Introduction
Materials and Methods
Plant Samples
Chromosome Spread Preparation
Probe Preparation
Fluorescence in situ hybridization
Results
Discussion
Introduction
Several species of the Cucurbitaceae family such as Cucumis sativus (cucumber), Cucumis melo (melon), Citrullus lanatus (watermelon) and Cucurbita spp. (pumpkins) are economically important, generating multi-billion dollars for food, cosmetic and pharmaceutical industry (Jeffrey, 2005; Weng and Sun, 2012). Additionally, other wild cucurbits possess economically and medicinally important traits that can be exploited for crop improvement of related species. For example, some Melothria and Trichosanthes species are good source of proteolytic enzymes and trichosanthin used as anti-HIV drug (McGrath et al., 1989; Lee-Huang et al., 1991; Mayer et al., 1992; Uchikoba et al., 2001; Antão and Malcata, 2005; Qian et al., 2012). Sicyos angulatus, a wild invasive species introduced to Korea in the mid-20th century (Tzonev, 2005), is used as a rootstock for grafting cucumbers and watermelons to
enhance resistance to nematodes and improve yield (Lee, 1994; Shengping et al., 2006).
Cytogenetic studies offer complementary and novel information on genome organization, gene activity and meiotic recombination (Heslop-Harrison, 1991). The development of cytogenetic tools has been highly utilized in breeding programs to manipulate both the genetic make-up and the crop-to-wild gene transmission system (Sain et al., 2002; Sybenga, 2012); including those of wheat, maize (Benavente et al., 2008; Albert et al., 2010; Icso et al., 2014) and watermelon (Sain et al., 2002). Tropical and subtropical cucurbits, like C. melo var flexuosus, M. pendula and T. anguina, are good genetic resources for the breeding of Korean cucurbits considering that the climate of Korean peninsula is gradually changing from temperate to subtropical (Bohnert et al., 1995). Hence, several cucurbits have been improved using cytogenetic-based breeding approach. For example, recent introgression breeding from a wild relative of cucumber, Cucumis hystrix, has increased the genetic diversity of cucumber (Zhuang et al., 2006; Delannay et al., 2010). Additionally, introgression between cultivars and wild relatives of bitter gourds (Momordica charantia) (Liao et al., 2012) and Citrullus (Sain et al., 2002) have also been reported.
In spite of the potential of the exotic cucurbit species of C. melo var. flexuosus (L.) Naudin, M. pendula L., and T. anguina L. as genetic resources for medicinal use and crop production (Lira and Caballero, 2002; Ojiako and Igwe, 2008), cytogenetic-based research investigating chromosome characteristics and genome composition is very limited. Most cucurbit species have small chromosomes that are difficult to distinguish (Waminal and Kim, 2012), however, fluorescence in situ hybridization (FISH) has revealed the identity of chromosome pairs and provided information that can be used in breeding programs (Huang et al., 2009; Waminal et al., 2011; Waminal and Kim, 2012). We have carried out triple-color FISH karyotype analysis with Korean wild or crop cucurbit species that will be useful for phylogenetic comparative studies and breeding programs focusing on Korean Cucurbitaceae family (Waminal et al., 2011; Waminal and Kim, 2012; Waminal and Kim, 2015). Here, we determined chromosome composition and distribution patterns of rDNA and telomeric DNA, employing triple-color FISH analysis, in three exotic cucurbit species of C. melo var. flexuosus (L.) Naudin, M. pendula L., T. anguina L. found in tropical and subtropical habitats providing major repeat-based karyotypes of these species and potentially improve their medicinal and vegetably use.
Materials and Methods
Plant Samples
Three tropical or subtropical exotic cucurbit species were used. The seeds of Armenian cucumber Cucumis melo var. flexuosus (L.) Naudin was purchased from Botanical Interest, Inc. Broomfield; the creeping cucumber or the Guadeloupe cucumber Melothria pendula L. and Trichosanthes anguina L. were collected from Opol, Misamis Oriental Province, Philippines and cultivated field in South Chungcheong Province, Korea, respectively. Each species was identified based on distinct morphological characteristics (Lee, 2003) and confirmed by wild plant taxonomist. Seeds were germinated on moist petri dish at 25°C. Harvested roots (~2 cm in length) were pretreated with 2 mM 8-hydroxyquinoline at 18°C temperature for five hours, fixed in Carnoy’s fixative and stored in 70% ethanol at 4°C until further use.
Chromosome Spread Preparation
Somatic chromosome spreads were prepared according to the procedure of Waminal and Kim (2012). In brief, root tips (-2 mm) were digested in pectolytic enzyme solution [2% Cellulase R-10 (C224, Phytotechnology Laboratories) and 1% Pectolyase Y-23 (P8004.0001, Duchefa) in 100 mM citrate buffer] at 37°C for 75 minutes. The root tips were washed in distilled water and transferred into Carnoy’s solution and vortexed for 30 s. Pellets were resuspended in 9:1 (v/v) aceto-ethanol and subsequently spread on pre-cleaned slides in a humid chamber before being air-dried. Slides were fixed with 2% formaldehyde (Vrana et al., 2012), dehydrated in a series of ethanol treatments (70%, 90% and 100%) and air-dried.
Probe Preparation
The 18S rDNA was obtained from genomic DNA of Triticum aestivum by PCR with primers 5’-AACCTGGTT GATCCTGCCAGT-3’ (forward) and 5’-CCATCAGAACTCCGCAGTTA-3’ (reverse) (Matoba et al., 2007) and labelled through nick translation with DEAC-5-dUTP (Perkin Elmer, NEL455001EA). For 5S rDNA, produced from genomic DNA of Brassica oleraceae by PCR with primers 5’-GATCCCATCAGAACTCC-3’ (forward) and 5’-GGTGCTTTAGTGCTG GTAT-3’ (reverse) (Koo et al., 2002) and labelled by nick translation with Alexa Fluor ® 488-5-dUTP (Invitrogen, Carlsbad, California, USA). Following the methods of Abd El-Twab and Kondo (2006), Arabidopsis-type telomere sequence was PCR-amplified, in the absence of the genomic template DNA, using primers (TTTAGGG)5 and (CCCTAAA)5 and amplicons were labelled with Texas Red-5-dUTP (Perkin Elmer, NEL417001EA).
Fluorescence in situ hybridization
The hybridization mixture consisted of 100% formamide, 50% dextran sulfate, 20×SSC, 50 ng·µL-1 of each DNA probe and DNase-free water (Sigma, W4502). The mixture was denatured at 90°C for 10 min and placed on ice for at least 5 min. 40 µL volume of mixture was placed on microscope slide and sealed with cover slip. Slides were further denatured at 80°C for 5 min and incubated overnight in a humid chamber at 37°C. Following the overnight hybridization, slides were washed in 2×SSC at room temperature for 10 min, 0.1×SSC at 42°C for 25 min, 2×SSC at room temperature for 5 min and before being dehydrated with ethanol series (70%, 90% and 100%) at room temperature. Slides were counterstained with DAPI (1 µg·µL-1) in Vectashield (Vector Labs, H-1000).
Chromosomes were observed under an Olympus BX53 fluorescence microscope (Olympus, Japan) using oil lens (x100 magnification) equipped with a built-in CCD camera (CoolSNAP™ cf). Images were captured and analyzed with Cytovision version 7.2 (Leica Microsystems, Germany) and images were processed using Adobe Photoshop CS6. Chromosomes were measured using ImageJ software (Scheider et al., 2012) and classified according to the methods described by Waminal and Kim (2012). Homologous chromosomes were identified based on their FISH signals, morphological characteristics and length.
Results
We used the three highly conserved DNA repeat sequences in plants, the 45S and 5S rDNAs and the Arabidopsis-type telomeric repeat, as probes for triple-color FISH karyotype analysis. Different distribution pattern on somatic metaphase chromosomes was observed among the three curcubit species. In Cucumis melo var. flexuosus, two pairs of 45S rDNA were observed on the paracentromeric region of the long arm of chromosome 3 and almost on the entire region of the short arm of chromosome 10 (Fig. 1A1). The 5S rDNA loci were observed on the intercalary region of the long arm of chromosome 8 (Fig. 1A2). Telomeric repeats were detected on the terminal region of both short and long arms of all chromosomes (Fig. 1A3).
In Melothria pendula, four pairs of 45S and two pairs of 5S rDNA loci were detected. Three pairs of 45S rDNA were independently localized on the paracentromeric region of chromosome 1, the terminal region of chromosome 2 and the intercalary region of chromosome 7 (Fig. 1B1). The largest 45S rDNA signal was in chromosome 1 (Fig. 2B). The 5S rDNA loci were observed on chromosomes 5 and 11 (Fig. 1B2). In chromosome 5, 5S rDNA was juxtaposed with 45S rDNA on the intercalary region of short arm (Fig. 2B). Li et al. (2016) have reported the presence of only one pair of 5S rDNA and two pairs of 45S rDNA in M. pendula using FISH. This discrepancy is likely due to experimental conditions and poor hybridization of probe that was considered as one major disadvantage of in situ hybridization method (Schubert et al., 1998; Kubota et al., 2006; Cuyacot et al., 2016). Telomeric signals were distributed on all M. pendula chromosome termini (Fig. 1B3).
Different rDNA distribution pattern was observed in the genome of Trichosanthes anguina with three pairs of 45S and two pairs of 5S rDNA loci. The 45S rDNA loci were detected on the terminal region of the long arm of chromosomes 5-7 (Fig. 1C1). One pair of 5S rDNA localized on the intercalary region of the long arm of chromosome 9 (Fig. 1C2). In chromosome 6, the 5S rDNA was co-localized with 45S rDNA on the terminal region of the chromosome (Fig. 3C). Telomeric signals were all detected on the terminal region of all chromosomes (Fig. 1C3).
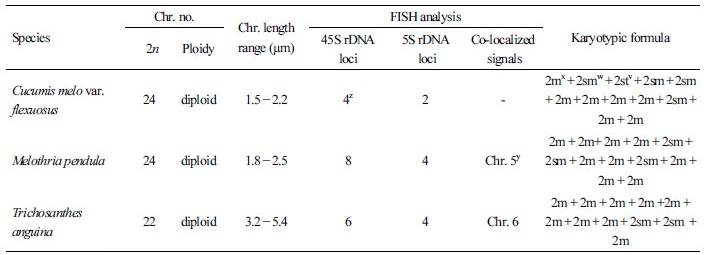
The FISH-based karyotypes of the three exotic cucurbit species were analyzed to elucidate their chromosome compositions. All three cucurbit species were diploid with considerably small chromosomes, with this result being consistent with previous reports investigating different cucurbit species (Waminal et al., 2011; Waminal and Kim, 2012; Waminal and Kim, 2015). The chromosomes were paired and arranged according to the probe signals and length in descending order (Fig. 2). Karyotypic idiograms of the three species are shown in Fig. 3. The chromosome complement of C. melo var. flexuosus was 2n=2x=24, with lengths ranging from 1.5 to 2.2 µm and composed of seven metacentric, four submetacentric and one subtelocentric pairs. In M. pendula, the chromosome complement was 2n=2x=24 with chromosome lengths ranging from 1.8 to 2.5 µm. M. pendula was comprised of nine metacentric and three submetacentric chromosomal pairs. The chromosome complement of T. anguina was 2n=2x=22 with chromosome lengths ranging from 3.2 to 5.4 µm, and it was composed of nine metacentric and two submetacentric homologous pairs. Chromosome number, length, karyotype formula and FISH signal distribution are summarized in Table 1.

Fig. 2. Triple-color FISH karyogram of Cucumis melovar. flexuosus (A), Melothria pendula (B) and Trichosanthes anguina showing chromosomes in descending length order. 45S rDNA, 5S rDNA and telomeric repeat DNA signals are indicated in blue, green and red, respectively. Co-localization of 45S and 5S rDNA signals were observed in chromosome 5 of M. pendula and chromosome 6 of T. anguina, a common feature within Cucurbitaceae family. Scale bar = 5 µm.
Discussion
The three exotic cucurbit species, Cucumis melo var. flexuosus (L.) Naudin (2n=24), Melothria pendula L. (2n=24), and Trichosanthes anguina L. (2n=22), are distributed across the tropical and subtropical Korean peninsula and have been utilized for their medicinal properties and in crop production (Bullock, 1990; Lira and Caballero, 2002; Ojiako and Igwe, 2008; Rai et al., 2008).
FISH procedures, using ribosomal rRNA genes (rDNAs) and other DNA repeat probes, have allowed the characterization of chromosome and genome composition of various species (Leitch and Heslop-Harrison, 1992; Devi et al., 2005; Kato et al., 2005; Belandres et al., 2015). Additionally, FISH is a powerful method to revealing chromosomal rearrangements over evolutionary time and enabling the description of phylogenetic relationships between species (Devi et al., 2005; Coluccia et al., 2011; Li et al., 2016). For example, cytogenetic reports using FISH supported the possible chromosome fusion of ancestral chromosomes in Cucumis sativus (x=7) after divergence from C. melo (Koo et al., 2010), and revealed the close evolutionary distance among cultivated watermelons (Li et al., 2016).
Tandem arrays of 45S and 5S ribosomal DNAs are highly conserved in eukaryotes (Hasterok et al., 2001). Distribution of telomeric repeats, although not as conserved as the rDNA, is of particular interest to describing evolutionary relationship (Blackburn, 1991; Fuchs et al., 1995). Recently, rDNA loci were investigated by FISH in more than 1,000 species (Garcia et al., 2012) providing key information on genome structure and inter-specific relationships that will in turn improve breeding designs.
The family Cucurbitaceae has the most common basic chromosome number of 11 or 12 (Whitaker, 1933; Beevy and Kuriachan, 1996; Schaefer and Renner, 2010; Waminal and Kim, 2015). The three cucurbits, investigated in this study, also verify this chromosome number. The observed variation in chromosome number in Cucurbitaceae family (Whitaker, 1933; Jeffrey, 1967; Jeffrey, 1980) is a common feature among angiosperms (Dodsworth et al., 2015). Polyploidization and subsequent diploidization events are often related to genome amplifications and shrinkage, chromosomal translocations or even loss of chromosomal segments (Bruggmann et al., 2006; Dodsworth et al., 2016). As a consequence, such events are often associated with loss or gain of rDNA signals (Mishima et al., 2002). For example, chromosome fusion of several chromosomes in the ancestral genome of cucumber resulted in the reduction of the chromosome number in C. sativus (x=7) after the divergence from C. melo (x=12) ( Koo et al., 2010). A fusion event is likely to associate with the chromosome number of T. anguina.
The most frequent localization of 45S and 5S rDNA in the three cucurbits species were in distal and interstitial regions, respectively. Independent rDNA position most often leads to a more efficient gene conversion and unequal crossing-over without disruptive interference (Martins and Galetti Jr, 1999; Martins and Galetti Jr, 2001; Koo et al., 2010). The interstitial localization of 5S rDNA loci could stabilize the tandem repeats, whereas 45S rDNA resides on terminal region are prone to dispersion in the genome during chromosomal rearrangement (Waminal and Kim, 2012; Reddy et al., 2013). The polymorphism of rDNA gene loci and merging of some repeats indicate the occurrence of genomic reorganization during speciation within the Cucurbitaceae family (Datson and Murray, 2006; Książczyk et al., 2010, Cuyacot et al., 2017).
The rDNA site distribution of the three cucurbits studied here provide information on their genome structure and organization that may be useful to improve breeding programs of other related cucurbit species (Renner et al., 2007). Advanced cytogenetic molecular analysis employing either cucurbit- or species-specific DNA sequence as cytogenetic markers could refine chromosome identification and further elucidate phylogenetic relationships among species.