB12, this gene is still a possible candidate (Y) for controlling fruit peel color of domesticated tomatoes. In addition, although CAPS-456 and 38123 did not perfectly cosegregate with the Y locus, these flanking markers would be ideal for selection of fruit peel color. Therefore, we assessed the utility of these markers by evaluating 64 SolCAP tomato accessions of diverse genetic backgrounds.
Phenotyping of 64 SolCAP tomato germplasm
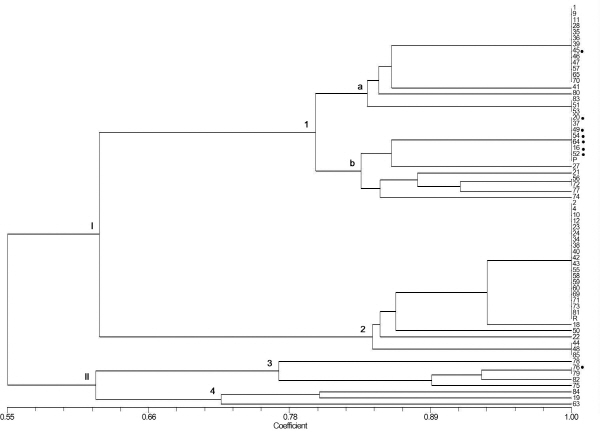
Genetic association between fruit peel color and SNPs in the SlMYB12-harboring genomic block flanked by CAPS-456 and 38123 (40ch01:71090567…71476848) was evaluated using the genome-wide SNP information of 62 SolCAP accessions and two parental lines (Table S2). Fruit peel color of each SolCAP accession was phenotyped in this study. Eight accessions produced pink fruits with transparent epidermal peels (y), while all other accessions produced yellow, orange, or red fruits with yellowish epidermal peels (Y) (Table S2). SNP information describing these germplasms was retrieved from the SolCAP Tomato Infinium SNP anno-tation table (http://solcap/msu/edu/). We found that seven SolCAP SNPs (solcap_snp_sl_456, 457, 38096, 25922, 38116, 38119, and 38123) located in this genomic block were highly conserved specific to pink tomato SolCAP accessions (Table 2). A phenetic tree was then constructed using these seven SNPs to examine the genetic diversity of the 64 accessions, ‘FCR’ and ‘FCP’ relating to this SlMYB12-harboring genomic region. The phenetic tree showed that six pink fruit accessions have the same seven SNPs and clustered as an independent group (subgroup I-2 in Fig. 5), while two other pink fruit accessions were positioned in two different groups (subgroup I-1a and II-3 in Fig. 5). These findings implied that pink tomato accessions evaluated in this study possess a genomic block introgressed, possibly from a common genetic source for the y allele.
Discussion
Tomato fruit color is an economically important trait for aesthetic and antioxidant effects of pigments and for seed markets that is dissected depending on consumer preference for fruit appearance. Pink tomatoes caused by transparent peels lacking yellow pigments were first described in 1925 (Lindstrom, 1925), and the yellow pigment was identified as yellow-colored NGC of flavonoid (Ballester et al., 2010). NGC is a chalcone composed of an aromatic ketone synthesized from 4-coumaroyl-CoA by chalcone synthase (CHS) that is converted into naringenin of flavanone by chalcone isomerase (CHI). CHS gene expression is induced in plants under stress conditions such as UV light or bacterial and fungal infection and may cause accumulation of flavonoid and isoflavonoid phytoalexin in the salicylic acid and defense pathway (Dao et al., 2011). The association of SlMYB12, which is orthologous to the gene encoding MYB12 tran-scription factor in Arabidopsis (Mehrtens et al., 2005) with the peel color locus (Y), has been well documented in wild tomato species accession LA1480 (S. chmielskii) (Ballester et al., 2010). The mutant pink fruit color in LA1480 (y) was attributed to indel mutations in the intron and exon of SlMYB12 and consequently malfunctioned production of NGC in fruit peel (Ballester et al., 2010). Structural phenyl-propanoid and flavonoid gene expression profiling also revealed correlation of the expression levels of SlMYB12 and CHS (Ballester et al., 2010).
Interestingly, no indel mutation or other sequence varia-tions were detected for SlMYB12 in pink-fruited accessions of S. lycopersicum, including potential promoters, implying that transparent peels of modern tomato cultivars were not caused by sequence variation in this gene (Adato et al., 2009; Ballester et al., 2010). These results are in accordance with our finding that cloning of SlMYB12 from ‘FCR’ (red fruit, Y_) and ‘FCP’ (pink fruit, yy) revealed no nucleotide polymorphisms in the genomic region from 700 bp upstream of the 5’-UTR (possibly including promoter) to 800 bp downstream of the 3’- UTRs of the gene. Thus, the direct effects of SlMYB12 promoter on gene expression may not be the cause of y mutation and reduced production of NGC in pink-fruited tomatoes of S. lycopersicum. However, it should be noted that these findings are not conclusive, since characterization of the exact promoter region of SlMYB12 and gene expression studies for pink-fruited S. lycopersicum are still needed.
Development of SlMYB12-based markers was difficult, since no polymorphisms in this gene were detected between ‘FCR’ and ‘FCP’. Instead, we identified several SNPs close to SlMYB12 by genotyping via the SolCAP SNP array. Genetic inheritance and linkage analysis of the F2 population based on those SNPs showed that the pink fruit color of ‘FCP’ may be controlled by the homozygous recessive allele (yy) at the Y locus on chromosome 1. In our genetic map, two SNP markers, CAPS-456 and CAPS-38123, were flanking the Y locus (SlMYB12), and showed cosegregation of 94.1% (4.62 cM) and 97.4% (2.11 cM) with the peel phenotype in the F2 population, respectively. When anchored by physical location, CAPS-456 and 38123 were 165,033 bp and 217,966 bp apart from SlMYB12, respectively. Although CAPS-38123 is located physically further from SlMYB12, its genetic distance is closer to the gene relative to CAPS-456. These findings indicate that chromosomal recombination would occur with lower frequencies between the gene and CAPS- 38123, and that CAPS-38123 can be more efficient for selecting pink fruit tomato.
The SolCAP tomato germplasm genotyped by the SolCAP Tomato SNP array represents a diverse gene pool of dome-sti-cated tomato cultivars (S. lycopersicum) (Sim et al., 2012). In the present study, the SlMYB12-harboring genomic block flanked by CAPS-456 and 38123 (40ch01:71090567…71476848) was evaluated using SNP information and fruit color of 62 SolCAP accessions and two parental lines. Our results showed that seven SolCAP SNPs are located in this block, and these SNPs are highly conserved in pink fruit accessions. Furthermore, a phenetic tree of 64 red and pink accessions constructed based on these seven SNPs demonstrated that most pink accessions grouped together (genetic similarity = 1) independently from all red accessions. These findings indicate that the combination of seven SNPs in the SlMYB12- harboring genomic block are only specific to pink SolCAP accessions tested in this study and provides useful infor-mation for developing SlMYB12-linked markers that could be applicable to diverse breeding programs.
In conclusion, our genetic mapping showed that the y locus located on chromosome 1 confers the transparent fruit peel and pink fruit color of elite tomato cultivars, and that SlMYB12 may be the gene for the locus. Sequencing of SlMYB12 alleles (Y and y) including a potential promoter region revealed no sequence variations between alleles, suggesting that SlMYB12 gene-based marker development cannot be straightforward. Nevertheless, an array of pink fruit-specific SNPs in a SlMYB12-harboring genomic block, as assessed using the SolCAP tomato germplasm, implies that a detailed study of this region may provide useful information for development of markers tightly linked to SlMYB12 and practically applicable for MAS of the peel trait for pink fruit tomato.