Introduction
The Cucurbitaceae family consists of economically impor-tant species such as Cucumis melo (melon), Citrullus lanatus (watermelon), Cucurbita spp. (pumpkin), and Cucumis sativus (cucumber) (Jeffrey, 2005), and breeding programs involving these plants have generated multi-billion dollar food, cosmetic, and medicine industries (Waminal et al., 2011; Weng and Sun, 2012). Genome assembly studies, particularly of cucum-ber, melon, and watermelon, have revealed phylogenetic relationships among members of this family and provided opportunities and resources to expedite crop improvement (Garcia-Mas et al., 2012; Guo et al., 2013; Huang et al., 2009).
Introgression of important agronomic traits, such as biotic and abiotic stress resistance, from wild related species is an important tool for crop improvement (Anamthawat- Jónsson, 2001; Zamir, 2001). This approach has been demonstrated in wheat, rice, rapeseed, and other crops (Anamthawat-Jónsson, 2001; Brar and Khush, 1997; Humphreys et al., 1997; Jiang et al., 1993; Lashermes et al., 2000; Snowdon et al., 1997). Some cucurbits have also benefitted from this breeding approach. For example, recent intro-gression breeding from a wild relative of cucumber, Cucumis hystrix, has increased the genetic diversity of cucumber (Delannay et al., 2010; Zhuang et al., 2006). Additionally, introgression between cultivars and wild relatives of bitter gourds (Momordica charantia) (Liao et al., 2012) and Citrullus (Sain et al., 2002) have also been reported.
Other wild cucurbits possess economically and medicinally important traits. For example, Melothria japonica is a good source of proteolytic enzymes (Antão and Malcata, 2005, Uchikoba et al., 2001), and Trichosanthes kirilowii, which has long been used in China as a medicinal plant, is the source of the anti-HIV drug trichosanthin and the less cytotoxic protein TAP 29 (Lee-Huang et al., 1991; Mayer et al., 1992; McGrath et al., 1989; Qian et al., 2012). Sicyos angulatus, a wild invasive species introduced to Korea in the mid-20th century (Tzonev, 2005), is used as a rootstock for grafting cucumbers and watermelons to enhance resistance to nematodes and improve yield (Lee, 1994; Shengping et al., 2006). Like S. angulatus, Citrullus lanatus var. citroides also provides nematode- and fungus-resistant rootstock for watermelon (Thies and Levi, 2007). Efficient breeding programs that produce crops with established desirable traits will maximize the exploitation of these agriculturally and economically significant traits.
Cytogenetic information plays an important role in breeding programs for some of the world’s most important crops, such as wheat, maize (Albert et al., 2010; Benavente et al., 2008; Icso et al., 2014) and watermelon (Sain et al., 2002). Considering the potential of the four wild cucurbits (i.e., Citrullus lanatus (Thunb.) Mansf. var. citroides L. H. Bailey, Melothria japonica Maxim., Sicyos angulatus L., and Trichosanthes kirilowii Maxim.) as genetic resources, their limited cytogenetic information prompted us to investigate their chromosome characteristics to provide basic information of their genome structure. Most cucurbit species have small chromosomes that are difficult to distinguish (Waminal and Kim, 2012); however, fluorescence in situ hybridization (FISH) has revealed the identity of chromosome pairs and provided information that can be used in breeding programs (Huang et al., 2009; Waminal et al., 2011; Waminal and Kim, 2012).
Here, we determined the ploidy level, chromosome con-stitution, and rDNA distribution patterns of four wild cucurbit species using FISH analysis. Additionally, we report the hexaploid and decaploid cytotypes of T. kirilowii for the first time. The role of polyploidy in crop improvement is discussed.
Materials and Methods
Plants growing in uncultivated lands were collected from different areas in South Korea. Mature fruits of Melothria japonica Maxim. and Trichosanthes kirilowii Maxim. were collected from South Jeolla Province, while those of Sicyos angulatus L. were from Sky Park, Seoul. The seeds of Citrullus lanatus (Thunb.) Mansf. var. citroides L. H. Bailey were kindly provided by Professor Gung Pyo Lee, Plant Genomics & Breeding Laboratory, School of Bioresource and Bioscience in Chung-Ang University, Ansung, Korea. Seeds were germinated and root tips with actively dividing cells were harvested.
Chromosome spread preparations and FISH procedures were performed according to Waminal and Kim (2012). Briefly, root tips treated with 2 mM 8-hyroxyquinoline and fixed with Carnoy’s solution were enzymatically digested with pectolytic enzyme solution (2% Cellulase R-10 [C224, Phytotechnology Laboratories] and 1% Pectolyase Y-23 [P8004.0001, Duchefa] in 100 mM citrate buffer) for 1 h. Root tips were then squashed on slides pre-cleaned with 70% ethanol. Air-dried slides were fixed in 2% formaldehyde (Vrana et al., 2012) for 5 min and dehydrated with a series of ethanol treatments (70, 90, and 100%). A 5S rDNA probe was obtained by PCR amplification of Brassica oleracea 5S rDNA using primers from Waminal et al. (2014), while a 45S rDNA probe was obtained from a clone from Gerlach and Bedbrook (1979). DNA probes were labeled with either Alexa Fluor 488-5-dUTP (C11397, Life Technologies) or Texas Red-5-dUTP (NEL417001EA, Perkin Elmer). Images were captured with an Olympus BX53 epifluorescence micro-scope equipped with a Leica DFC365 FS CCD camera, and processed using Cytovision version 7.2 (Leica Microsystems, Germany). Further image enhancements were performed using Adobe Photoshop CS6.
Results and Discussion
FISH karyotype analysis is an excellent method for pro-viding information about a genome’s chromosomal organization. It has contributed to genomic studies and breeding programs by providing chromosomal information about a species of interest (Macas et al., 2007; Mayer et al., 2011; Szinay et al., 2010). In addition, chromosome composition can be used to verify and explain phylogenetic relationships of closely related taxa (Guerra, 2008; Mandáková et al., 2015,). For instance, although the usual basic chro-mosome number of the family Cucurbitaceae is either 11 or 12 (Robinson and Decker-Walters, 1997), cucumber (Cucumis sativus) shows an atypical number of x = 7, where five of the seven chromosomes have resulted from a fusion of ten ancestral chromosomes after diverging from melons (Huang et al., 2009; Koo et al., 2010). Chromosome infor-ma-tion was also critical in identifying the taxonomic relationship of C. hystrix, a wild Cucumis species with morphological and biochemical features similar to those of cucumber, but with a basic chromosome number of 12, as observed for melon (Zhuang et al., 2006). Additionally, chromosome number information describes the association of certain phenotypic characteristics to the ploidy level (Qian et al., 2012; Singh, 1979; Soltis et al.; 2012, Xiong and Pires, 2011).
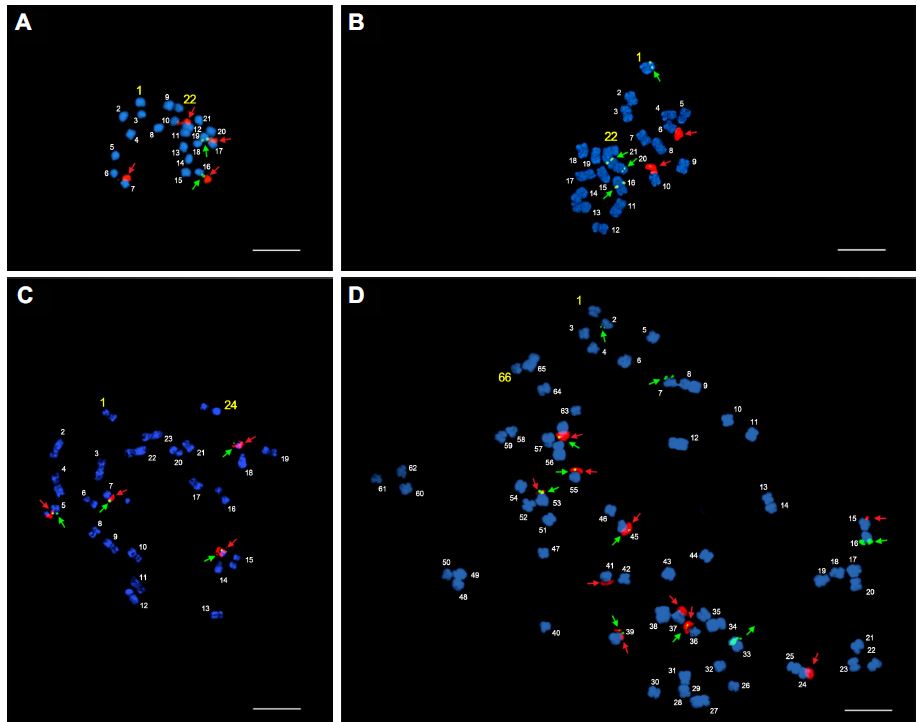
In this study, we analyzed four wild cucurbit species, i.e., Citrullus lanatus (Thunb.) Mansf. var. citroides L. H. Bailey, Melothria japonica Maxim., Sicyos angulatus L., and Trichosanthes kirilowii Maxim., using FISH karyotype analysis with 5S and 45S rDNA probes. All four species showed a typical basic chromosome number of 11 or 12. C. lanatus var. citroides and M. japonica, which belong to the tribe Benincaseae, revealed diploid complements of 2n = 22, and S. angulatus from the tribe Sicyeae showed 2n = 24 (Fig. 1A-C). However, T. kirilowii, which is from the tribe Trichosantheae, appeared as polyploids of hexa-, octa-, and decaploid complements of 2n = 66, 88, and 110, respectively, with a basic number of 11 (Figs. 1D, 2 and 3). Cucurbitaceae chromosomes are relatively small (Waminal and Kim, 2012; Waminal et al., 2011). The chromosome length of C. lanatus var. citroides ranged from 3.1 to 4.7 μm, while those of M. japonica, S. angulatus, and T. kirilowii ranged from 2.4 to 3.4 μm, 1.9 to 4.6 μm, and 2.3 to 4.5 μm, respectively (Table 1). Chromosome numbers were assigned based on decreasing order of total chromosome lengths. Our previous results of chromosome composition in Cucurbitaceae species showed variation from the usual basic chromosome number of 11 and 12; for instance, Luffa cylindrica has 13 (2n = 26) and Cucurbita moschata has 10 (2n = 40), while others, such as Lagenaria siceraria and Momordica charantia, have the usual basic chromosome number of 11 (2n = 22) and Benincasa hispida has 12 (2n = 24). This suggests a dynamic genomic history laden with genome downsizing or amplifi-cation at the chromosomal level within Cucurbitaceae, and this is particularly exemplified by the genomic relationship between cumber and melon (Huang et al., 2009, Koo et al., 2010). Genomic downsizing or amplification is not a rare phenomenon among angiosperms. In fact, it is commonly associated with genomic hybridizations that result in poly-ploids and their corresponding subsequent diploidization events, which can be demonstrated by rDNA loci reduction (Bruggmann et al., 2006; Clarkson et al., 2005; Kovarik et al., 2012; Mun et al., 2009).
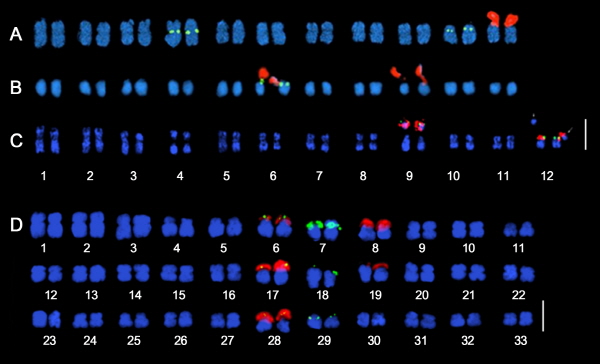
The locations and distributions of ribosomal DNA can be useful for deducing species history and phylogenetic relationship (Clarkson et al., 2005). In several cucurbit species studied previously, 5S and 45S rDNA co-localization were commonly observed (Waminal et al., 2011; Waminal and Kim, 2012). Similarly, we observed co-localization of rDNA signals in three out of four wild species (Fig. 3, Table 1). M. japonica revealed one and two 5S and 45 rDNA loci, respectively, with one co-localized signal (1:2:1); while the corresponding numbers for S. angulatus, T. kirilowii, and C. lanatus var. citroides were 2:2:2, 5:5:3, and 2:1:0, respectively (Table 1). These co-localized signals could indicate syntenic regions from a shared origin (Bertioli et al., 2009; Kellogg 2013), and thus may carry some phylogenetic implications in the study of Cucurbitaceae species. Additionally, comparative genomics at the sequence level, similar to those done in grasses (Mayer et al., 2011), should provide insight into their phylogenetic relationships. Hexaploid T. kirilowii showed a disproportional rDNA loci number in relation to its ploidy. This can be explained by a loss of 45S rDNA loci after polyploidization, considering that 45S rDNA loci have been shown to be rapidly gained or lost even after a single generation (Wendel, 2000; Waminal et al., 2012).
A hexaploid species has been reported in Trichosanthes bracteata (Karmakar et al., 2013) and diploid and octaploid cytotypes have been reported in T. kirilowii species (Qian et al., 2012). However, the hexaploid and decaploid cytotypes in T. kirilowii found here were not previously reported. Qian et al. (2012) reported diverse morphological polymorphism in diploids and octaploids of T. kirilowii collected from different regions of China. While leaf morphologies varied between the diploids and octaploids, there was even more variation among the octaploids. The fruit color, however, showed distinct difference between the two ploidy groups (Qian et al., 2012). Phenotypic variations need to be investi-gated in hexaploid and decaploid T. kirilowii cytotypes in a continued study. Most angiosperms are considered to have undergone ancient polyploidization events (McGrath and Lynch, 2012). While many cucurbits examined in previous studies exhibit disomic chromosome pairing, some have traces of ancient polyploidy in their chromosome number, such as the genus Cucurbita, which has 2n = 40, showing a multiple basic chromosome number of 10 (Weeden and Robinson, 1986), and some exhibit an ongoing species diversification through polyploidization, as observed in Gynostemma pentaphyllum (Jiang, 2009).
Geographical distribution plays an important role in facilitating hybridization between related species, or cyto-types, and creating natural polyploids (Balao et al., 2011; Wendel, 2000). Several mechanisms could have resulted in the present polyploid genomes. Based on chromosome number, hexaploid T. kirilowii may be a result of hybridi-zation between a diploid and a tetraploid. Furthermore, the decaploid could have resulted from a later hybridization between an octaploid and diploid, or tetraploid and hexaploid. Additionally, the disjunct distribution of this genus (Kocyan et al., 2007) may have facilitated several hybridization events, and those found in Korea may have established hexaploidy and decaploidy in addition to the other established ploidies. The resulting genome reorganization after polyploidization (Bennetzen, 2005; Bennetzen et al., 2005; Chen and Yu, 2013) is a mechanism for plants to stabilize genomes and often leads to adaptive mechanisms to a vast array of environmental conditions. Hence, most polyploids have more efficient survival machinery as compared to their diploid counterparts (Otto, 2007; Ramsey, 2011; Wang et al., 2012) and this may explain their adaptation to a wide range of environmental conditions (Comai, 2005).
|
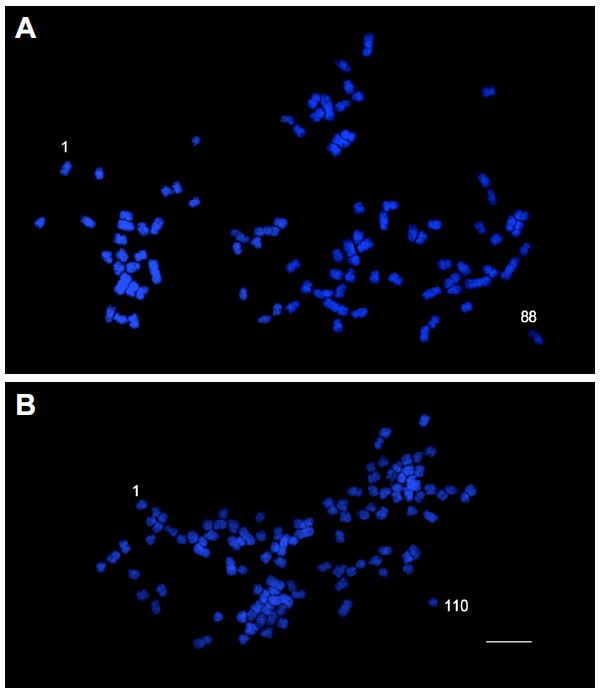
Fig. 2. Polyploid cytotypes of Trichosanthes kirilowii. The octaploid (A) and decaploid (B) are shown. Bars = 5 μm. |
T. kirilowii has been used as a medicinal herb in oriental countries and is known to have important medicinal pro-perties (Lee-Huang et al., 1991; Mayer et al., 1992; Qian et al., 2012). Breeding and cultivation of elite polyploid lines may be useful for the mass production of these medicinals. Presently, much work remains to be done to identify and develop the best cytogenetic lines with optimal medicinal and agronomic traits. The cytogenetic results presented here complement molecular and biochemical studies on T. kirilowii (Minh et al., 2015; Ni et al. 2015; Seo et al., 2015). Similarly, basic chromosome information of C. lanatus var. citroides, M. japonica, and S. angulatus may be useful in efforts to improve Cucurbitaceae crops. Understanding the genomic history of these species may further facilitate the efficient exploitation of their desirable agricultural and medicinal traits. As desirable traits are partially influenced by ploidy (Balao et al., 2011; Comai, 2005), the medicinal properties of these hexaploid and decaploid cytotypes should be further studied (Minh et al., 2015; Ni et al., 2015; Seo et al., 2015).