Introduction
Materials and Methods
Construction of an Integrated Abiotic Stress-related Gene Expression Network Model
Analysis of Specific Expression of B. rapa Genes During Cold, Salt, or Drought Stress
Analysis of Changes in the Expression of Abiotic Stress Resistance-related Genes in Transgenic B. rapa Using Network Models
Results
Construction of a Gene Expression Network Model for Integrated Expression Analysis of B. rapa Under Cold, Salt, and Drought Stress
Cold Stress-specific Responses
Salt Stress-specific Responses
Drought Stress-specific Responses
Shared Responses to Cold, Salt, and Drought Stress
Characterization of New Abiotic Stress Tolerance-related Genes using the Network Model
Introduction
In the past decade, gene expression data have been generated for model plants and important crops, such as thale cress (Arabidopsis thaliana), rice (Oryza sativa), potato (Solanum tuberosum), and maize (Zea mays). These data have been analyzed to help design forward genetic screens and other studies (Goodstein et al., 2012; Provart, 2012). There is also a growing need to develop in silico platforms to integrate these publicly available data and to construct network models of protein-protein interactions, as well as to correlate gene expression data with plant molecular mechanisms (Serin et al., 2016). These computational analysis methods to identify novel genes involve the identification of correlation patterns
among different mechanisms, environments, and biological processes (Usadel et al., 2009; Provart, 2012). Hence, the importance of gene correlation analysis is expected to increase functional genomics area that genome wide association studies (GWAS).
Several tools and software, such as Pajek (Batagelj and Mrvar, 1998), Cytoscape (Shannon et al., 2003), and Biolayout (Theocharidis et al., 2009), are available for network visualization and analysis. Cytoscape, one of the most highly utilized open-source tools for visualizing co-expression networks, includes a feature extension function using Java programming (Cline et al., 2007). These tools can be used to construct networks to identify genes or proteins showing similar expression patterns (Provart, 2012).
In previous studies aimed at constructing co-expression networks, abiotic stress-related data were collected from the KBGP-24K microarray Brassica rapa EST and microarray database (BrEMD), and time-based expression analyses of B. rapa plants were performed. Topological and functional analysis of gene co-expression networks revealed that abiotic stress is significantly correlated with the categories ‘activation of chitinase-antifreeze protein by WRKY transcription factor and salicylic acid (SA) signaling’ (cold stress; Lee et al., 2015), ‘Na+ transport by reactive oxygen species (ROS) signaling and accumulation of proline’ (salt stress; Lee et al., 2014), and ‘ROS removing process by raffinose family oligosaccharides (RFOs) and the inositol metabolic process’ (drought stress; Lee et al., 2017).
In the present study, we utilized the KBGP-24K microarray BrEMD to obtain microarray information related to cold, salt, and drought stress (RDA, 2008) via statistical analysis of this information, the visualization of relevant genes, and integrated analysis of the correlation of each abiotic stress network. We obtained an integrated and/or independent analytical co-expression network model for abiotic stress, providing basic information about the various important genes involved in abiotic stress responses in Chinese cabbage. Understanding the associated regulatory networks will facilitate research aimed at enhancing abiotic-stress resistance in Chinese cabbage.
Materials and Methods
Construction of an Integrated Abiotic Stress-related Gene Expression Network Model
In the present study, microarray data were compiled and annotated with respect to the responses to cold, salt, or drought stress, as previously described (Lee et al., 2014; Lee et al., 2015). Of these, the group of genes with a correlation value >0.85 in the PlantArrayNet analysis was selected. An integrated abiotic stress-related gene expression network was constructed using the Cytoscape program (version 3.4.0, Cytoscape Consortium; Smoot et al., 2011), and the network structure was visualized using a spring-embedded layout (Barnes and Hut, 1986).
Analysis of Specific Expression of B. rapa Genes During Cold, Salt, or Drought Stress
The constructed network was utilized to analyze specific expression patterns of genes via Venn diagrams. The Venn and Euler diagrams were utilized as a plug-in to Cytoscape to compare cold-, salt-, and drought-specific correlation networks. Changes in plant metabolic pathways involving these genes were ascertained using the KEGG (Kyoto Encyclopedia of Genes and Genomes, http://www.genome.jp/kegg) and TAIR (The Arabidopsis Information Resource, http://www.arabidopsis.org) resources. The DAVID (The Database for Annotation, Visualization and Integrated Discovery, http://david.abcc.ncifcrf.gov) bioinformatics resource was then use to extrapolate the abiotic stress resistance functions of B. rapa by predicting the possible functions of the expressed genes (Huang et al., 2009; Kanehisa et al., 2012). Based on these data, gene ontology analysis and visualization were performed using the Cytoscape plug-in program ClueGO (Bindea et al., 2009).
Analysis of Changes in the Expression of Abiotic Stress Resistance-related Genes in Transgenic B. rapa Using Network Models
Three transgenic Chinese cabbage (Brassica rapa ssp. pekinensis inbred line ‘CT001’) lines were selected for the present study. These transgenic lines, which were obtained by Agrobacterium-mediated transformation, overexpressed B. rapa genes Bra000358, Bra033354, and Bra026701, which were previously identified in plants showing cold, salt, and drought stress resistance, respectively (Yu et al., 2014a; Yu et al., 2014b; Yu et al., 2016). Expression profiling was conducted with the B. rapa genome 135K cDNA chip version 3.0 (NimbleGen, Madison, WI, USA; GreenGene Bio Tech Inc., Yongin, Korea). The microarray was scanned using a GenePix 4000B scanner (Axon Instruments, Union, CA, USA), and the signals were digitalized using NimbleScan software (NimbleGen Systems, Madison, WI, USA). The signals were then normalized using the Loess normalization method and the false discovery rate (FDR) controlling method (Benjamini and Hochberg, 1995), while the microarray data were analyzed against the cold, salt, or drought stress-gene expression network using the Cytoscape program (version 3.4.0, Cytoscape Consortium; Smoot et al., 2011).
Results and Discussion
Construction of a Gene Expression Network Model for Integrated Expression Analysis of B. rapa Under Cold, Salt, and Drought Stress
We constructed an expression network model for integrated abiotic stress response genes in B. rapa using Brassica 300K microarray analysis. We selected B. rapa genes that function in the responses to cold, salt, and drought stress; 4,306 probes were chosen out of a possible 23,937 that were significantly differentially expressed under abiotic stress conditions. To analyze the functions of the genes selected from the network, we performed functional annotation clustering using the DAVID bioinformatics resource (Table 1). Of the 306 functional clusters analyzed, we selected five clusters with the highest enrichment scores. Cluster 1, which had the highest enrichment score among functional clusters, contained genes related to the functional categories ‘Response to abiotic stimulus’, ‘Response to osmotic stimulus’, and ‘Response to salt stress’. The remaining clusters (Clusters 2-5) contained many genes related to ‘Response to inorganic substance’, ‘Response to organic substance’, ‘Response to endogenous stimulus’, and ‘Cell wall’. When a plant is exposed to abiotic stress, such as cold, salt, or drought conditions, the cell membrane induces transient influxes of osmolytes and antioxidants into the cytoplasm, which provide membrane fluidity and lead to cytoskeletal reorganization (Hasegawa et al., 2000; Orvar et al., 2000; Sangwan et al., 2001; Xiong et al., 2002). This finding indicates that the majority of changes induced by cold and osmotic stress are shared by all treatments; a similar finding was obtained through expression profiling analysis of Arabidopsis (Seki et al., 2001; Kreps et al., 2002). These mechanisms are consistent with the functional annotation clusters described above, indicating that the group of probes selected to construct the network was strongly related to abiotic resistance in Chinese cabbage.
In the gene expression network model, the genes constitute the nodes, and a significant correlation in expression levels between genes is represented by an edge (Fig. 1A and 1B). After constructing the network model, we identified and eliminated multi-edge node pairs and self-loop nodes, resulting in a network containing 4,306 nodes, 30,889 edges, and 98 connected components. The largest independent network among the suborder connected components contained 94.37% of all nodes (4,064 nodes) and 99.50% of all edges (30,736 edges), demonstrating that the network model is highly relevant. When we utilized the constructed network model to analyze the expression patterns of significantly differentially expressed cold, salt, and drought stress-related genes among the integrated set of 4,309 abiotic stress-related genes, significant interactions with cold stress (4°C treatment), salt stress (250 mM NaCl treatment), and drought stress (dehydration treatment) were detected for 2,030 genes (47.11%), 1,351 genes (31.35%), and 1,549 genes (35.95%), respectively. We also performed Venn diagram analysis to investigate abiotic stress responses in Chinese cabbage (Fig. 1C and 1D). To analyze the functions of the genes selected from the network, we performed functional classification of the genes using the DAVID bioinformatics resources. Of the functional clusters analyzed, we selected three clusters with the highest enrichment scores (Table S1). However, the data about changes in gene expression values and each stress-specific response in the Venn diagram analysis disagreed, which is in contrast to the results obtained by Lee et al. (2008) from their transcriptome analysis of B. rapa via Brassica 24K oligo microarray analysis using the BrEMD. In their study, the majority of abiotic stress-related genes with >5-fold change at any treatment time point were selected, while in the present study, B. rapa genes that showed an expression correlation value >0.85 in response to cold, salt, and drought stress were selected, which were significantly differentially expressed in each abiotic stress-related co-expression network.
Table 1. Functional annotation and clustering of genes related to cold, salt, and drought stress tolerance in Chinese cabbage 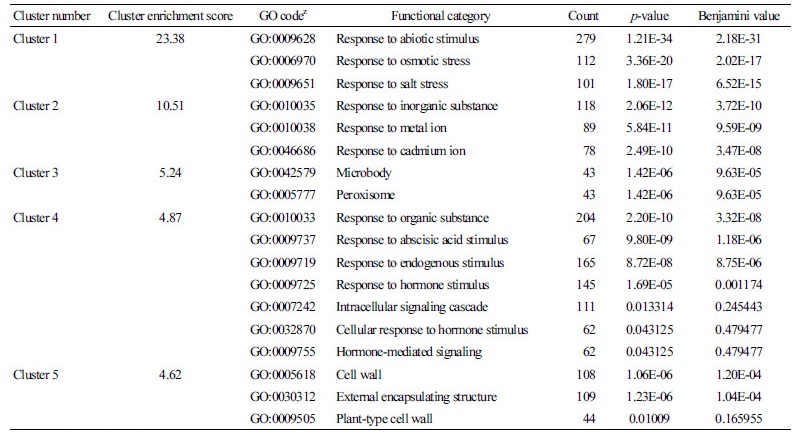 | |
zGene ontology code number. | |
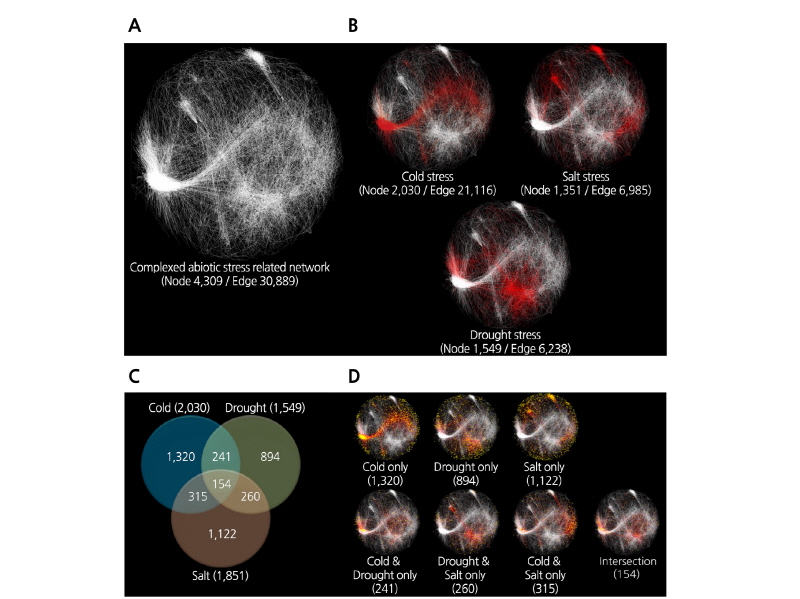
Fig. 1. Properties and topologies of gene co-expression networks related to integrated abiotic stress tolerance in Chinese cabbage. Analysis of experimental data in the co-expression network indicates that the average correlation coefficient is >0.85. The network is displayed in a Cytoscape spring-embedded layout. Nodes represent genes, and edges represent significant expressional interactions between the genes (A), distribution of genes associated with cold, salt, and drought stress (B), Venn diagrams showing the distribution of stimulus-specific and shared stress responses (C), and distribution of genes (D). Colored dots and lines indicate significantly differentially expressed genes during each stress respons
Cold Stress-specific Responses
We identified 1,320 cold stress-specific genes by Venn diagram analysis, representing 65% of the total cold stress-related co-expression network. After functional classification of specific gene expression-related cold-specific response clusters, we selected the four clusters with the highest enrichment scores, which represent genes involved in ribosomal RNA processing, ribosomal protein, calcium-binding, and calmodulin like protein (Table 1). We also used the gene ontology-based network to analyze major changes in the function and expression of genes involved in cold stress-specific responses (Fig. 2A and Table S2).
The gene ontology clusters contained many genes related to the ‘Response to temperature stimulus’, ‘Ribonucleoprotein complex biogenesis and assembly’, ‘Organ development’, and ‘Phenylpropanoid biosynthetic process’. In particular, genes in the ‘Response to temperature stimulus’ group were found to activate ‘vegetative to reproductive phase transition of meristem’ mechanisms, which act to convert the plant from vegetative to reproductive growth. The organic acid metabolic process genes in the ‘phenylpropanoid biosynthetic process’ group mediate the transition of resistance mechanisms against hypoxia and the succinate metabolic process to provide cold-specific resistance responses when exogenous cold stress is encountered. These results match those from informatics-based analyses of the proteins and transcriptomes expressed when cold stress is applied to A. thaliana (Leyva et al., 1995), Lotus japonicas (Calzadilla et al., 2016), and rice (Christie et al., 1994). These studies describe the accumulation of anthocyanin via phenylpropanoid metabolism as the one essential process used by plants to attain cold resistance. In B. rapa, FLAVONOID 3'-MONOOXYGENASE and FLAVANONE 3-HYDROXYLASE were overexpressed by up to 2.7-16.2-fold relative to the control after 24-48 h of cold treatment, suggesting they play a major role in the acquisition of cold resistance.
Salt Stress-specific Responses
Based on Venn diagram comparisons, we found that the salt stress-specific response network involves 1,851 genes, representing 61% of all salt stress-related genes in the co-expression network. We performed functional classification of specific gene expression-related salt-specific response clusters, finding that the three clusters with the highest enrichment scores represent gene groups encoding heat shock protein, endoplasmic reticulum (ER)-luminal binding protein, and beta-galactosidase, which play a role in stress responses, Cd2+-induced programmed cell death via endoplasmic reticulum stress, and catalyzing the hydrolysis of β-galactosides into monosaccharides, respectively (Table 1). Heat shock proteins play essential roles in protein folding, assembly, translocation, and degradation as defense mechanisms against abiotic stress in plants (Wang et al., 2004). Specifically, the salt stress-related small heat shock protein (sHsp) family, consisting of low-molecular-mass Hsps of 12-40 kDa, not only assist in protein folding in the ER (thereby preventing aggregation and stabilizing non-native proteins), but they are also expressed during certain developmental stages and provide molecular adaptation to abiotic stress conditions in plants (Vierling, 1991; Waters et al., 1996; Ehrnsperger et al., 1997; Wang et al., 2004). Therefore, the heat shock protein genes that function in salt stress-specific responses represent the initial sHsp-mediated mechanism of salt resistance acquisition in response to salt stress.
The gene ontology clusters contain numerous genes related to the ‘regulation of biological quality’, ‘Translation’, ‘Response to stimulus’, ‘Response to external stimulus’, and ‘Transport’. These clusters also contain many genes in the categories ‘Related to the cellular process’ and ‘Response to stimulus’ (Fig. 2B and Table S3). During salt-specific responses, activated jasmonate produces an ROS signal, and the activation of the abscisic acid (ABA) pathway and cell wall biosynthesis provide the plant with salt resistance. This finding indicates that the same salt stress response mechanisms are active in the plants over time.
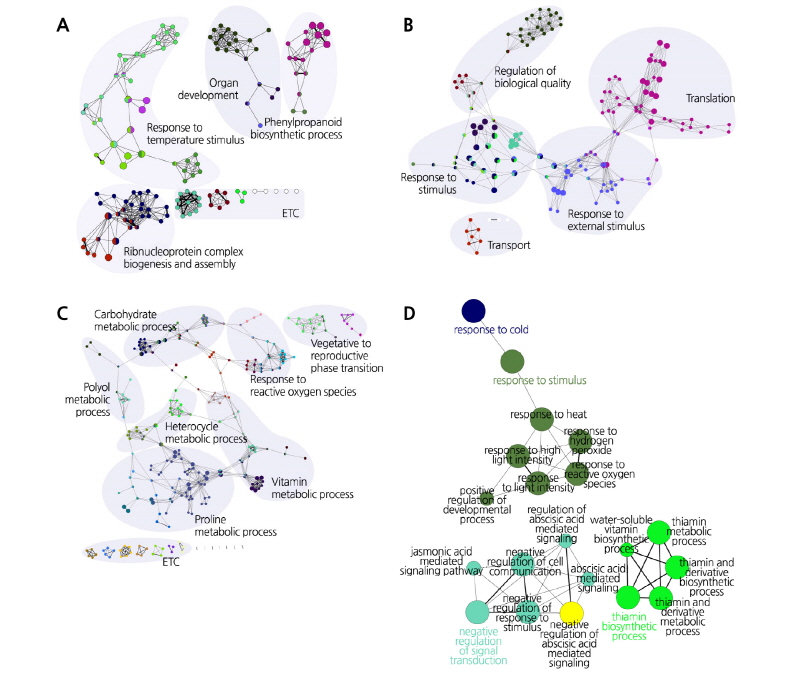
Fig. 2. Topological analysis of gene ontology categories with abiotic stress-specific or shared gene co-expression networks. Distribution of gene ontological clustering is associated with cold specific (A), salt specific (B), drought specific (C), and shared responses (D). The networks were displayed using a Cytoscape ClueGO plug-in layout.
Drought Stress-specific Responses
Venn diagram analysis revealed that the drought stress-specific response network contains 1,549 genes, representing 57% of all drought stress-related genes in the co-expression network. We performed functional classification of co-expressed genes in drought-specific response clusters, finding that the three clusters with the highest enrichment scores represent gene groups encoding ABA pathway-related protein, transcription factor, and arabinogalactan protein, which regulate plant growth and development (Table 1). These gene expression interactions promote cell wall adjustments by inducing the expression of arabinogalactan proteins. These results are similar to those obtained by Clauw et al. (2015) and Zang et al. (2015), who reported that mild drought stress suppresses the activity of cell wall hydrolase in the cell wall, which loosens cell walls to maintain them in a growth-ready state, while in dehydration tissues, cell walls stiffen to provide the plants with drought resistance. During this drought stress-specific response, the genes ARABINOGALACTAN PROTEIN 10 (AGP10) and ARABINOGALACTAN PROTEIN 20 (AGP20) were expressed at levels of up to 2.8-7.0-fold relative to the control after 24-48 h drought treatment, suggesting they play a major role in the acquisition of drought resistance.
The gene ontology clusters contain numerous genes related to the categories ‘Carbohydrate metabolic process’, ‘Response to reactive oxygen species’, ‘Vegetative to reproductive phase transition’, ‘Polyol metabolic process’, ‘Heterocycle metabolic process’, ‘Proline metabolic process’, and ‘Vitamin metabolic process’. These clusters also contain numerous genes in the categories ‘Carbohydrate metabolic process’ and ‘organic acid metabolic process’, which show similar expression patterns in response to various mechanisms over time in the drought stress network model (Fig. 2C and Table S4). Continuous drought stress triggers the production of ROS in plants, followed by the activation of RFOs and vitamin metabolism via carbohydrate metabolic processes. Moreover, the ‘Polyol metabolic process’ group was not identified during our time-based analysis. Polyol, a product of photosynthesis, not only functions as a compatible solute, but it also functions in the anti-oxidant response and plays a role in molecular adaptation to drought stress conditions in plants (Williamson et al., 2002).
Shared Responses to Cold, Salt, and Drought Stress
A comparison of Venn diagrams showed that 154 genes are common to the cold, salt, and drought stress-related co-expression networks, representing only 7.6%, 8.3%, and 9.9% of all cold, salt, and drought stress-related co-expressed genes, respectively. We performed functional classification of shared stress-related response clusters, finding that the three clusters with the highest enrichment scores represent gene groups involved in the response to abiotic stress, metal ion binding, and transcription factors (Table 1).
The gene ontology clusters contain many genes related to the ‘Negative regulation of signal transduction’, ‘Thiamin biosynthetic process’, and ‘Response to temperature stimulus’. The clusters also contain numerous genes in the category ‘Temperature stimulus’ that showed similar expression patterns in response mechanisms over time in each stress network model (Fig. 2D and Table S5). In the shared abiotic stress response, the core mechanism of abiotic stress tolerance in B. rapa is the inactivation of ABA metabolism to trigger the activation of proline biosynthesis. When confronted with abiotic stresses such as cold, salt, and drought, plants close their stomata to prevent moisture loss. This mechanism can be divided into two types: slow modulation of stomata size by ABA, and rapid modulation in response to environmental changes independent of ABA. In terms of genes related to stomata regulation and ABA inactivation, in the present study, ABA INSENSITIVE 1 (ABI1) and HOMEOBOX PROTEIN 6 (HB6) showed up to 4-fold increases in expression relative to the control after each treatment (Fig. 3A and 3B). HB6 has a negative impact on plant growth or inactivation of ABA under abiotic stress, i.e., it provides resistance by closing the stomata (Lechner et al., 2011). ABI1 plays a role in the activation of the DELTA1-PYRROLINE-5- CARBOXYLATE SYNTHASE (P5CS1) gene; triggering ABI1 expression under abiotic stress conditions induces the production of hydrogen peroxide (H2O2) via an ABA-mediated catalyzing reaction (Strizhov et al., 1997; Szabados and Savouré, 2010). This hydrogen peroxide signaling via ABI1 leads to the activation of proline biosynthesis (Fig. 3C). Proline is essential for primary metabolism, regulating cell homeostasis and osmotic balance in the cytoplasm and protecting the plant from excess ROS accumulation through its scavenging activity as a singlet oxygen quencher (Hare and Cress, 1997; Matysik et al., 2002; Szabados and Savouré, 2010). In addition, in the ‘Thiamin biosynthetic process’ group, representative genes that function in the response to abiotic stimulus included THIAMIN BIOSYNTHESIS PROTEIN (ThiaminC), which was upregulated up to 2-fold in response to stress treatment (Fig. 3D). Tunc-Ozdemir et al. (2009) demonstrated that thiamin and thiamin pyrophosphate play an additional role in protecting cells against oxidative damage caused by abiotic stress. In summary, under abiotic stresses such as cold, salt, and drought stress, the accumulation of ABA is inactivated by HB6, and hydrogen peroxide signaling by ABI1 catalyzed ABA translocation leads to the activation of proline biosynthesis by P5CS1 and thiamin biosynthesis by ThiaminC. These results suggest that these genes play a major role in the acquisition of abiotic stress resistance (Fig. 3E).
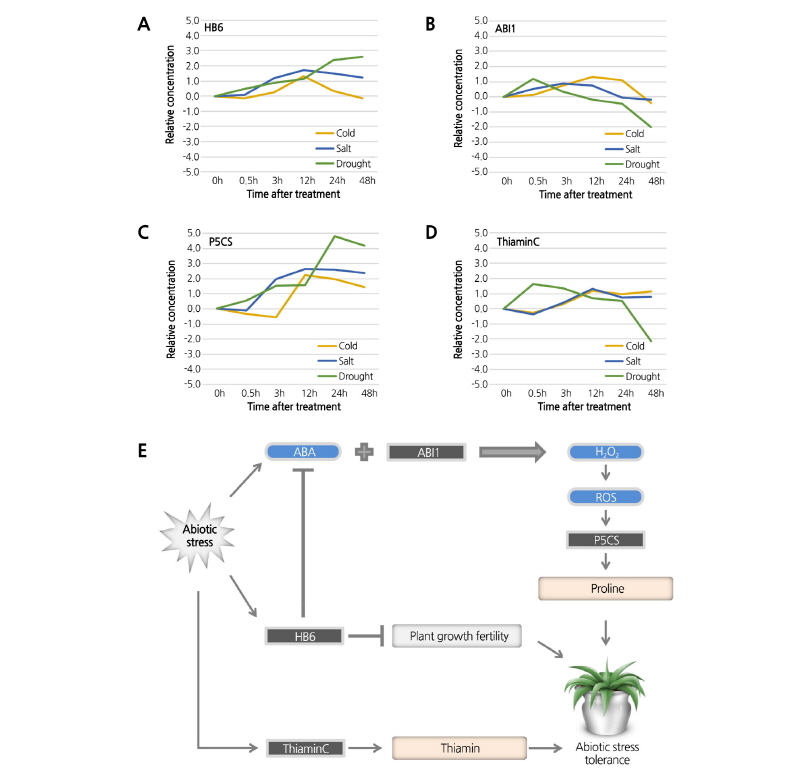
Fig. 3. Hypothetical abiotic stress tolerance model in Chinese cabbage. Time-based expression levels of major abiotic stress-responsive genes HB6 (A), ABI1 (B), P5CS (C), and ThiaminC (D) after cold, salt, or drought treatment in the KBGP-24K oligo chip data. (E) Model of abiotic stress tolerance in Chinese cabbage.
Characterization of New Abiotic Stress Tolerance-related Genes using the Network Model
We constructed an expression network model for cold, salt, and drought stress response genes to compare cold, salt, and drought stress tolerance in transgenic B. rapa lines overexpressing B. rapa genes Bra000358, Bra033354, and Bra026701, respectively, whose functions are currently unknown (Yu et al., 2014a; Yu et al., 2014b; Yu et al., 2016). We selected cold, salt, and drought stress-related genes that showed an expression change of >2-fold in response to each stress treatment. Of the 3,589 genes examined, only 37, 143, and 67 genes were significantly differentially expressed under cold, salt, and drought stress conditions, respectively. A partial explanation for this difference is that we used Gene-Chip data compared to the constructed abiotic stress-related expression network models (Fig. 4).
To characterize the abiotic stress-related genes selected from the networks, we performed functional annotation clustering using the ClueGO plug-in database. Of the cold stress-resistance genes analyzed, we selected two gene network groups with shared interactions (Fig. 4A). Group 1 contained genes related to succinate biosynthesis, thiamin biosynthesis, and glutamate biosynthesis. The representative succinate biosynthesis genes that responded to cold stimulus included the cold resistance-related gene GAMMA-AMINOBUTYRATE TRANSAMINASE (GABA-T), which not only participates in succinate biosynthesis during the tricarboxylic acid (TCA) cycle, but is also related to cold resistance (Krasensky and Jonak, 2012). Group 2 contained genes that were annotated as unknown or predicted to be directly involved in cold stress-related processes. Moreover, the ETC group contained genes related to the ETHYLENE-RESPONSIVE TRANSCRIPTION FACTOR (ERF) gene, which also functions in cold stress, as described previously.
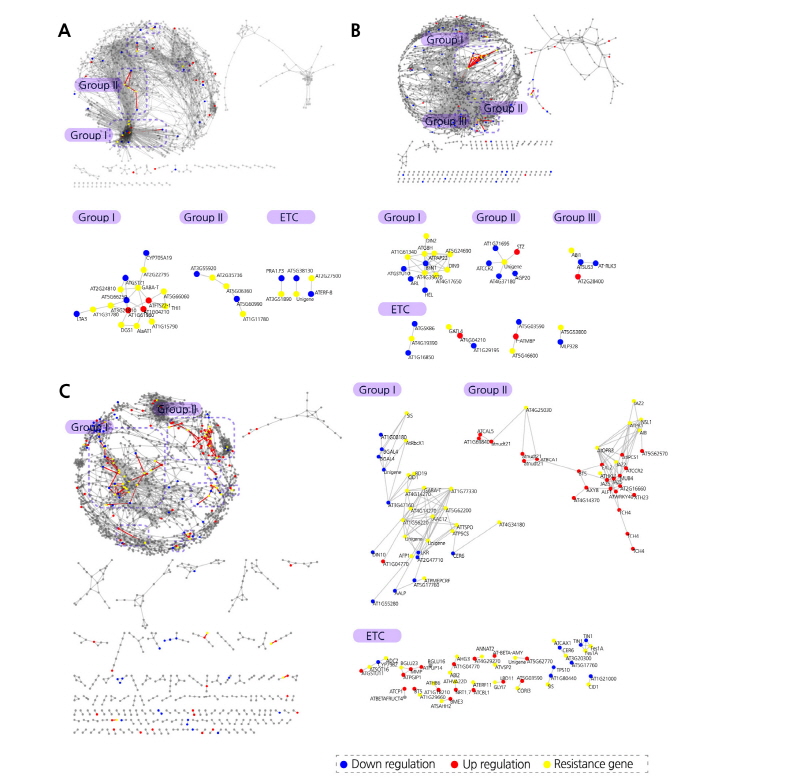
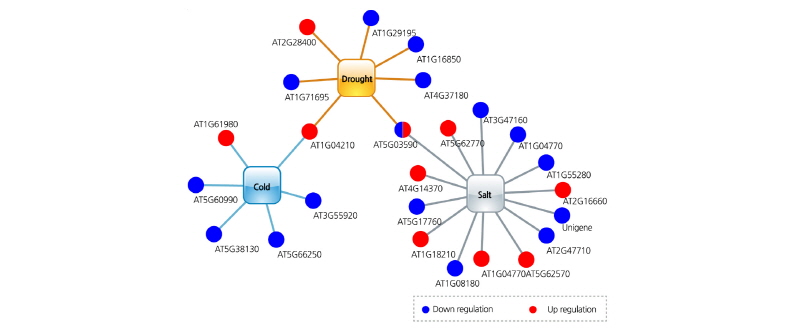
Fig. 4. Examination of Brassica rapa abiotic stress-related network topology using abiotic stress-tolerant transgenic Chinese cabbage. Distribution of downregulated (blue), upregulated (red), and stress-resistance genes (yellow) in the cold (A), drought (B), and salt (C) stress-related gene networks.
Of the drought stress-resistance genes analyzed, we selected three gene network groups with shared interactions (Fig. 4B). Group 1 contained genes possibly related to drought stress, including the representative genes AUTOPHAGY 8H (ATG8H) and DARK INDUCIBLE 2 (DIN2), which are involved in leaf senescence and autophagy in plants, respectively (Baena-González et al., 2007; Devarenne, 2011). Group 2 contained genes related to proline biosynthesis, which is a core abiotic stress response in the shared gene co-expression networks. The accumulation of proline is an essential process that helps plants acquire cold resistance via ROS detoxification; SALT TOLERANCE ZINC FINGER (STZ) in particularly important in this process (Mehterov et al., 2012). In transgenic B. rapa under drought stress, STZ was upregulated up to 2.7-fold relative to the control, suggesting it plays a major role in the acquisition of drought resistance.
Of the salt-stress resistance genes analyzed, two gene network groups with shared interaction were selected (Fig. 4C). Group 1 and Group 2 included down- and upregulated gene networks, respectively. Group 1 contained genes also related to abiotic stress, including GABA-T, a succinate metabolism-related gene in plants. These genes displayed similar expression patterns to genes involved in cold stress resistance in transgenic B. rapa. In particular, the unknown B. rapa gene BRAS0001S00011581 (AT1G04770), which appears to participate in signal transduction during photosynthesis (Ruckle et al., 2012), was overexpressed 6.28-fold in plants under salt stress. Continuous salt stress is a major factor in the downregulation of photosynthesis in plants, which triggers leaf senescence. Therefore, when salt stress-resistant transgenic B. rapa plants were exposed to salt, changes in the expression of the unknown B. rapa gene BRAS0001S00011581 had an unknown modulatory effect on signal transduction during photosynthesis, which led to the acquisition of salt-stress resistance by activating the TCA cycle via GABA-T. The additional representative downregulated genes, LYSINE-KETOGLUTARATE (LKR) and 3-KETOACYL-COA SYNTHASE 6 (CER6), function in the inactivation of the TCA cycle and the activation of transpiration via regulating cuticular wax formation in plants, respectively (Schulz et al., 2007; Angelovici et al., 2010). Hence, our results confirm that negative effects in metabolism affecting plant growth, such as the inactivation of photosynthesis and the TCA cycle, occur in B. rapa during salt stress, which leads to the inactivation of genes involved in these mechanisms and signal transduction related to salt resistance.
In conclusion, our analysis using an abiotic stress-related network model resulted in the characterization of unknown abiotic stress tolerance-related genes, including 6, 14, and 7 genes for cold, salt, and drought stress tolerance, respectively, in Chinese cabbage (Fig. 5 and Table 2). The network model constructed in the present study verifies that changes in gene expression occur in B. rapa in response to abiotic stress, increasing our understanding of abiotic stress resistance mechanisms and abiotic sensitivity in B. rapa. We discovered numerous abiotic stress resistance genes, which could be utilized in the development of genetically modified crops.
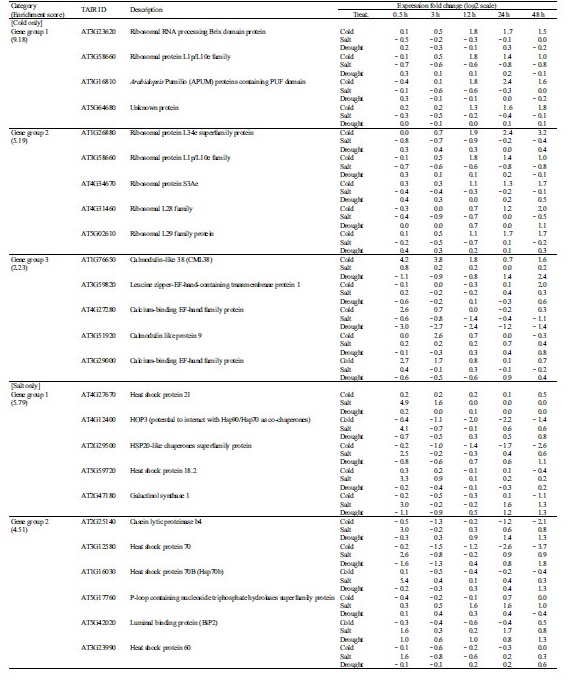
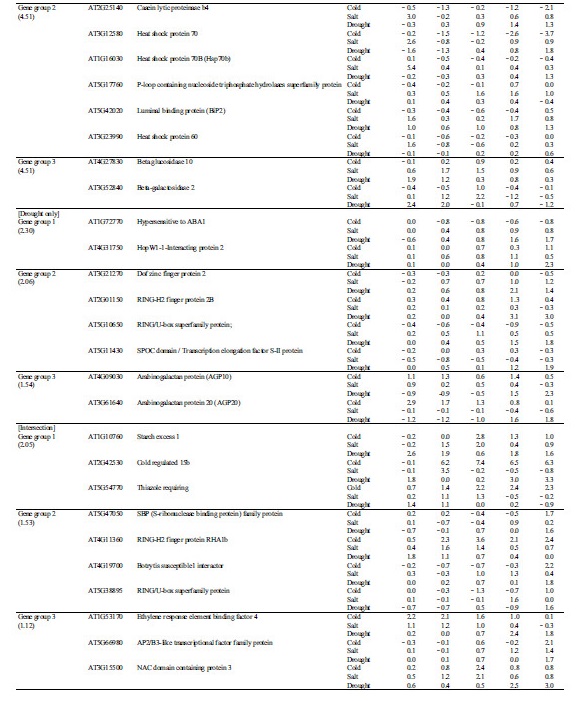
Table 2. Characterization of abiotic stress tolerance-related unknown genes in Chinese cabbage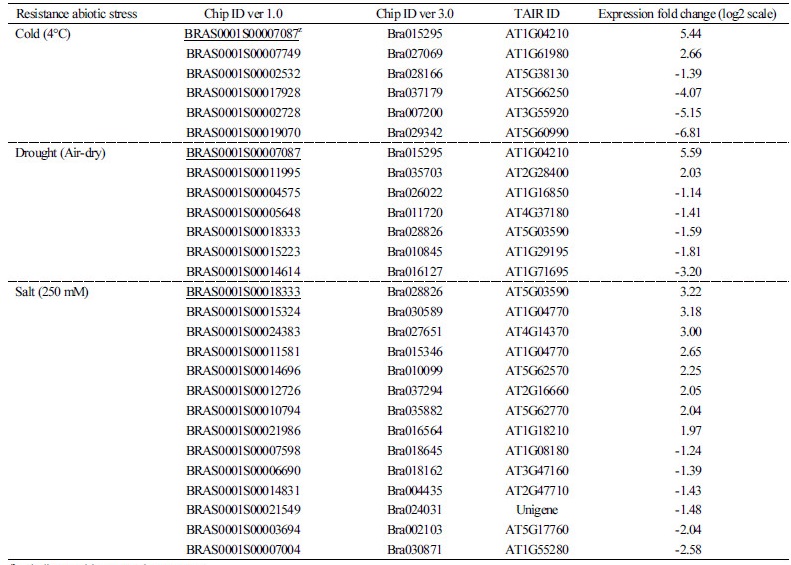 | |
zunderline: multi-stress resistance gene. | |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |
Table S2. Major groups of gene ontology categories for genes in the cold stress-specific gene co-expression networks
Gene ontological |