Introduction
Materials and Methods
Plant Materials
Preparation of Somatic Metaphase Chromosome Spreads
Probe Preparation
Fluorescence in Situ Hybridization (FISH) and Genomic in Situ Hybridization (GISH)
Meiotic Chromosome Pairing Analysis
Imaging and Karyotype Analysis
Results
Karyotype Analysis
Meiosis Pairing Analysis
Discussion
Introduction
Brassica and Raphanus species have been cultivated world-wide as important vegetables, fodders, and sources of functional compounds. Owing to their usefulness, many studies have been carried out to investigate breeding of these species (McNaughton, 1979; Namai et al., 1980; Olsson and Ellerstrom, 1980; Prakash et al., 2009).
Synthetic hybrid species between Brassica and Raphanus, xBrassicoraphanus or xRaphanobrassica, have been developed and studied by many researchers using morphological and cytological methods (Dolstra, 1982; Kato and Tokumasu, 1983; Tokumasu, 1976). In Korea, Lee et al. (1999, 2002) developed a new intergeneric allotetraploid, xBrassicoraphanus, by a cross between B. rapa L. ssp. pekinensis and R. sativus L. and named it ‘Baemoochae’. Initially, this new synthetic hybrid was so unstable that low seed fertility and poor uniformity were continued through generations. However, a microspore mutagenesis using N-nitroso N-methyl urethane (NMU) created stabilized progenies from this unstable material (Lee et al., 2011). They also developed another intergeneric allotetraploid xBrassicoraphanus between Korean land races of B. rapa L. ssp. pekinensis and R. sativus L. var. rafiphera and the F1 hybrid was crossed again with a mutagen-induced stabilized line. Using the same microspore mutation technique, another stable line of xBrassicoraphanus showing quite high seed fertility and morphological uni-formity was obtained, which was named BB#5.
Chromosome-based cytogenetic studies using conventional staining method have been applied to Brassica species since 1920 to investigate chromosome composition and genome structure. Chromosome characterization and genome identification have advanced significantly with the develop-ment of molecular cytogenetics through fluorescence in situ hybridization (FISH) and genomic in situ hybridization (GISH) techniques (Capdeville et al., 2008; Fukui, 2005; Hwang et al., 2010, 2012; Levsky and Singer, 2003; Park et al., 2010). Indeed, these methods have been used to investigate genome structure and inter-genomic relationship of hybrid plants (Jellen et al., 1994; Kenton et al., 1993), allopolyploid species (Cao, 2003; Devi et al., 2005; Yang et al., 1999), and recombinant breeding lines (Hwang et al., 2012; Lou et al., 2010; Vasconcelos et al., 2010).
Tandem repeat DNA sequences including 5S and 45S ribosomal DNAs (rDNAs) have primarily been used as cytogenetic markers in FISH (Kato et al., 2004; Lim et al., 2005, 2012). In Brassica, Maluszynska and Heslop-Harrison (1993) first reported the number of 45S rDNA loci in diploids (B. rapa, n = 10, AA genome; B. nigra; n = 8, BB genome; B. oleracea, n = 9, CC genome,) and allo-tetraploids (B. carinata, n = 17, BBCC genome; B. juncea, n = 18, AABB genome; B. napus, n = 19, AACC genome). Genomic dis-tributions of rDNA sites on prometaphase and metaphase chromosomes were described more precisely upon further investigations (Fukui et al., 1998; Hasterok and Muluszynska, 2000a, 2000b; Hasterok et al., 2006; Hwang et al., 2009; Kim et al., 1998; Snowdon et al., 1997). Some chromosomes of the complements were identified exactly, enabling a detailed karyotype analysis for the genus Brassica.
Lim et al. (2012) analyzed the karyotype of the somatic metaphase of the xBrassicoraphanus line BB#4 by conventional Giemsa-staining, but no cytogenetic studies of the new synthetic intergeneric hybrid line BB#5 have been conducted to date. Here, we carried out FISH karyotype analysis of BB#5 using 5S and 45S rDNA probes, with reference to its parental species B. rapa and R. sativus. In addition, meiotic chromosome pairing patterns were analyzed and the Brassica and Raphanus genomes were discriminated by GISH method.
Materials and Methods
Plant Materials
Seeds and flower buds of the xBrassicoraphanus line BB#5 were provided by the Bio Breeding Institute, Ansung, Korea and seeds of its parental species, Brassica. rapa L. ssp. pekinensis and Raphanus. sativus L. var. rafiphera were provided by the RDA Genebank, Suwon, Korea.
Seeds were sown on moist filter paper in Petri dishes and germinated at 25°C for 48 hours. Approximately 2 cm long root tips were harvested from the germinated seeds, pretreated in 2 mM 8-hydroxyquinoline at 18°C for 5 hours, fixed in aceto-ethanol (1:3 v/v) solution for 2 to 24 hours, and then stored in 70% ethanol until use. The xBrassicoraphanus flower buds were fixed in the same solution for 24 hours and stored at -20°C in 70% ethanol.
Preparation of Somatic Metaphase Chromosome Spreads
A modified version of the method described by Kato et al. (2004) was used to prepare chromosome spreads. Briefly, fixed root tips were thoroughly washed with distilled water, after which the meristem section of the roots was cut out and digested in 2% cellulase (MB Cell, Korea), 1.5% macerozyme (Maxim Bio, USA) and 1% pectolyase (Sigma, Japan) in 150 mM citrate buffer, pH 4.5 for 90 minutes at 37°C. The meristems were then thoroughly washed in ice-cold distilled water, after which the root epidermises were removed and the remaining section containing the dividing cells was pipetted into a tube with chilled aceto- ethanol (1:3 v/v) solution, then suspended by gentle vortexing for 30 seconds. The cells were subsequently collected in the bottom of the tube by centrifugation and re-suspended in aceto- ethanol (9:1 v/v) solution. Finally, the cell suspension was dropped on pre-cleaned glass slides, placed in a humid chamber (37°C) and slowly dried.
Probe Preparation
The procedure described by Lim et al. (2005) was used to obtain the 5S rDNA sequences of genomic DNA from B. rapa L. ssp. pekinensis and the fragment was labelled with dig-11-dUTP (Roche, Germany) by nick translation. The 45S rDNA sequence, containing a 9-kb fragment of 18S-5.8S-25SrDNA genes from wheat (Gerlach and Bedbrook, 1979) was isolated and labelled with biotin-16-dUTP (Roche, Germany) by nick translation. The 200-500 bp labelled DNA fragments were used as probes.
For GISH analysis, total genomic DNA was extracted and purified from young leaves of B. rapa L. ssp. pekinensis and R. sativus L. using the CTAB- extraction method (Allen et al., 2006). The genomic DNA from B. rapa was labelled with dig-11-dUTP (Roche, Germany) by nick translation and used as a probe, while the unlabelled DNA of R. sativus was fragmented at 100°C for 7 minutes and used as a block. The lengths of the probe and the blocking DNA fragments were determined by agarose-gel electrophoresis within the range of 200-500 bp, after which samples were stored at -20°C until use.
Fluorescence in Situ Hybridization (FISH) and Genomic in Situ Hybridization (GISH)
The FISH procedure described by Lim et al. (2005) and Waminal et al. (2012) was used in this study, with some modifications. Briefly, slides were pretreated with RNase A buffer (100 µg・mL-1 RNase A in 2x SSC) at 37°C for 1 hour, after which they were soaked in 0.01 M HCl for 2 minutes. Next, samples were treated with 10% (w/v) pepsin in a 1:100 dilution of 0.01 M HCl for 10 minutes at 37°C and then washed for 5 minutes with 2x SSC. The slides were subsequently post-fixed with 4% paraformaldehyde in 2x SSC, washed with 2x SSC, and dehydrated in ethanol with a series of different concentrations. A hybridization mixture that consisted of 50% formamide, 10% dextransulfate, 2x SSC, 200 ng salmon sperm DNA or 5 ng・µL-1 unlabelled genomic DNA for GISH, and 50 ng・µL-1 of each DNA probe (5S rDNA and 45S rDNA for FISH, or labelled genomic DNA for GISH) was prepared and then adjusted to 40 µL per slide with nuclease free water. The FISH/GISH mixture was denatured at 90°C and cooled on ice for another 10 minutes, then mounted on slides and denatured at 80°C for 2.5 minutes, after which it was placed in an incubator at 37°C overnight (to 16 hours). The next day, the slides were washed in 2x SSC for 30 minutes at RT, 0.1x SSC for 35 minutes at RT and finally in 2x SSC for 15 minutes at 42°C. Dig- labelled and biotin- labelled probes were detected with anti-dig-FITC conjugate (Sigma, USA) and streptavidin-Cy3 conjugate antibodies (Zymed Lab., USA), respectively. The antibodies were reacted on slides at 37°C for 1 hour, then washed in the detection buffer at 37°C. Next, the slides were dehydrated in ethanol series (70, 90, and 100%, 3 minutes each) and air-dried. A 40 µL aliquot of 1:100 DAPI [f.c. 1 µg・mL-1 (Roche, Germany)] in Vectashield (Vector Lab., Inc., USA) reagent was mounted to counterstain the chromosomes and covered with a glass coverslip.
Meiotic Chromosome Pairing Analysis
Meiotic chromosomes were prepared according to Zhong et al. (1996) and Ge and Li (2007) with minor modifications. Young flower buds from 1.6 to 2.0 mm and anthers from 0.8 to 1.0 mm long were used. Fixed flower buds were washed with distilled water for at least 10 minutes. One anther from a flower bud was squashed in 60% acetic acid, air-dried, stained with 4',6-diamidino-2-phenylindole (DAPI), and analyzed using a fluorescence microscope. Microsporocytes in prophase I were selected, and the remaining anthers were then treated with an enzyme mixture of 2% cellulase (MB Cell, Korea), 1% cytohelicase (Yakult, Japan), and 1% pectolyase (Sigma, Japan) for 90 minutes at 37°C. Next, the treated anthers were washed in distilled water for 5 minutes, transferred to a glass slide, then squashed after adding 60% acetic acid and air-dried. Meiotic chromosomes entering late prophase I were scored and meiotic pairing configurations and frequency were analysed using 100 pollen mother cells (PMCs) from different flower buds.
Imaging and Karyotype Analysis
The FISH images of each chromosome were measured three times to obtain the mean values. Homologous chro-mosomes were confirmed based on chromosome length, morphology and, FISH signal patterns. The chromosome arrangements of B. rapa and R. sativus were conducted according to the rDNA distribution patterns, according to the methods described by Lim et al. (2005) and Hwang et al. (2012), respectively. The overall chromosome size of xBrassicoraphanus line BB#5 was estimated by measuring the lengths of short and long arms of chromosomes, excluding satellites and nucleolar organizing regions due to considerable variations in their size in each satellite chromosome and the chromosomes were arranged from longest to shortest. Images were captured using an Olympus BX-51 fluorescence microscope equipped with a charge- coupled device (CCD) camera (CoolSNAP™ cf) and analyzed using the Genus software (version 3.1). Final images were enhanced using Adobe Photoshop CS3. The method described by Levan et al. (1964) was used to classify the chromosome types. Homologous chromosomes were determined based on their lengths, morphological characteristics, and rDNAs signals.
Results
Karyotype Analysis
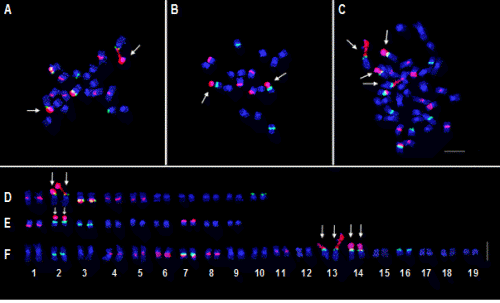
The mitotic chromosome number of Brassica rapa L. ssp. pekinensis was 2n = 20, (Fig. 1A), with lengths ranging from 2.75 to 4.75 µm and a total length of 36.22 µm (Table 1). The chromosome complement is comprised of eight metacentric and two submetacentric homologous chromosomes, respectively. Three and five pairs of 5S and 45S rDNA loci, respectively, were observed near the centromeric regions of B. rapa chromosomes. The 5S rDNA loci were observed on the terminal parts of the short arm of chromosome #2 and #10 and near the centromeric part of the long arm of chromosome #3 (Figs. 1A and 2). In the case of chromosomes #2 and #3, 5S rDNA was juxtaposed with 45S rDNA on the short and long arms, respectively. The 45S rDNA loci of chromosomes #1, #3, #4, and #5 were observed near the centromeric regions of the long arms, while that of chromosome #2, which showed the most intense signal, was on the satellite including the nucleolar organizing region (NOR). Previous studies reported three and five pairs of 5S and 45S rDNA loci in B. rapa (Hwang et al., 2009; Koo et al., 2011; Lim et al., 2005; Xiong and Pires, 2011;) and the same rDNA numbers and distribution patterns were observed in the present study.
The mitotic chromosome number of Raphanus sativus L. var. rafiphera was 2n = 18 (Fig. 1B), with lengths ranging from 2.04 to 3.31 µm and a total length of 24.66 µm (Table 2). The chromosome complement consisted of eight metacentric pairs and one submetacentric homologous chromosome pair. Two pairs of 5S and three pairs of 45S rDNA loci were observed in the R. sativus genome (Figs. 1B and 2). The 5S rDNA loci were observed on the terminal part of the short arm and near the centromeric part of the long arm of chromosomes #2 and #3, respectively. Two 45S rDNA loci were observed near the centromeric region of the long arm of chromosome #1 and the intercalary region of the short arm of chromosome #7. The largest 45S rDNA signal appears on the satellite segment that contains NOR on chromosome #2 to which 5S rDNA is juxtaposed (Fig. 1B). These results coincided with those reported by Hwang et al. (2012).
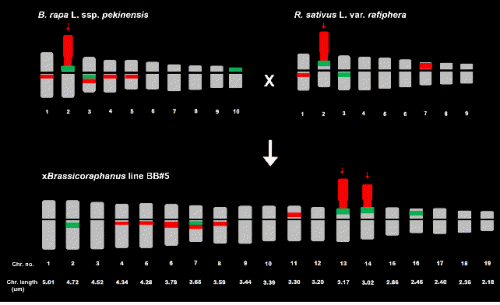
The mitotic chromosome number of xBrassicoraphanus BB#5 was 2n = 38 (Fig. 1C), with lengths ranging from 2.18 to 5.01 µm and a total length of 65.68 µm (Table 3). The chromosome complement is comprised of 17 metacentric and two submetacentric homologous chromosome pairs, respectively. There were five 5S and eight 45S rDNA loci observed on BB#5 chromosomes (Figs. 1C and 3). The 5S rDNA loci were observed near the centromeric regions of the long arms of chromosomes #2 and #7, at the terminal parts of the short arms of chromosomes #13 and #14 while being juxtaposed to the NOR, and on the intercalary part of the short arm of chromosome #16. The 45S rDNA loci were observed near the centromeric regions of the long arms of chromosomes #4, #5, #6, #7, and #8, near that of the short arm of chromosome #11, and on the satellite segment including NOR of chromosomes #13 and #14. According to the rDNA signal patterns, chromosome #2 of BB#5 was similar to chromosome #3 of R. sativus; BB#5 chromosomes #4, #5, #6, and #8 to chromosomes #1, #4, and #5 of B. rapa and chromosome #1 of R. sativus; BB#5 chromosome #7 to chromosome #3 of B. rapa; BB#5 chromosome #11 to chromosome #7 of R. sativus; BB#5 chromosome #16 to chromosome #10 of B. rapa; and BB#5 satellite chromosomes #13 and #14 to satellite chromosome #2 of B. rapa and R. sativus, respectively (Fig. 2).
Due to the small size and similar morphology of the chromosomes, accurate identification of the homologous chromosome using only DAPI staining without any chromosome- specific sequence markers was relatively difficult. Upon FISH karyotype analysis of B. rapa, R. sativus, and xBrassicaraphanus line BB#5 using the 5S and 45S rDNA probes, the difficulties associated with identification of each chromosome were overcome by the presence, absence, position, or size of the rDNA signals. In hybrid line BB#5, all rDNA signals from both the B. rapa and R. sativus genomes were detected, including two pairs of nucleolar organizing chromosomes that each resembled either that from the B. rapa or R. sativus genome. FISH karyotype analysis revealed that 10 of 19 chromosomes of the BB#5 genome were identified as inherited chromosomes from each parental species.
When the FISH karyotype of BB#5 was compared with that of each parental species, no apparent variation in chromosomal number or structure was detected at the microscopic level. Rather, complete sets of the two parental chromosomes appeared to reside in the same nucleus, strongly supporting that xBrassicoraphanus line BB#5 is a true allotetraploid derived from a cross between different genera, which is an extremely rare event in nature.
Meiosis Pairing Analysis
Although B. rapa and R. sativus belong to different genera, they are closely related species in the Brassicaceae family; accordingly, there must be significant homology between the two genomes. Recent genomic studies revealed that B. rapa and R. sativus share a high degree of synteny in their genomes (Kitashiba et al., 2014; Moghe and Shiu, 2014). This suggests the possibility that nonhomologous chromosome pairing can take place via homeologous regions during the early stages of meiotic prophase I, which may lead to the formation of multivalent chromosome pairs between chromosomes of B. rapa and R. sativus.
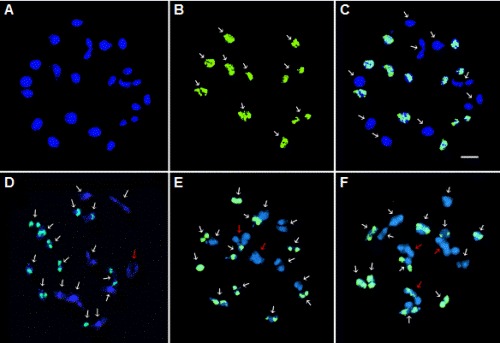
In the present meiotic pairing analysis at prophase I, 19 bivalent pairs were most frequently observed (Figs. 3A and 3C), which accounted for 42% of the 100 PMCs examined (Table 2), whereas the rest consisted of a mixture of bivalent and tetravalent chromosomes. For example, 28% of the PMCs were found to have one tetravalent and 17 bivalents (Fig. 3D), while 24% contained two tetravalents and 15 bivalents (Fig. 3E). Moreover, 6% of the PMCs formed three tetravalents and 13 bivalents (Fig. 3F). These findings indicate that significant homology exists between B. rapa and R. sativus chromosomes and they are able to physically interact with each other to form multivalents during meiosis. Multivalent chromosome formation can generate various chromosomal changes via abnormal segre-gation in meiosis (Comai, 2005; Fujii and Ohmido, 2011; Leitch and Leitch, 2008). These findings suggest that there could be a high chance of chromosome rearrangement, such as translocation and deletion, when two nonhomologous chromosomes exchange segments, which conceivably leads to genome instability and infertility in newly synthesized allopolyploid plants. Despite the formation of multivalent chromosome pairs during meiosis, GISH analysis of BB#5 clearly showed 10 bivalent chromosome pairs with green fluorescence signals, although the remaining chromosome pairs displayed no signals when B. rapa genomic DNA was used as a probe and R. sativus DNA as blocking DNA. These findings indicate that 10 pairs of chromosomes are B. rapa- specific and there is no massive chromosome rearrangement in xBrasscoraphanus.
Poor fertility is always an obstacle that must be overcome in allopolyploid breeding (Prakash et al., 2009). However, the intergeneric hybrid line BB#5 displays exceptionally high seed fertility and stable inheritance of morphological characters as demonstrated in another xBrassicoraphanus line (Lee et al., 2011). Overall, 42% of pollen mother cells analysed showed 19 complete bivalent chromosomes even though some appeared to have different numbers of tetra-valents with different frequencies. These results strongly supports that stabilization of the intergeneric polyploid xBrassicoraphanus line BB#5.
Discussion
We analyzed the karyotype of somatic metaphase chro-mosomes by fluorescence in situ hybridization (FISH) and meiotic chromosome pairing of pollen mother cells (PMCs) by genomic in situ hybridization (GISH) in the newly synthesized allotetraploid xBrassicoraphanus line BB#5.
Cytogenetic studies in natural and synthetic allopolyploids have been reported by many researchers (Bennett et al., 1992; Fujii and Ohmido, 2011; Yao et al., 2010). The FISH technique has been shown to be especially useful for more accurate karyotype analysis because it provides good diagnostic markers for identification of each chromosome. Tandem repeat sequences such as 5S and 45S rDNAs have primarily been used as probes for FISH (Fujii and Ohmido, 2011; Fukui et al., 1998; Hasterok et al., 2006; Kim et al., 1998; Lim et al., 2007; Snowdon et al., 1997). Several researchers applied FISH in Brassica rapa using 5S and 45S rDNA probes and reported three and five 5S and 45S rDNA pairs, respectively (Hwang et al., 2009; Koo et al., 2011; Lim et al., 2005; Xiong and Pires, 2011). Hwang et al. (2012) analyzed the FISH karyotype of Raphanus sativus and reported two and three 5S and 45S rDNA pairs, respectively. In our FISH karyotype analyses of Brassica rapa and Raphanus sativus using 5S and 45S rDNA probes, the same rDNA distribution patterns were observed and xBrassicaraphanus line BB#5 appeared to have both combined rDNA patterns of two parental species. In addition, GISH was successful at identifying subgenomes of interspecific hybrid in Brassicaceae (Howell and Armstrong, 2013; Lim et al., 2012; Wang et al., 2006; Yao et al., 2010). Lim et al. (2012) applied GISH to discriminate each parental subgenomes on somatic metaphase chromosomes of the xBrassicoraphanus line BB#4 and confirmed 20 chromosomes of B. rapa in the hybrid genome. Here, we applied GISH to the meiotic prophase I cell of BB#5. Ten pairs of Brassica bivalent chromosomes could clearly be discriminated in the intergeneric hybrid line.
Allopolyploids generally show genetic and phenotypic changes due to gene dosage effects, the existence of different genomes in a nucleus, variations in chromosome number, unbalanced parental genome contributions, and chromosomal rearrangements (Leitch and Leitch, 2008). Fujii and Ohmido (2011) observed several chromosome variations and aberrant meiosis in a synthetic hybrid and suggested that the genetic instability of the hybrid polyploid was due to abnormal chromosome number and structure caused by aberrant meiosis. In the present study, intergeneric allotetraploid xBrassicoraphanus line BB#5 showed very stable mitotic chromosome constitution and meiotic pairing pattern, supporting the high seed fertility and morphological uniformity of BB#5 even though we could not rule out the possibility of chromosomal rearrangement or structural changes in the next generations because of the appearance of different number of tetravalent with different frequencies according to the PMCs. However, such stable behaviour in somatic and meiotic cells is very rare in natural and synthetic intergeneric hybrids. Gaeta et al. (2007) observed that genomic changes in resynthesized Brassica napus were rare in the first generation, but that the changes were much more frequent in the fifth generation. Continuous cytogenetic and morphological study of BB#5 should be conducted to confirm whether the chromosomal stability will be maintained during successive generations.
Consequently, a very stable xBrassicoraphanus line BB#5 was newly developed by microspore mutation, which was certified by stable chromosome composition and meiotic pairing patterns in this study. Further molecular cytogenetic studies including FISH and GISH using more detailed chro-mosome-specific markers and comparative studies related to the genome sequencing are necessary to elucidate a more accurate genome structure of the hybrid line. The results presented herein will be useful for further genomic study of xBrassicoraphanus lines and their improvement as new promising breeding varieties.