Introduction
Materials and Methods
Plant Material
cDNA Cloning and Sequencing of Psy, Chy2, and Ccs
Ccs-based Drived Cleaved Amplified Polymorphic Sequence (dCAPS) Marker for Selecting ‘K146465’
Carotenoid Extraction and High Performance Liquid Chromatography (HPLC) Analysis
Results
New Nonsense Mutation in Ccs in Orange Pepper C. annuum ‘K146465’
Co-segregation Analysis between Orange Fruit Color and Ccs Mutation
Carotenoid Profiling in ‘Sweet Banana’ and ‘K146465’ by HPLC
Discussion
Introduction
Fruit color, a trait that attracts the consumer, is associated with differential pigment levels. Carotenoids, the major pigments of fruits and flowers, are synthesized in plastids (Bartley and Scolnik, 1995). They play a crucial role in photo-protection of photosynthesis and act as a precursor of the plant hormone abscisic acid (ABA) (Tanaka et al., 2008). In humans, they are important dietary nutrients as they provide many health benefits. For instance, β-carotene, α-carotene, γ-carotene, and β-cryptoxanthin are the precursors of vitamin A (Yonekura and Nagao, 2007); lutein and zeaxanthin have a significant role in delaying and preventing cataract and age-related macular degeneration, both of which are the causes of blindness in 56% of cases worldwide (Nwachukwu et al., 2016); capsanthin acts as an antioxidant (Taylor and Ramsay, 2005). Although carotenoids are essential nutrients , they cannot be synthesized by the human body (Fiedor and Burda, 2014), and therefore their intake from fruits and vegetables is important for improving human health.
Diverse fruit colors often are due to variation in the functionality and expression of carotenoid biosynthetic genes. Pepper fruits (Capsicum spp.) are typically red, but in some cultivars their color varies such as orange, yellow and brown (Lightbourn et al., 2008). A three-locus model has been proposed to explain variation in pepper fruit colors, in which three independent genes encoded by the y, c1, and c2 loci control the ripe fruit color (Hurtado-Hernandez and Smith, 1985). The y locus is responsible for yellow fruit color and is controlled by capsanthin-capsorubin synthase (Ccs) (Lefebvre et al., 1998); the c2 locus is responsible for orange fruit color and is controlled by phytoene synthase (Psy) (Huh et al., 2001; Kim et al., 2010) the role of the c1 locus is still unknown. Different combinations of alleles in these three loci can generate additional fruit colors, such as white and lemon (Thorup et al., 2000).
Among the various colors of pepper fruits, orange fruits are known to be associated with three independent genes that are involved in carotenoid biosynthesis. The first gene is Psy, which is involved in the early steps of the carotenoid biosynthesis pathway (Bartley and Scolnik, 1995). Because Psy encodes a rate-limiting enzyme, impaired activity of Psy decreases the amount of total carotenoids (Cazzonelli and Pogson, 2010), resulting in orange fruit color of ripe peppers; a typical example is C. chinense ‘Habanero’ (Huh et al., 2001; Kim et al., 2010). The second gene is Ccs, which synthesizes the red carotenoid compounds capsanthin and capsorubin in the last step of the carotenoid biosynthesis pathway. Interestingly, impaired activity of Ccs is associated with not only yellow fruit color (Lefebvre et al., 1998), but also orange fruit color (Lang et al., 2004). The third gene is β-carotene hydroxylase2 (Chy2), which catalyzes the conversion of α- and β-carotene into lutein and zeaxanthin, respectively. A study using ethyl methane sulfonate (EMS) mutagenesis of the wild type red pepper C. annuum ‘Maor’ identified a mutant C. annuum ‘E-172-3’ with orange fruit color and high levels of β-carotene accumulation, which resulted from a mutation in Chy2 (Borovsky et al., 2013).
In the present study, we explored the genetic variation associated with fruit color and carotenoid variation in the orange pepper C. annuum ‘K146465’. This germplasm and the new causal variation discovered will facilitate marker-assisted breeding for selection of orange fruit color and improvement of carotenoid levels in pepper.
Materials and Methods
Plant Material
Two pepper accessions, C. annuum ‘Sweet Banana’ (‘Sweet Banana’) and C. annuum ‘K146465’ (‘K146465’), were obtained from the National Agro Biodiversity Center, Rural Development Administration, Korea. A mapping population was constructed from a cross between ‘Sweet Banana’ and ‘K146465’. An F2 population of 217 individuals was used to conduct co-segregation analysis of fruit color and candidate genotypes. All plant material was grown under greenhouse conditions at Kyungpook National University, Daegu, Korea. Pericarp from ripe fruits of ‘Sweet Banana’ and ‘K146465’ were collected, immediately frozen in liquid nitrogen, and ground with a chilled mortar and pestle for total RNA and carotenoid extractions.
cDNA Cloning and Sequencing of Psy, Chy2, and Ccs
Total RNA from the ripe fruit pericarp was extracted using a TRI Reagent (Molecular Research Center, Inc., Cincinnati, OH, USA), and cDNA was synthesized using a DiaStarTM RT Kit (SolGent Co., Ltd., Daejeon, Korea). Primers for Psy, Chy2, and Ccs cloning were designed and synthesized by Neoprobe (Daejeon, Korea). The Psy (CA04g04080) cDNA cloning was conducted using two sets of primers, Psy-1 and Psy-2, and primer sequences are as follows. Psy-1 forward: 5ʹ- ATGTCTGTTGCCTTGTTATGGGTT-3ʹ, reverse: 5ʹ-TCTGTATCTTGACTTCCTCAAGTC-3ʹ and Psy-2 forward: 5ʹ-GAAGACAGGCTAGAAGATGTTTTC-3ʹ, reverse: 5ʹ-TCATGTTCTTGTAGAAGGCACAAG-3ʹ. The Chy2 (CA03g25820)
primer sequences included the forward primer: 5ʹ-ATGGCTGCTGAAATTTCAATCTC-3ʹ, and reverse primer: 5ʹ-CATAATCTCTTCGAACTTTTAATTCTTC -3ʹ. The Ccs (CA06g22860) cDNA cloning was also conducted using two sets of primers, Ccs-1 and Ccs-2, and primer sequences are as follows. Ccs-1 forward: 5ʹ-ATGGAAACCCTTCTAAAGCC TTTTC-3ʹ, reverse: 5ʹ-CCAAATTCCTATCAAATGGCATTGC-3ʹ and Ccs-2 forward: 5ʹ-TGGAGGGATTCTCATTTAG GTAATG-3ʹ, reverse: 5ʹ-TCAAAGGCTCTCTATTGCTAGATTG-3ʹ.
PCR amplification was carried out using e-Taq DNA polymerase (SolGent Co., Ltd) according to the manufacturer’s instructions and under the following conditions: 95°C for 3 min, followed by 35 cycles at 95°C for 30 s, 58°C for 30 s, and 72°C for 1 min, and a final extension step at 72°C for 5 min. All cDNA amplicons were cloned using a T-blunt PCR Cloning Kit (SolGent Co., Ltd). Plasmid DNA was isolated using a HiGene™ Plasmid Mini Prep Kit (BIOFACT Co., Ltd., Daejeon, Korea), and three clones per each gene were sequenced to verify sequence fidelity. Sequencing was conducted bi-directionally (SolGent Co., Ltd), and sequence alignment was performed in Cluster-X (http://www.cluster-x.org/) and GeneDoc (Nicholas, 1997), whereas protein translation was conducted using ExPASy Translate tool (http://www.expasy.org/resources).
Ccs-based Drived Cleaved Amplified Polymorphic Sequence (dCAPS) Marker for Selecting ‘K146465’
dCAPS marker was generated based on the cDNA variation in Ccs between ̒Sweet Banana̓ and ̒K146465̓ (Neff et al., 1998); we designed one Ccs/dCAPS primer pair and the sequences are as follows. Forward primer: 5ʹ-GAAGAGAAGTGTGTGATCACTATGGGAGGACGAATT-3ʹ and reverse primer: 5ʹ-CATTCTCTAACACGTCTTCTATCC-3ʹ. The PCR product obtained was 255-bp. The underlined nucleotides in the primer sequence were mutations introduced to create an artificial restriction site. They are 32 C > G and 34 C > A. Genomic DNA was used as a template for amplification, and was extracted from young leaves of ̒Sweet Banana̓ , ̒K146465̓ , and their F2 progeny by the cetyltrimethylammonium bromide (CTAB) method (Murray and Thompson, 1980). The purity and concentration of extracted genomic DNA were monitored by a Nanodrop 2000 Spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). PCR amplification was carried out using e-Taq DNA polymerase (SolGent Co., Ltd., Daejeon, Korea) according to the manufacturer’s instructions with the following conditions: 95°C for 3 min, followed by 35 cycles at 95°C for 30 s, 58°C for 30 s, and 72°C for 1 min, with a final extension step at 72°C for 5 min. The PCR amplicons obtained by the dCAPS marker were digested with EcoRI (New England Biolabs, Ipswich, MA, USA) for 4 h and electrophoresed on a 2.5% agarose gel.
Carotenoid Extraction and High Performance Liquid Chromatography (HPLC) Analysis
Approximately 100 mg of frozen pericarp powder from five biological replicates was used for carotenoid extraction. Carotenoid extraction and quantification were performed according to the method described by Yoo et al. (2017), except for the saponification procedure.
For saponification, 300 µL of 20% KOH were added to the dried sample and incubated for 1 h at room temperature. Next, 250 µL of methyl t-butyl ether were added to the saponified sample and incubated for 15 min. Finally, 242.5 µL of MeOH were added, samples were vortexed vigorously, then filtered through a 0.45 µm Smart-Por nylon filter (Woongki Science, Seoul, Korea) and transferred to a HPLC vial (Agilent Technologies, Santa Clara, CA, USA).
After the extraction, carotenoids were analyzed by HPLC (Agilent 1260 Infinity) using a C30 YMC Carotenoid S-5 (4.6 × 250 mm) and Agilent ChemStation software (Agilent Technologies). Carotenoids werWe separated using two solvents [MeOH including 0.1% ammonium acetate 100% (A) and methyl tert-butyl ether (B)]. All chemical reagents were HPLC grade. Carotenoids were detected under five spectrums (286, 348, 434, 450, and 471 nm), and each carotenoid was identified by the absorption maxima spectrum (Britton, 1985; Gupta et al., 2015). The concentration of carotenoid compounds were calculated using β-carotene (Sigma-Aldrich, St. Louis, MO, USA) and capsanthin (Sigma-Aldrich) calibration curves. Statistically significant differences were determined by t-test.
Results
New Nonsense Mutation in Ccs in Orange Pepper C. annuum ‘K146465’
To identify genetic variation regulating orange fruit color in ̒K146465̓ (Fig. 1), the cDNAs of Psy, Chy2, and Ccs in ripe fruits of ̒K146465̓ and ̒Sweet Banana̓ were cloned and sequenced. The cDNA sequences of Psy and Chy2 in ̒K146465̓ and ̒Sweet Banana̓ were identical, whereas three single-nucleotide polymorphisms (SNPs) were found in the Ccs sequence of ̒K146465̓ when compared to the sequence of ̒Sweet Banana̓ . The SNP (G to C) at 235 bp from the start codon is a missense mutation (glycine to arginine), and the SNP (C to T) at 630 bp from the start codon is a silence mutation. A T insertion at 1026 bp from the start codon results in a premature termination codon at the 370 th position of the amino acid sequence (Fig. 2). These results suggest that ̒K146465̓ carries a new type of nonsense mutation in Ccs.
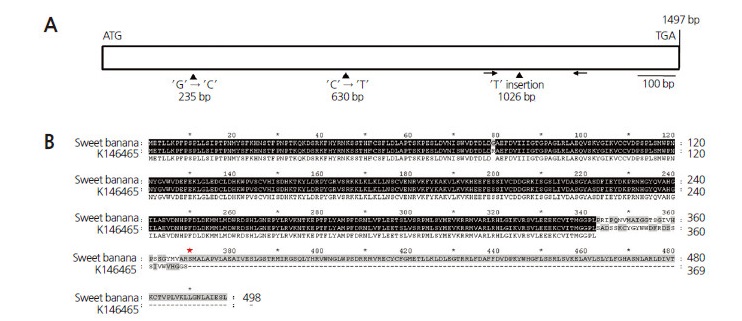
Fig. 2.
Allelic variation in Ccs in Capsicum annuum ‘K146465’. (A) Gene structure of Ccs. It consists of a 1497-bp intronless exon. ‘ATG’, start codon and ‘TGA’, stop codon. In ‘K146465’, the Ccs coding sequence showed a G (guanine) to C (cytosine) substitution at 235 bp, a C (cytosine) to T (thymine) substitution at 630 bp, and a T insertion at 1026 bp. A pair of arrows indicates the Ccs/dCAPS primer site. The bar represents 100 bp. (B) Alignment of the Ccs amino acid sequences from C. annuum ‘Sweet Banana’ and ‘K146465’. A missense mutation, glycine to arginine, and a premature termination codon are induced at positions 79 and 362 of the amino acid sequence by the G-to-C substitution and T insertion in the Ccs sequence in ‘K146465’. A red asterisk indicates the premature termination codon.
Co-segregation Analysis between Orange Fruit Color and Ccs Mutation
All fruits of F1 plants (̒Sweet Banana̓ × ̒K146465̓) were red, indicating that orange fruit color is a recessive trait. We hypothesized that orange fruit color in ̒K146465̓ is controlled by a nonsense mutation in Ccs, unlike the previous fruit color models in pepper (Lefebvre et al., 1998; Thorup et al., 2000). To verify this, we conducted a co-segregation analysis between the trait and the mutation in 217 individuals of the F2 population derived from a cross between ̒Sweet Banana̓ and ̒K146465̓ . Among the F2 progeny, 164 individuals had red fruits and 53 individuals had orange fruits, thus fitting the expected phenotypic ratio of 3:1 (Table 1).
To distinguish the nonsense mutation in Ccs, a dCAPS marker was developed based on the T1026 insertion and used to amplify the sequences in ̒Sweet Banana̓ and ̒K146465̓ . Agarose gel electrophoresis of the resulting PCR amplicons digested by EcoRI detected a polymorphism. The alleles from Sweet Banana were digested by EcoRI, whereas the alleles from ̒K146465̓ were not. The co-dominant dCAPS marker was used to identify the Ccs genotypes of 217 F2 progenies . We identified 49 homozygous individuals of ̒Sweet Banana̓ type (red fruits), 115 heterozygous individuals (red fruits), and 53 homozygous individuals of ̒K146465̓ type (orange fruits), fitting the expected genotypic ratio of 1:2:1 (Table 1). Therefore, the nonsense mutation in Ccs detected by the dCAPS marker co-segregated with orange fruit color in the 217 individuals of the F2 progeny (Fig. 3).
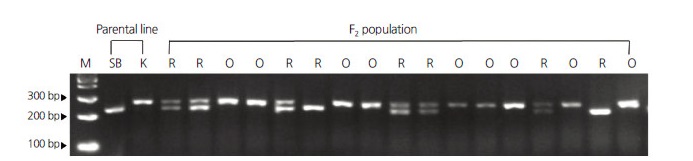
Fig. 3.
T1026 insertion identified by the dCAPS marker co-segregates with fruit color in an F2 population derived from a cross between Capsicum annuum ‘Sweet Banana’ (SB) and ‘K146465’ (K). PCR products amplified by dCAPS primers were digested with EcoRI and separated on 2.5% agarose gels. Numbers on the left side indicate the DNA ladder size; M indicates the 1 kb DNA ladder; R and O indicate red and orange fruit color in the F2 population, respectively.
Carotenoid Profiling in ‘Sweet Banana’ and ‘K146465’ by HPLC
Carotenoid profile was analyzed in ripe fruit pericarps of ̒Sweet Banana̓ and ̒K146465̓ by HPLC. As expected, capsanthin was the most abundant carotenoid in ̒Sweet Banana̓ , comprising approximately 74.6% of total carotenoids. Zeaxanthin, β-carotene, β-cryptoxanthin, phytoene, and phytofluene were identified, while lutein and violaxanthin were not detected in ̒Sweet Banana̓ . In contrast, lutein and zeaxanthin were the most abundant carotenoids in ̒K146465̓, while capsanthin was not detected.
Five types of carotenoids were detected in both pepper varieties. Zeaxanthin was detected at the similar levels [40 μg·g-1 fresh weight (FW) in ̒Sweet Banana̓ and 53 μg·g-1 FW in ̒K146465̓]. The β-carotene content in ̒K146465̓ was approximately 1.5-fold higher than in ̒Sweet Banana̓ . The levels of phytoene and phytofluene were 10-fold higher in ̒K146465̓ compared to ̒Sweet Banana̓ , whereas the amount of β-cryptoxanthin was similar in both pepper varieties (Table 2). These results suggest that the lack of capsanthin and dominance of other carotenoids are metabolic determinants of the orange fruit color in ̒K146465̓.
Discussion
Pepper fruit color is primarily determined by the concentration and composition of carotenoids (Gómez-García and Ochoa-Alejo, 2013), with capsanthin being the most abundant red carotenoid. Capsanthin and capsorubin are the end products of the carotenoid pathway and are synthesized from antheraxanthin and violaxanthin, respectively. These re actions are catalyzed by capsanthin-capsorubin synthase (Ccs) (Bouvier et al., 1994). Groups of orange (α- and β-carotene, β-cryptoxanthin, antheraxanthin, etc.) and yellow (violaxanthin, lutein, neoxanthin, etc.) carotenoids are minor compounds in pepper fruits (Gómez-García and Ochoa-Alejo, 2013). Red peppers contain high amounts of capsanthin, whereas yellow peppers contain primarily lutein and zeaxanthin and no capsanthin (Kim et al., 2016). Orange peppers accumulate diverse carotenoids, such as capsanthin, lutein, zeaxanthin, and β-carotene, depending on the cultivar (Guzman et al., 2010; Rodriguez-Uribe et al., 2012). Two major loci controlling carotenoid composition in orange peppers were reported as c2 and y, corresponding to genetic variation in Psy (Huh et al., 2001) and Ccs (Lang et al., 2004), respectively. An EMS mutant in β-carotene hydroxylase also showed orange fruit color (Borovsky et al., 2013). Formerly, it was believed that Psy is responsible for orange fruit color (Huh et al., 2001) and Ccs for yellow fruit color (Lefebvre et al., 1998). Here, we provide fundamental genetic evidence that Ccs can control orange fruit color as well. Two types of genetic variation are related to these recessive alleles: sequence variation and gene expression level variation.
A point mutation was identified in the orange C. chinense ‘Habanero’ at the splice-acceptor site of Psy; this mutation leads to a frame shift and introduces a premature termination codon (Kim et al., 2010). No expression without coding sequence variation in Psy was detected in ripening fruits of the orange C. annuum ‘NuMex Sunset’ (Guzman et al., 2010; Rodriguez-Uribe et al., 2012). The distribution of each carotenoid component in the orange pepper C. chinense ‘Habanero’ was similar to that of the red pepper. Capsanthin was the major carotenoid, and carotenoid content was 6-fold lower than in red pepper (Huh et al., 2001). Similarly, only small amounts of capsanthin were detected in C. annuum ‘NuMex Sunset’, which contains unidentified carotenoids (Rodriguez-Uribe et al., 2012).
An EMS mutant, orange C. annuum ̒E-172-3̓, carries a missense mutation (glycine 237 > arginine) in Chy2, which results in increased levels of β-carotene, a precursor of CHY2 (Borovsky et al., 2013). The orange C. annuum ‘msGTY-1’ contains a deletion in the upstream region of Ccs (Lang et al., 2004), and the orange C. annuum ‘Fogo’ has a single nucleotide deletion in Ccs, resulting in a premature termination codon at position 423 of the amino acid sequence (Guzman et al., 2010). No expression without sequence variation in Ccs was reported in C. annuum ‘Oriole’ and C. annuum ‘Orange Grande’ (Guzman et al., 2010; Rodriguez-Uribe et al., 2012).
Carotenoid content variation in these orange peppers was also identified. C. annuum ‘Fogo’ accumulates high amounts of violaxanthin, followed by lutein, zeaxanthin, and β-carotene, but no detectable amounts of capsanthin, whereas C. annuum ‘Oriole’ and ‘Orange Grande’ contain high amounts of zeaxanthin and lutein, but no capsanthin (Rodriguez-Uribe et al., 2012). Similar to C. annuum ‘Fogo’, C. annuum ‘K146465’ examined in this study has a point mutation (T insertion) at 1026 bp from the 5’ end, and the carotenoid profile was similar to that of Fogo. Both cultivars accumulate relatively high amounts of lutein, zeaxanthin, and β-carotene and no detectable amounts of capsanthin when compared to red pepper.
Ccs sequence variation regulates both orange and yellow fruit color in pepper. For example, the yellow cultivar C. annuum ‘Ramuga’ was reported to have a sequence deletion in Ccs (Lefebvre et al., 1998). In addition, two yellow Capsicum cultivars show sequence variation in the coding region of Ccs; an 8-bp insertion at position 1431 and a single nucleotide substitution at position 599, which introduces premature stop codon (Ha et al., 2007). Carotenoid profiling of these cultivars revealed high levels of lutein and the absence of capsanthin, although the total carotenoid amount was significantly lower than in red peppers. Down regulation of Ccs via virus-induced gene silencing also reduced the amount of β-cryptoxanthin, zeaxanthin, and other carotenoids, in particular capsanthin (Tian et al., 2014). The same study reported that the fruits of Ccs - silenced peppers were yellow, indicating that Ccs can regulate the yellow fruit color in pepper.
β-carotene is one of the most abundant carotenoids in ̒K146465̓ when compared to the red pepper cultivar Sweet Banana. However, it seems that higher β-carotene content may not be a direct effect of the Ccs mutation. The lycopene β-cyclase (Lcy-b ) and lycopene ε-cyclase (Lcy-e ), which are responsible for β-carotene and α-carotene biosynthesis, respectively, evolved from the same ancestor of the lycopene cyclase family. Ccs also belongs to this family (Krubasik and Sandmann, 2000). CCS showed lycopene β-cyclase activity in vivo (Hugueney et al., 1995) and Ccs showed micro-collinearity with the Beta locus in tomato (Ronen et al., 2000; Thorup et al., 2000). Beta is the overexpressor of the chromoplast-specific lycopene beta cyclase (CYC-B), which accumulates high amounts of β–carotene in tomato fruits (Ronen et al., 2000). The amino acid sequence of Cyc-b is very similar to that of Ccs (86.1% identity) (Ronen et al., 2000) and neoxanthin synthase (Nxs) (99% identity) (Hirschberg, 2001). Nxs converts violaxanthin to neoxanthin, which is known as the final product of plant xanthophylls and a precursor of the plant hormone abscisic acid (ABA) (Cutler and Krochko, 1999). In previous studies, neoxanthin was detected equally in yellow and orange Capsicum fruits, but not in red peppers (Kim et al., 2016). Despite its detection in Capsicum spp., neoxanthin synthase has not been identified yet. In agreement with Guzman et al. (2010), we hypothesize that the truncated Ccs in ̒K146465̓ could partly function as Lcy-b and Nxs, increasing β-carotene and xanthophylls, but does not function as Ccs . Additional enzymatic analysis to determine Lcy-b and Nxs activity of the truncated Ccs would be necessary to prove this hypothesis.
Because of differences in carotenoid content, orange peppers carrying the Ccs mutation may be nutritionally superior to orange pepper varieties carrying the Psy mutation. Although previous reports described Ccs mutations in orange peppers with no clear genetic evidence (Guzman et al., 2010; Rodriguez-Uribe et al., 2012), we identified a new nonsense mutation in Ccs that determines the orange fruit color by the co-segregation analysis. Furthermore, the use of dCAPS marker will facilitate marker-assisted breeding of C. annuum ‘K146465’ to select for orange fruit color and improve lutein and β-carotene levels in pepper.





