Introduction
Materials and Methods
Plant and Pathogen Materials
CR Evaluation
Microscopic Observation of Infected Root Tissues
Results and Discussion
CR Evaluation in Chinese Cabbage Cultivars
Microscopic Comparison of Infected Root Tissues of Clubroot-Resistant and -Susceptible Lines
Inheritance of CR in the F1, F2, and BC1F1 Populations
Introduction
Clubroot disease in Brassica crops, which is caused by Plasmodiophora Brassicae, was first reported in Korea in 1928 and occurred in a mild form until the 1980s (Kim et al., 2003). Since the 1990s, this disease has become an important limiting factor for Chinese cabbage production in Korea (Kim et al., 1999).
Clubroot resistance (CR) in B. rapa has been found only among European turnip cultivars such as ‘Debra’, ‘Gelria R’, ‘Milan White’, and ‘Siloga’, in which at least eight CR genes were identified (Crute et al., 1980; Yoshikawa, 1993; Hirai, 2006; Diederichsen et al., 2009; Piao et al., 2009). Most of the known resistance of B. rapa to the clubroot pathogen is under dominant monogenic control (Strandberg and Williams, 1967; Kuginuki et al., 1997; Suwabe et al., 2003; Hirai et al., 2004; Piao et al., 2004; Cho et al., 2008). However, Yoshikawa (1981) reported that CR in European turnips is controlled by a major gene and several minor genes. Two independent major genes and one quantitative trait locus for CR have also been identified in ‘Siloga’ (Suwabe et al., 2003, 2006). To date, CR in B. rapa has been shown to be independently controlled by several genes (Piao et al., 2009).
Many CR cultivars of Chinese cabbage have been bred by introducing CR genes from European turnips (Yoshikawa, 1981; Hirai, 2006). However, most of these CR cultivars have become susceptible to clubroot in some regions (Tanaka et al., 1998; Kuginuki et al., 1999; Hirai et al., 2004; Piao et al., 2004), while the European turnips remain highly resistant to field isolates of P. Brassicae from the same area (Kuginuki et al., 1999). This discrepancy might be due to the evolution of pathotypes or the loss of CR genes during the breeding of Chinese cabbage CR cultivars (Kuginuki et al., 1999; Hirai et al., 2004). To breed durable CR lines, other CR genes for introgression into Chinese cabbage cultivars must be identified from European turnips. The inheritance patterns of CR genes must also be elucidated prior to Chinese cabbage breeding programs.
Interactions between P. Brassicae and its host have been studied using various host genotypes (Crute et al., 1980; Voorrips, 1995), such as Williams’ differential cultivar set (1966) and the European clubroot differential set (Buczacki et al., 1975). Multiple pathotypes of P. Brassicae can co-exist in the same infected fields (Jones et al., 1982; Kim et al., 2003; Xue et al., 2008) or even a single root gall (Jones et al., 1982). The race specificity of CR genes is difficult to define using genetic analyses of resistance to field isolates due to the heterozygosity of the pathogen races themselves (Manzanares-Dauleux et al., 2000). Single-spore isolates (SSI), i.e., homozygous pathogen isolates, are required for precise genetic analysis and breeding programs for CR. Of all known sixteen physiological races of P. Brassicae, fourteen races except races 10 and 12 are known to be present in the Chinese cabbage cultivation areas in Korea, and races 4, 5, and 8 of P. Brassicae are widely distributed (Cho et al., 2003; Kim et al., 2003; Jang et al., 2007). Among these, race 4 has the highest pathogenicity for all four members of Williams’ differential cultivar set (Kang, 2005).
In the present study, we evaluated the resistance of Chinese cabbage (B. rapa ssp. pekinensis.) cultivars to race 4 of P. Brassicae by comparing them to the clubroot-susceptible Chinese cabbage inbred line ‘BP079’ and the clubroot-resistant European turnip (B. rapa ssp. rapifera) inbred line ‘IT033820’. We compared infected root tissues of ‘BP079’ and ‘IT033820’ by both light microscopy and scanning electron microscopy (SEM). We investigated the inheritance of CR to P. Brassicae race 4 in ‘IT033820’ to examine whether this line could be used as a new genetic resource for breeding Chinese cabbage cultivars with durable CR, because ‘IT033820’ was also found to be resistant to races 2 and 9 of P. Brassicae (data not shown).
Materials and Methods
Plant and Pathogen Materials
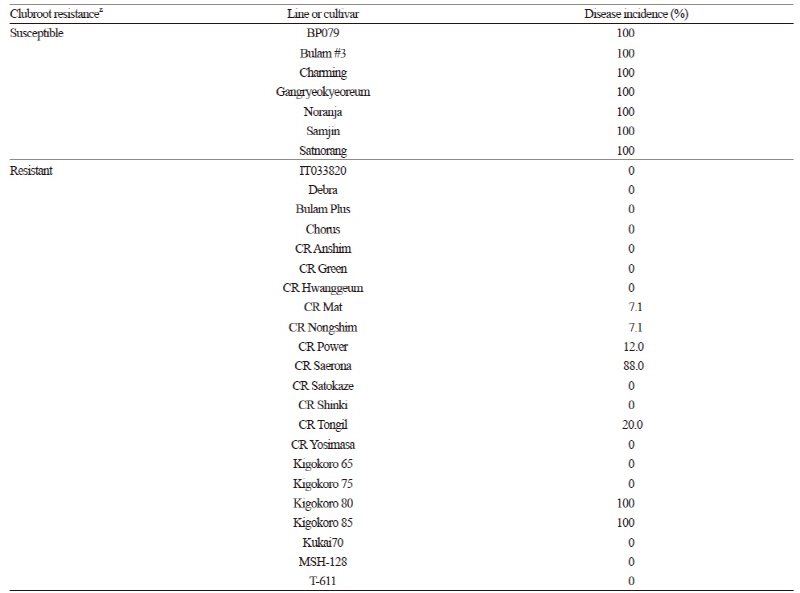
Along with European turnip line ‘IT033820’ and cultivar ‘Debra’, one Chinese cabbage line, ‘BP079’, and 26 commercially available F1 cultivars were evaluated for CR (Table 1). Seeds of ‘BP079’ and ‘IT033820’ were obtained from the National Agrobiodiversity Center of the Rural Development Administration, Suwon, Korea. ‘BP079’ is highly susceptible to diverse races of P. Brassicae, whereas ‘IT033820’ is highly resistant (Kang, 2005). Using the two lines as pollen and seed parents, respectively, ‘IT033820’ and ‘BP079’ were crossed to produce F1 hybrids, and F2 plants were obtained by self-bud pollination of an F1 plant. The F1 hybrids were backcrossed with each parental line to produce BC1F1 populations. An SSI of P. Brassicae (SSI79-2-1) identified as race 4 based on Williams’ classification (Williams, 1966) was used for CR evaluation.
CR Evaluation
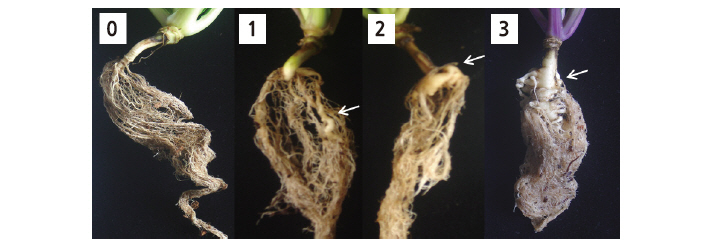
Inoculum preparation of P. Brassicae and evaluation of CR were performed using the root dip inoculation method (Johnston, 1968) with some modifications. Seeds were sown in a Petri dish (9.5 cm × 4.5 cm × 4.5 cm, W × L × H) containing moistened filter paper. When the seedlings in the dish were 1 week old, they were inoculated with a spore suspension of race 4 of P. Brassicae at 1 × 106 spores/mL by soaking their roots for 20 min. The inoculated seedlings were transplanted into pots containing commercial horticulture nursery medium (Plant World, Nongwoo Bio, Yeoju, Korea). To ensure infection, after soaking the roots the remaining suspension was distributed around the inoculated seedlings so that the spore concentration in the soil was high. Pots containing the inoculated seedlings were saturated with water for the first week to maintain an optimum environment for infection. The seedlings were grown in a greenhouse maintained at a minimum temperature of 25°C under natural daylight conditions. Four weeks after inoculation, clubroot disease symptoms in the seedlings were evaluated using the grading system described by Suwabe et al. (2003): disease index (DI) 0, no symptoms; 1, a few small, separate globular clubs on lateral roots; 2, intermediate symptoms; 3, severe clubs on main roots (Fig. 1).
Microscopic Observation of Infected Root Tissues
The infected main root tissues of clubroot-susceptible ‘BP079’ and clubroot-resistant ‘IT033820’ and ‘Kigokoro 75’ were observed under a light microscope as described by Luft (1973). The root samples (approximately 1 mm3) were fixed with 2.5% glutaraldehyde in 0.1 M sodium phosphate buffer (pH 7.2) at 4°C for 2 h. The specimens were rinsed, post-fixed with 1% osmium tetroxide at 4°C for 2 h, and held overnight in the same sodium phosphate buffer. After fixation, the specimens were dehydrated in a graded series of ethanol (40, 60, 80, 90, 95, and 100% in distilled water [v/v]). To ensure complete dehydration, the specimens were processed through three changes of propylene oxide for 15, 15, and 30 min per change and gradually infiltrated for 3 h each with 30, 50, and 100% Epon embedding medium in propylene oxide. The specimens were incubated overnight in 100% Epon before polymerization at 60°C for 72 h. The specimens were sectioned (1.5 μm), stained with periodic acid-Schiff staining, and viewed under a light microscope (Axioskop 2; Carl Zeiss Inc., Oberkochen, Germany).
For SEM examination, root tissues (main root, lateral root, and boundary tissues between main and lateral roots) were cut in median sections (2.0 mm × 2.0 mm × 2.0 mm), adhered onto stubs using carbon tape, and sputter-coated with gold using an ion coater (K-450 Sputter Coater; Emitech Ltd., Ashford, UK) to approximately 15 nm in thickness. The specimens were examined and photographed in natural mode with a Hitachi S-2460 (Hitachi Co., Tokyo, Japan).
Results and Discussion
CR Evaluation in Chinese Cabbage Cultivars
Upon infection with race 4 of P. Brassicae, no symptoms of clubroot disease were found in the European turnip cultivar ‘Debra’, but many of the Chinese cabbage cultivars exhibited disease symptoms to varying degrees (Table 1 and Fig. 2). The Chinese cabbage cultivars that were reported to be clubroot-susceptible were susceptible to race 4 of P. Brassicae; however, seven of the 20 Chinese cabbage cultivars reported to be clubroot-resistant were suceptible to race 4 of P. Brassicae to some extent. In particular, all ‘Kigokoro 80’ and ‘Kigokoro 85’ plants were severely infected (Table 1). Similary, Jo et al. (2011) examined 25 CR Chinese cabbage cultivars bred in Korea for their resistance to isolates of P. Brassicae collected from ten different regions in Korea and found that the cultivars exhibited various responses to pathogens of the same race. This suggests that only a limited number of CR genetic resources might have been used for CR breeding of Chinese cabbage cultivars in Korea, and that the CR of these cultivars might have been lost during cultivation (Jo et al., 2011). To increase the durability of CR cultivars, different CR genes should be combined into a single line, as suggested by Piao et al. (2009). Thus, more CR genes from Brassica germplasm need to be identified.
| |
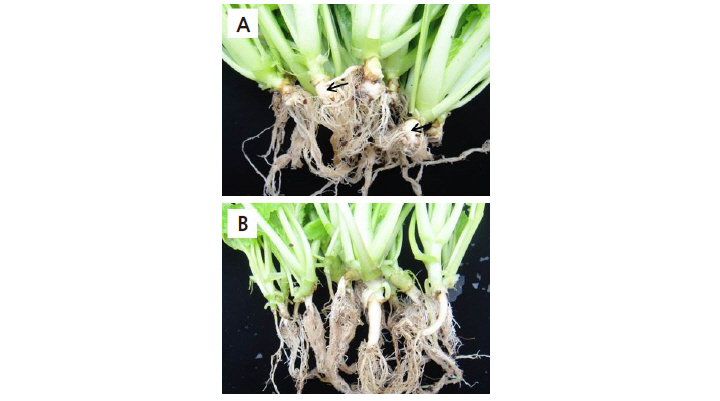
Fig. 2. Clubroot development in Brassica rapa lines ‘BP079’ (A) and ‘IT033820’ (B) inoculated with race 4 of Plasmodiophora Brassicae. Arrows indicate globular clubs. | |
Microscopic Comparison of Infected Root Tissues of Clubroot-Resistant and -Susceptible Lines
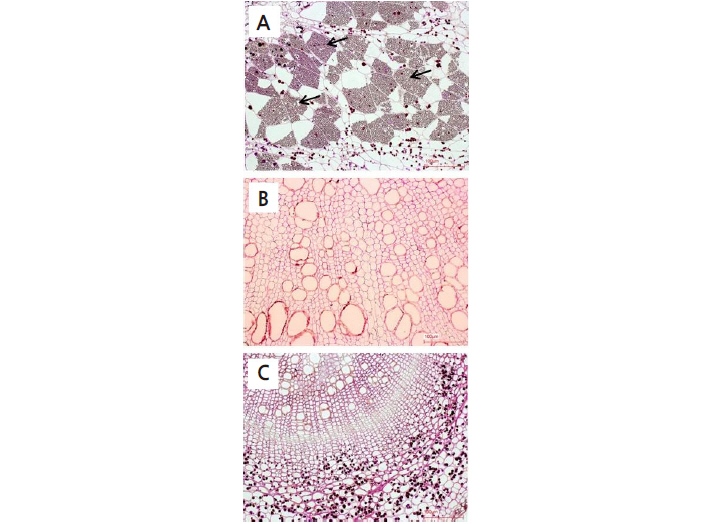
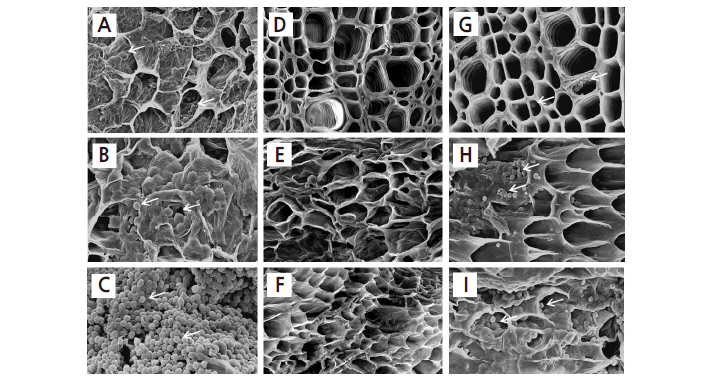
We compared the infected root tissues of clubroot-susceptible ‘BP079’ and clubroot-resistant ‘IT033820’ and ‘Kigokoro 75’ by light microscopy and SEM. Resting spores of P. Brassicae were abundant within the main root tissues of ‘BP079’ (Fig. 3A), but were not observed within root tissues of ‘IT033820’ (Fig. 3B) or ‘Kigokoro 75’ using light mincroscopy (Fig. 3C). Furthermore, many resting spores were observed under SEM in root tissues of ‘BP079’ (Fig. 4A, B, and C), but no resting spores were found in root tissues of ‘IT033820’ (Fig. 4D, E, and F). More resting spores were found in the boundary tissues between the main and lateral roots than within these roots. A few resting spores were also observed in the root tissues of ‘Kigokoro 75’ (Fig. 4G, H, and I) under SEM, although they were not detected by light microscopy. The cell walls of the main root tissues of ‘Kigokoro 75’ were intact (Fig. 4G), while those of ‘BP079’ had collapsed (Fig. 4A). Similarly, upon examination of CR in B. oleracea, Donald et al. (2008) observed no degradation of the secondary thickenings and cell walls of the xylem in the resistant host. Little or no cell wall breakage may have restricted the movement of P. Brassicae in amoeboid form, but not in the resistant host. In the present study, symptoms of root galling were not observed in ‘Kigokoro 75’, despite the presence of some resting spores. However, the precise mechanism underlying CR remains unknown.
Inheritance of CR in the F1, F2, and BC1F1 Populations
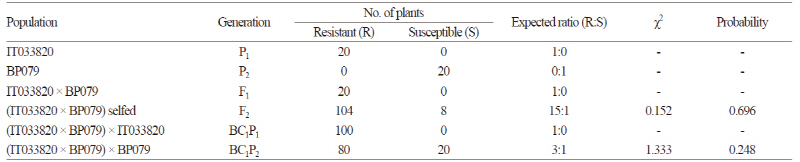
Following root dip inoculation of race 4 of P. Brassicae, the clubroot-susceptible parental line ‘BP079’ exhibited large globular clubs on its roots (DI of 3, Fig. 2A), while the clubroot-resistant parental line ‘IT033820’ and the F1 hybrids produced by crossing ‘IT033820’ and ‘BP079’ showed no disease symptoms (DI of 0, Fig. 2B). All seedlings of Chinese cabbage cultivars ‘Bulam #3’ and ‘Charming’, which were used as susceptible controls for CR evaluation, also exhibited large globular clubs on their roots (data not shown). All members of the BC1P1 population from the cross between the F1 hybrid and ‘IT033820’ exhibited a resistant phenotype (Table 2). In the BC1P2 population from the cross between the F1 hybrid and ‘BP079’, 80 out of 100 plants were resistant (R), but 20 plants were susceptible (S). The segregation ratio was 3(R):1(S) (χ2 = 1.333, p = 0.248) at a 5% significance level. Inoculated plants showed either a highly resistant or highly susceptible phenotype, indicating that CR carried by ‘IT033820’ to race 4 of P. Brassicae is dominant. By contrast, of the 112 F2 plants, 104 were highly resistant and eight were highly susceptible, with a segregation ratio of 15(R):1(S) (χ2 = 0.152, p = 0.696) at a 5% significance level, suggesting that CR in ‘IT033820’ is mainly controlled by two dominant genes, as reported by Kang (2005). Inheritance of CR to SSI of race 4 of P. Brassicae, identified based on Williams’ differential hosts, is thought to be controlled by one or two dominant genes depending on the susceptible parent (Kang, 2005). When resistant lines ‘ECD04’ or ‘IT033820’ were crossed with the susceptible line ‘ECD05’, the segregation ratio in the subsequent F2 generation was 3(R):1(S). When these lines were crossed with the susceptible parental line ‘BP079’, however, the segregation ratio in the F2 generation was 15:1. Otani et al. (1981) also reported that CR inheritance varies depending on the susceptible parents. European turnip cultivars ‘Gelria R’ and ‘Milan White’ were each crossed with Chinese cabbage and Chinese mustard (B. juncea ssp. juncea) to identify CR inheritance, revealing that CR in ‘Gelria R’ is controlled by a single, completely dominant gene. When ‘Milan White’ was crossed with Chinese mustard, CR inheritance was shown to be completely dominant; however, when it was crossed with Chinese cabbage, the resulting plants were completely or moderately resistant, depending on the susceptible parents.
Monogenic inheritance of CR genes in Chinese cabbage has often been reported (Strandberg and Williams, 1967; Hirai et al., 2004; Piao et al., 2002, 2004; Cho et al., 2008). Strandberg and Williams (1967) first reported monogenic inheritance of CR in Chinese cabbage cultivar ‘Michihili’ to races 6 and 7 of P. Brassicae. Matsumoto et al. (1998) and Piao et al. (2002, 2004) identified single dominant genes, including CRa in ‘T136-8’ derived from ‘ECD02’ for CR to race 2 and CRb in the doubled haploid line ‘CR Shinki’ derived from ‘Gelria R’ for CR to races 2, 4, and 8. Two independent major genes, Crr1 and Crr2, were also derived from ‘Siloga’ (Kuginuki et al., 1997; Suwabe et al., 2003, 2006). CR is stronger when the Crr1 and Crr2 loci are homozygous for resistant alleles than when they are heterozygous (Suwabe et al., 2003). These findings indicate that CR in B. rapa is under oligogenic control and that the cooperation of these loci is necessary to generate resistance. Another dominant gene, Crr3, was derived from ‘Milan White’ (Hirai et al., 2004). Segregation analysis of the CR gene in Chinese cabbage cultivar ‘CR Saerona’ against SSI race 4 of P. Brassicae revealed that the resistant parental line has a single, dominant gene (Cho et al., 2008).
In this study, we identified two dominant CR genes in a new CR resource, ‘IT033820’. This line can be used to develop durable CR breeding lines in Chinese cabbage, although the genetic behavior of CR is not completely understood. Further studies are needed to investigate the inheritance of CR in ‘IT033820’ more precisely and to determine the relationships between CR genes derived from ‘IT033820’ and previously reported CR genes.