Introduction
Materials and Methods
Plant Materials
RNA Extraction and Gene Cloning
Construction of Vectors
Preparation of Agrobacterium Cultures for Infiltration
Expression Analysis by qRT-PCR
Analysis of PA Content
Statistical Analysis
Results and Discussion
Transient ihpRNA-induced DkPDS Gene Silencing via Agroinfiltration
Transient Silencing of DkLAR Reduces PA Levels in Persimmon Leaves
Introduction
Persimmon (Diospyros kaki Thunb.) belongs to the genus Diospyros in the family Ebenaceae and is recognized as the most important species among fruit crops of this genus. Persimmon is generally classified into two major types, PCNA and non-PCNA, based on the fruit astringency at maturity and genetic characteristic. PCNA-type persimmons include Chinese PCNA (C-PCNA) and Japanese PCNA (J-PCNA), which are further subdivided into three groups, PVNA, PVA and PCA (Akagi et al., 2011). Condensed tannins (also known as proanthocyanidins, PAs) accumulate in the vacuoles of young and developing fruit. Accordingly, the astringency of persimmon fruit directly depends upon PA levels. In the past decade, our understanding of the metabolic networks of PAs in persimmon was greatly enhanced by the use of transcriptome sequencing technology (Nakagawa et al., 2008; Luo et al., 2014). However, although many candidate genes associated with PA metabolism have been identified, progress in elucidating their functions has been hampered due to the lack of an efficient system for functional characterization of this plant.
Hairpin RNA (hpRNA)-mediated RNA interference (RNAi) technology is an effective approach for gene silencing, thus providing a robust tool for studying gene function. The intron-containing, self-complementary hpRNA (ihpRNA) constructs are more effective for silencing target genes compared to sense and antisense RNAs, as revealed by the higher silencing rates in transgenic lines (Wesley et al., 2001; Smith et al., 2000). Due to its superior characteristics, including the ease of manipulation and the applicability to numerous plant species, Agrobacterium-mediated transient ihpRNA silencing has been widely employed to characterize various genes in tobacco (Yan et al., 2012), grapevine (Urso et al., 2013), strawberry (Hoffmann et al., 2006) and Dendrobium (Ratanasut et al., 2015). To the best of our knowledge, transient ihpRNA-induced gene silencing in persimmon has not previously been reported.
PAs are phenolic oligomers or polymers of flavan-3-ol units (catechin and epicatechin). Leucoanthocyanidin reductase (LAR) and anthocyanidin reductase (ANR), the key enzymes for flavan-3-ol biosynthesis, can convert leucoanthocyanidin to catechin and anthocyanidin to epicatechin, respectively (Xie et al., 2003; Tanner et al., 2003). DkANR converts anthocyanins to both 2,3-cis-flavan-3-ol and 2,3-trans-flavan-3-ol and is involved in PA biosynthesis in persimmon fruit (Akagi et al., 2009). The DkLAR, which was isolated from C-PCNA, is closely related to its homologs in other plant species. The expression pattern of DkLAR is consistent with tannin cell development in the C-PCNA genotype. DkLAR is potentially involved in PA accumulation in persimmon fruit (Wang et al., 2010). Therefore, transient silencing of DkLAR in persimmon leaves might allow us to study its function in PA biosythesis in persimmon fruit.
In this study, we established a simple and highly efficient transient gene silencing system for gene functional analysis in persimmon based on ihpRNA-induced gene silencing. To evaluate the efficiency of the transient silencing system, we chose DkPDS as a gene-silencing target. DkLAR in leaves was also the target of gene silencing induced by T-DNA constructs producing LAR ihpRNAs, which led to reduce PA levels. The system developed in this study can be utilized for rapid, high-throughput functional analysis of genes involved in PA accumulation.
Materials and Methods
Plant Materials
Three persimmon (Diospyros kaki Thunb.) cultivars,‘Eshi 1’ (C-PCNA), ‘Youhou’ (J-PCNA) and ‘Mopanshi’ (PCA), grown in the Persimmon Repository of Huazhong Agricultural University (Wuhan, China) were used in this study. Five-week-old leaves were sampled from the trees and used for analysis.
RNA Extraction and Gene Cloning,
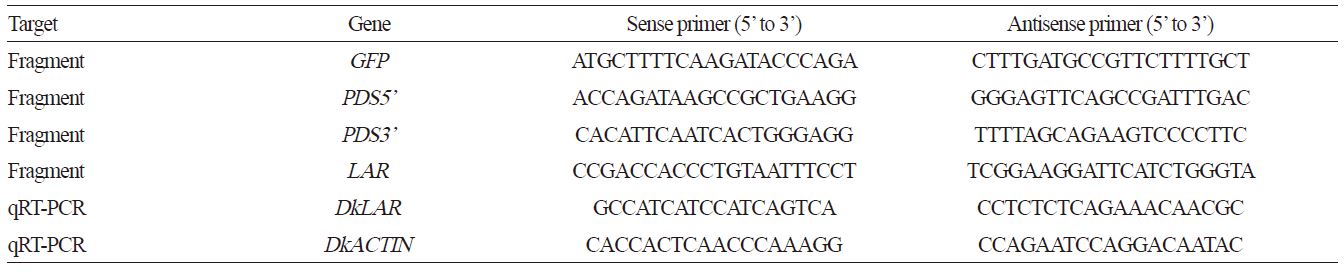
Total RNA was extracted with an RNA Prep Pure Plant Kit (Tiangen, Beijing, China) according to the manufacturer’s instructions. The cDNA was synthesized from approximately 1 μg of total RNA using a PrimeScriptTM RT Reagent Kit (TaKaRa, Japan) according to the manufacturer’s instructions. Two fragments (PDS5’ 310 bp and PDS3’ 304 bp) of DkPDS (GU112527) (Zhao et al., 2011) and a fragment (253 bp) of DkLAR (EU747876) (Wang et al., 2010) were amplified from the cDNA of young ‘Eshi 1’ fruit. The primers used in this study are shown in Table 1.
Construction of Vectors
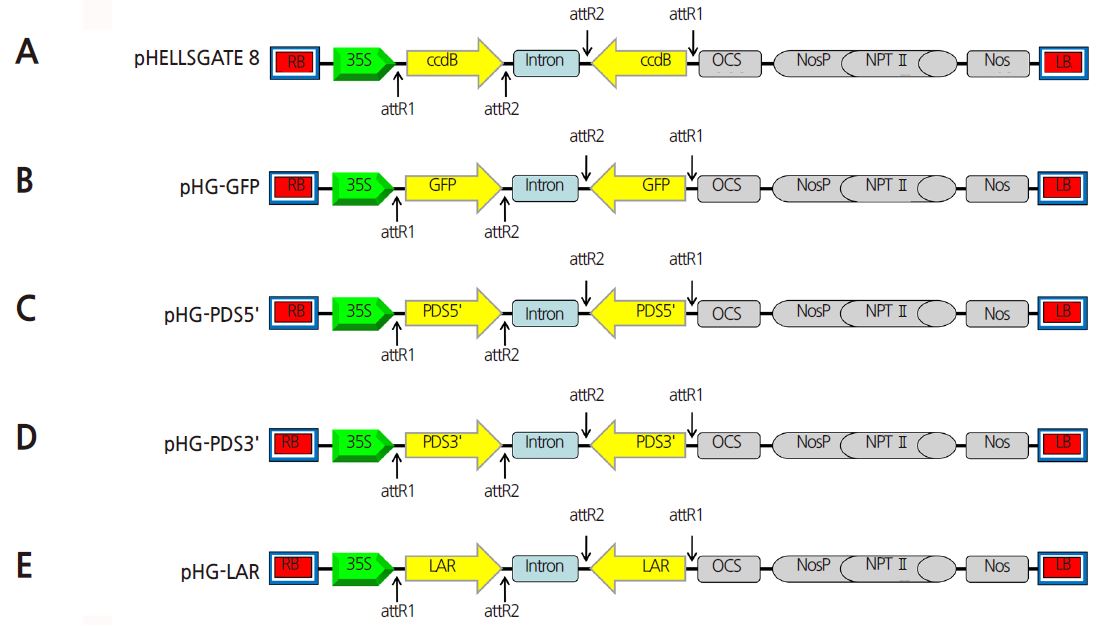
The fragments were introduced into the pDONRTM222 vector using the BP recombination reaction (Invitrogen, USA) and sequenced. After sequence verification, the products were transferred to the pHELLSGATE8 vector containing the coupled cell division B (ccdB) gene using the LR recombination reaction (Invitrogen, USA) based on Gateway® technology, generating three ihpRNA vectors: pHG-PDS5’, pHG-PDS3’ and pHG-LAR (Fig. 1). In addition, a fragment (281 bp) of GFP was amplified from pCAMBIA1302 and introduced into pHELLSGATE8 to create the pHG-GFP construct, which was used as a control (Fig. 1).
Preparation of Agrobacterium Cultures for Infiltration
The vectors were transferred into Agrobacterium strain GV3101. To establish Agrobacterium suspensions, a single bacterial colony was inoculated in 50 ml of LB medium containing the appropriate concentration of kanamycin and shaken overnight at 250 rpm at 28°C. The cultures were centrifuged at 4,000 × g for 10 min and re-suspended in infiltration medium composed of 10 mM MgCl2, 10 mM 2-(N-morpholino)ethanesulfonic acid (MES) and 150 μM acetosyringone to an optical density (OD) of 0.75 at 600 nm (Mo et al., 2015). The cultures were then incubated at room temperature for 3 h before infiltration. Agrobacterium-mediated leaf infiltration of the suspensions was carried out as described by Mo et al. (2015). The infiltrated leaves were collected at 7 or 15 days after agroinfiltration for DkPDS expression analysis of ‘Mopanshi’ transformed with pHG-PDS5’, pHG-PDS3’ and pHG-GFP, respectively. The pHG-LAR and pHG-GFP vectors were transformed into ‘Mopanshi’ and ‘Eshi 1’ leaves, respectively. The leaves were sampled at 10 and 15 days after agroinfiltration.
Expression Analysis by qRT-PCR
The primers used for qRT-PCR analysis are shown in Table 1. The qRT-PCR was performed on a LightCycler® 480 Real-Time PCR Instrument (Roche Diagnostics, Germany) with SYBR Pre-mix Ex-Taq (TaKaRa, Japan), and gene expression was normalized against the expression of the actin gene. For each reaction, the average threshold cycle (Ct) was automatically determined by the LightCycler® 480 Real-Time PCR Instrument as the default state. Quantitative RT-PCR was performed with three biological replicates per sample, and data represent means ± SD (n = 3).
Analysis of PA Content
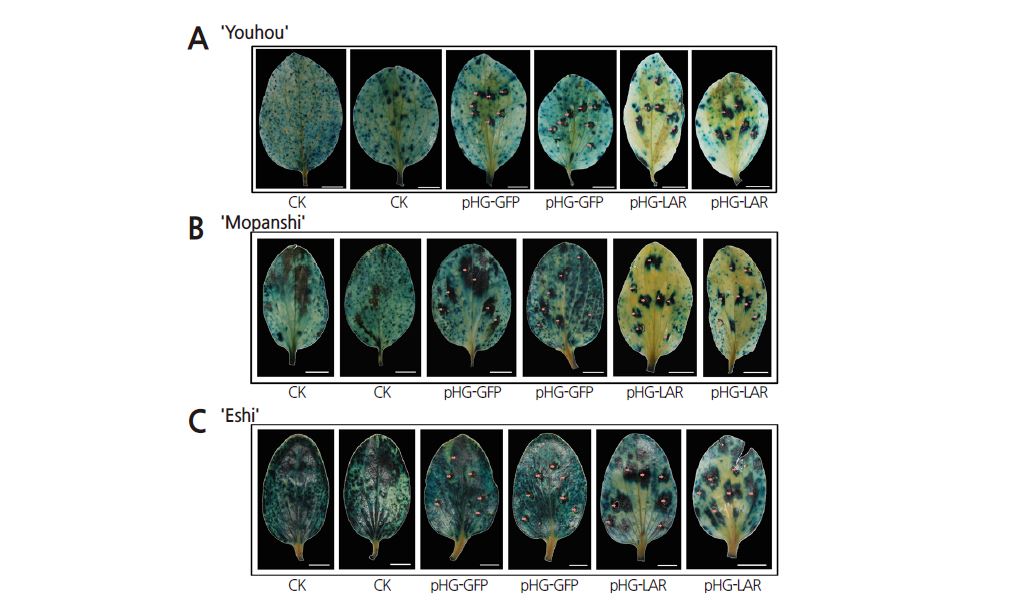
Leaves of three persimmon cultivars, ‘Mopanshi’, ‘Eshi 1’ and ‘Youhou’, transformed with pHG-LAR or pHG-GFP were sampled at 10 and 15 days after agroinfiltration. Histochemical staining with DMACA solution (0.6% DMACA and 1% 6N HCl in methanol) was performed to analyze PA contents in leaves. The infiltrated leaves were decolorized in 30% acetic acid in ethanol for 12–20 h, washed with 75% ethanol and stained blue in DMACA solution for 2 min (Li et al., 1996): the deeper the blue coloring, the higher the PA content. Furthermore, ‘Mopanshi’ and ‘Eshi 1’ leaves sampled at 15 days after infiltration were used for tannin content analysis. In addition, soluble and insoluble tannin contents were also measured using the Folin-Ciocalteau method (Oshida et al., 1996). Approximately 0.1 g of leaf tissues was used for soluble and insoluble tannin content analysis, with a total of ten single leaf replicates.
Statistical Analysis
Statistical significance of differences was calculated using Student’s test (p < 0.01). All statistical analysis was performed with SPSS 17.0.
Results and Discussion
Transient ihpRNA-induced DkPDS Gene Silencing via Agroinfiltration
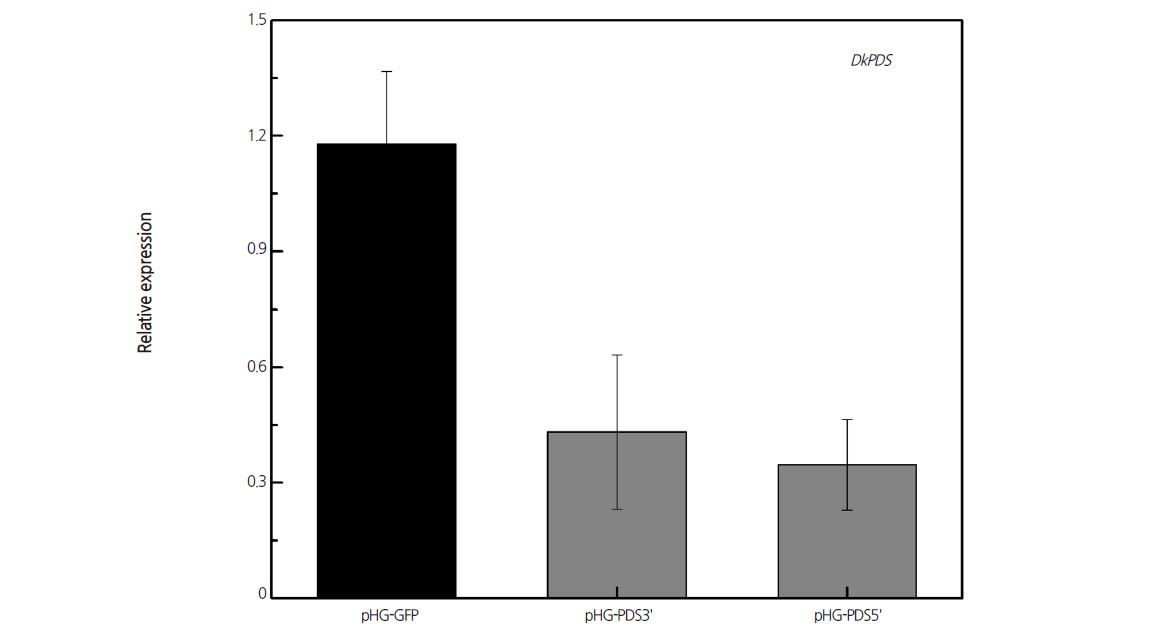
In order to develop an ihpRNA-based transient gene silencing system in persimmon leaves in vivo, we chose DkPDS as the gene-silencing target. Silencing PDS (encoding a key enzyme in the carotenoid biosynthesis pathway) (Zhao et al., 2011) caused photobleaching in plants. We cloned two fragments of DkPDS into the ihpRNA interference vector pHELLSGATE8 to examine the effects of different sequence fragments on transient silencing efficiency. Therefore, the DkPDS gene was targeted by two ihpRNA constructs, pHG-PDS5’ and pHG-PDS3’, under the control of the CaMV 35S promoter. These vectors were transiently transformed into ‘Mopanshi’ (PCA) leaves by agroinfiltration. As shown by qRT-PCR analysis, DkPDS expression in leaves infiltrated with either construct was reduced three-fold compared to empty vector at 7 days after infiltration (Fig. 2). These results indicate that Agrobacterium-mediated transient transformation of leaves with ihpRNA successfully induced DkPDS gene silencing, with no difference in silencing efficiency between constructs. However, we observed extensive leaf drop in plants infiltrated with pHG-PDS5’ and pHG-PDS3’ at 10 days after agroinfiltration (data not show). This observation indicates that DkPDS was successfully silenced, which resulted in dysfunctional leaf photosynthesis and led to leaf abscission. Therefore, we were unable to observe a photobleaching phenotype in these plants.
Transient Silencing of DkLAR Reduces PA Levels in Persimmon Leaves
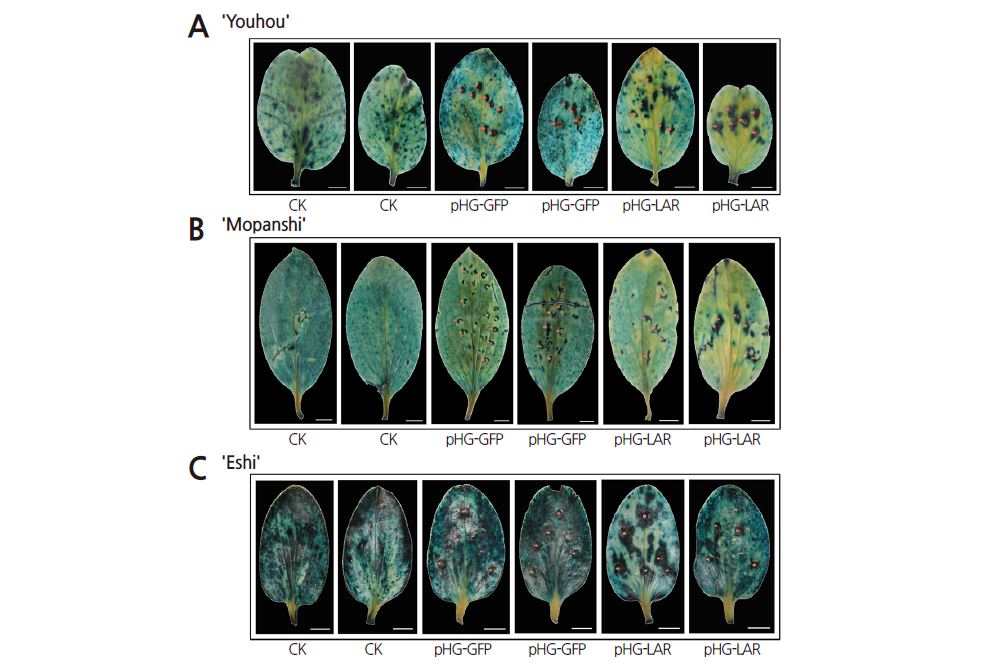
As described above, silencing of DkPDS caused leaf abscission. Therefore, we chose DkLAR, which is potentially involved in the accumulation of persimmon PAs (Wang et al., 2010), for transient silencing to evaluate the stability and reliability of the transient ihpRNA-mediated gene silencing system in persimmon. The ihpRNA interference vector pHG-LAR, which is targeted to the leucoanthocyanidin reductase (LAR) gene, was transformed into ‘Mopanshi’ (PCA), ‘Youhou’ (J-PCNA) and ‘Eshi 1’ (C-PCNA) leaves in vivo by agroinfiltration. DMACA staining revealed significantly reduced PA levels in leaves infiltrated with pHG-LAR compared to the control (Figs. 3 and 4). These results suggest that the transient silencing of DkLAR inhibits PA accumulation in persimmon leaves. Moreover, lower PA levels were detected in infiltrated leaves sampled at 15 days versus 10 days after agroinfiltration (Figs. 3 and 4). Therefore, our Agrobacterium-mediated transient ihpRNA-inducing gene silencing system is stable and should be appropriate for persimmon cultivars with different types of astringency.
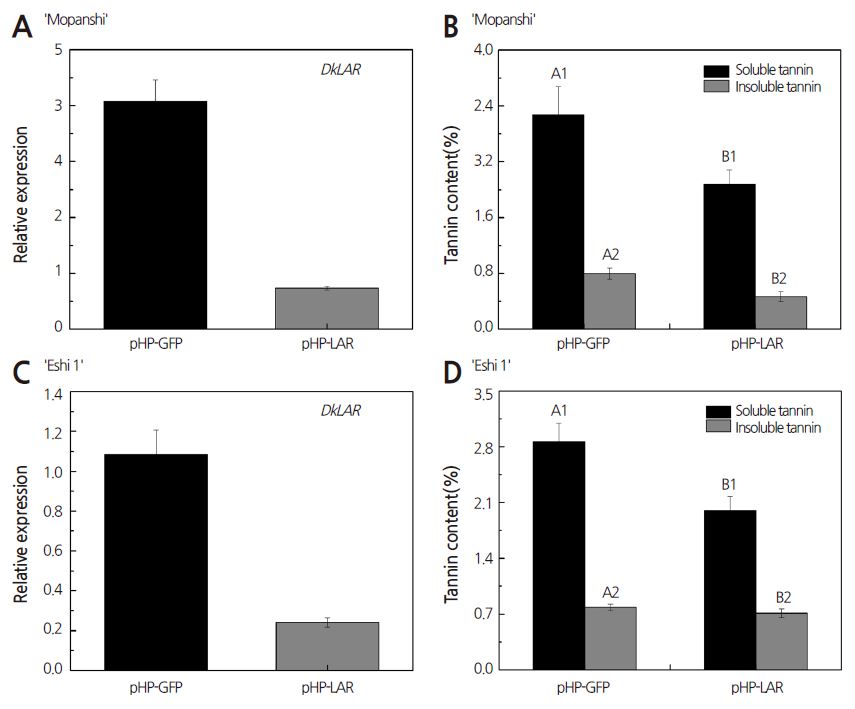
We also collected ‘Mopanshi’ and ‘Eshi 1’ leaves transformed with pHG-LAR for further experimental analysis at 15 days after agroinfiltration to confirm the notion that DkLAR is involved in PA accumulation (Wang et al., 2010). The qRT-PCR analysis showed that the expression of DkLAR was ca. 6-fold and ca. 5-fold lower in ‘Mopanshi’ and ‘Eshi 1’ leaves than in the positive control transformed with empty vector (pHG-GFP), respectively (Figs. 5A and 5C). In addition, the Folin-Ciocalteau assay revealed extremely significant reductions in the levels of soluble tannin in leaves of both persimmon cultivars infiltrated with pHG-LAR (Figs. 5B and 5D). The same tendencies were observed for insoluble tannin contents in ‘Mopanshi’ persimmon leaves, which were significantly reduced compared to the empty vector control (Fig. 5B). However, a slight reduction in insoluble tannin levels was observed in ‘Eshi 1’ persimmon leaves (Fig. 5D). These results indicate that DkLAR gene silencing leads to reduce PA levels in persimmon leaves, which is consistent with the results of DMACA staining (Fig. 4). LAR activity is clearly associated with PA accumulation (Joseph et al., 1998; Pang et al., 2007). To date, LAR has been isolated from many plant species, including grapevine (Bogs et al., 2005; Gagne et al., 2009) and poplar (Yuan et al., 2012; Wang et al., 2013). Previous studies have confirmed that VvLAR, which shares 71% sequence identity with DkLAR (Wang et al., 2010), contributes to PA accumulation in grape (Bogs et al., 2005). The overexpression of PtrLAR1 (Wang et al., 2013) and PtrLAR3 (Yuan et al., 2012) also causes strong accumulation of PAs in poplar. Together, these findings suggest that DkLAR, an LAR homolog, is required for PA accumulation in persimmon.
Functional studies of genes involved in PA biosynthesis in Arabidopsis are often based on the lose-of-function tt (transparent testa) mutants (Debeaujon et al., 2000; Devic et al., 1999), which can be easily identified using DMACA staining (Abrahams et al., 2002). However, it is not currently possible to create mutants in perennial woody plants such as persimmon for functional analysis of PA biosynthesis genes. Persimmon leaves can accumulate a large number of PAs. Thus, combined with DMACA staining, the in vivo transient gene silencing system for persimmon leaves developed in this study can serve as a model method for other perennial woody plants for rapid functional analysis of genes involved in PA biosynthesis.
Moreover, the current results show that the efficiency of transient silencing differs depending on genotype (Figs. 3-5). In general, syringe-mediated agroinfiltration requires forcing the bacterial suspension through stomata and into air spaces in the leaf (Sparkes et al., 2006). Leaf architecture traits including high density of palisade and spongy mesophyll cells, low density and/or small aperture of stomatal pores and overall fragility of leaf tissue often lead to poor infiltration (Simmons et al., 2009, 2012). Thus, we speculate that the difference in transient silencing efficiency in different persimmon genotypes is due to differences in leaf architecture. Similar results have been reported in other plant species (Wroblewski et al., 2005; Manavella and Chan, 2009; Andrieu et al., 2012).
In conclusion, we developed a simple, stable and efficient transient ihpRNA-induced gene silencing system based on Agrobacterium infiltration in persimmon leaves in vivo that can be utilized for rapid and high-throughput functional analyses of persimmon genes. Silencing of DkLAR using this system inhibited PA accumulation in persimmon.