Introduction
Materials and Methods
Plant Materials
RNA Isolation
Reverse Transcription (RT)-PCR
Stress Treatments
Analysis of DNA and Protein Sequences
Results
Isolation of the IbCAD Gene Family and Sequence Analysis
Structural Characteristics of IbCADs
Differential Expression of the IbCAD Family Under Environmental Stresses
Discussion
Introduction
Cinnamyl alcohol dehydrogenase (CAD) catalyzes the reduction of hydroxycinnamyl aldehydes (p-coumaraldehyde, coniferaldehyde, and sinapaldehyde) to their corresponding alcohols (monolignol) and is regarded as a key enzyme in lignin biosynthesis (Boerjan et al., 2003). It has been shown that CAD activity is regulated developmentally in various plant species. CAD is expressed abundantly in the developing xylem, which is consistent with the fact that the secondary walls of the xylem cells are
the major site of lignin synthesis during development.
On the other hand, it is likely that CAD is involved in defense response. In other words, CAD responds to stress by reinforcing the cell wall. For example, when wheat leaves were treated with a fungi elicitor, a dramatic increase in the activity of CAD that specifically oxidizes sinapyl alcohol was observed (Mitchell et al., 1994). In jack pine (Pinus banksiana) cell culture, a fungal elicitor induced lignification. Furthermore, an increase in CAD activity was measured in the seedlings of P. abies after ozone fumigation (Galliano et al., 1993). Therefore, the CAD protein is a possible molecular marker for lignification during development and plant stress responses.
Many CAD genes in several plants have been studied. This gene belongs to a multigene family, with 9 members in Arabidopsis (Kim et al., 2004), 14 members in sorghum (Saballos et al., 2009), 12 members in rice (Tobias and Chow, 2005), 16 members in poplar (Barakat et al., 2009), and 26 members in pear fruit (Cheng et al., 2017). These CAD isoforms showed differential expression during plant development and environmental conditions and might have different kinetics and substrate preferences. The CAD enzyme isoforms identified so far showed that one isoform might be involved in developmental lignifications, whereas the other isoforms could affect the composition of defensive lignins and wall-bound phenolics. The developmental and defense lignins appear to form according to separate signal transduction pathways. Different functional and biochemical properties, such as substrate specificities, of CAD proteins have been attributed to the different isoforms of the CAD enzymes identified in various plant species (Li et al., 2001; Kim et al., 2004, 2007; Bomati and Noel, 2005). However, no studies have been conducted on CAD isoforms in root crops.
The sweet potato CAD gene (Ipomoea batatas Cinnamyl Alcohol Dehydrogenase, IbCAD1) and its promoter were reported previously (Kim et al., 2010). The suspension-cultured cells of sweet potato can be grown under high oxidative stress conditions. The most significantly abundant expressed sequence tag (EST) in the suspension-cultured cells was a CAD gene (Kim et al., 2006). The IbCAD1 promoter was strongly induced by both abiotic (cold) and biotic (wounding) stresses, including the reactive oxygen species (ROS) inducers (methyl viologen and hydrogen peroxide). The role and specificity of different CAD isoforms involved in the lignification process and expressed in response to environmental factors in root crops are still unclear. Therefore, the purpose of this study was to identify 13 CAD isoforms in sweet potato suspension cells and to enhance the knowledge of the role of each CAD gene. The structure features of the IbCAD family genes were characterized, and their expression patterns were also analyzed under environmental stresses.
Materials and Methods
Plant Materials
Sweet potato (Ipomoea batatas ‘Yulmi’) slips were provided by the National Honam Agriculture Experiment Station and maintained in a pot at 25 - 28°C in a greenhouse under a 16/8-h (light/dark) photoperiod. The apical shoot was excised, planted for propagation, and grown for 3 or 4 weeks.
RNA Isolation
Sweet potato plant tissues (1 g) were ground to a fine powder in liquid nitrogen and homogenized in extraction buffer according to Kim et al. (2013). The aqueous phase containing nucleic acids was precipitated with isopropanol. The pellet was washed and resuspended in 1 mL Tris-EDTA buffer. RNA was precipitated with 2 M LiCl overnight at 4°C. The precipitated RNA was centrifuged, and the resulting pellet was washed with 70% ethanol and resuspended in TE buffer. The RNA was stored at - 70°C until use.
Reverse Transcription (RT)-PCR
RT-PCR was conducted to analyze the transcription pattern of the IbCAD family. Group-specific primers were designed and confirmed the primer specificity. For cDNA synthesis, the reverse transcription was performed using total RNA (2 µg), oligo (dT)20 as a primer, and SuperscriptTM II reverse transcriptase (Invitrogen, Carlsbad, CA, USA). One-tenth of the cDNA reaction was used as a template for each of the following PCR amplifications. The PCR cycling conditions were as follows: initial denaturation at 94°C for 5 min, followed by 30 cycles of denaturation at 94°C for 30 s, annealing at 50 - 60°C (depending on the annealing temperature of each group-specific primer), extension at 72°C for 1 min, and a final extension step at 72°C for 7 min. As an internal control, TUBULIN gene was amplified. All group-specific primers are listed in Table 1. The PCR products were electrophoresed on a 1% agarose gel.
Stress Treatments
For stress treatments, the apical shoots (15 cm) of sweet potato plants were excised and then planted for 3 weeks to synchronize their growth. Among them, 16 plants of similar growth stage in pots were selected and subjected to 15°C cold treatment. Treatment of plants at 10°C was performed the same way as the 15°C cold treatment. Afterward, the second and third leaves from apical tips, petioles, and roots were collected from 3 plants out of 16 plants for each sample.
For H2O2 stress, the second and third leaves from the apical tips were detached and incubated in 0.44 M H2O2 solution at 25°C for 24 h. For wounding stress, the leaves were detached and cut into pieces. The petioles were cross-cut to 10 mm in length, and roots were cut into 10-mm-long sections after washing and put on a moistened filter paper. Petioles were incubated at 25°C under continuous illumination, whereas roots were incubated in the dark (Sasaki et al., 2002). After stress treatment, the plant samples were collected and then frozen in liquid nitrogen and maintained at - 70°C until further analysis. Three plants were used for each time point as replicates. An aliquot of 2 µg of RNA of three biological replicates extracted from leaves, petioles, and roots at an indicated time was pooled and used for RT-PCR analysis.
Analysis of DNA and Protein Sequences
A sequence similarity search was performed using the Basic Local Alignment Search Tool (BLAST) program against protein sequence databases. The nucleotide and deduced amino acid sequence analysis was performed using the DNAStar software. A multiple amino acid sequence analysis was performed with the GENEDOC and CLC Workbench 4 program using the ClustalX algorithm (Nicholas et al., 1997). The phylogenetic tree was drawn using the Tree View program (Page, 1996).
Results
Isolation of the IbCAD Gene Family and Sequence Analysis
In a previous study, the cDNA library was constructed from sweet potato suspension cells, and 1,411 ESTs were obtained from the cDNA library (Kim et al., 2006). The most abundant EST was a CAD gene (IbCAD1). The exhaustive BLAST searches of 13 cDNAs showed high sequence similarity to the known CAD genes. The full sequences of cDNA clones encoding CAD were obtained by sequencing from both ends and designated as IbCAD1 to IbCAD13.
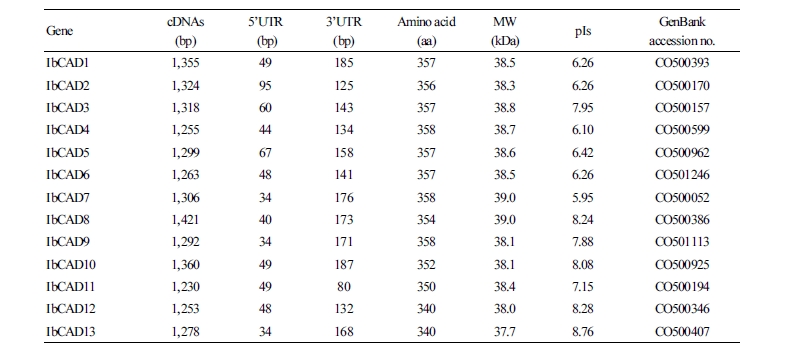
The length of cDNA was 1,230 to 1,421 bp, and the cDNA contained a 5'-untranslated region of 34 to 95 nucleotides and a 3'-end of 80 to 187 nucleotides. The open reading frame (ORF) encodes a protein of 340 to 358 amino acids with a calculated molecular mass of 37.7 to 39.0 kDa and an isoelectric point (pI) of 5.95 to 8.76 (Table 2).
To examine the similarity of IbCADs to the known CAD proteins in detail, the full-length protein sequences were compared to the sequences of previously reported CADs from other plant species (Fig. 1). The phylogenetic tree showed that all CAD protein sequences were divided into four classes, probably reflecting their biological similarities. Of the 13 IbCADs, 12 (IbCAD1-IbCAD12) belonged to Class II and 1 (IbCAD13) belonged to Class IV.
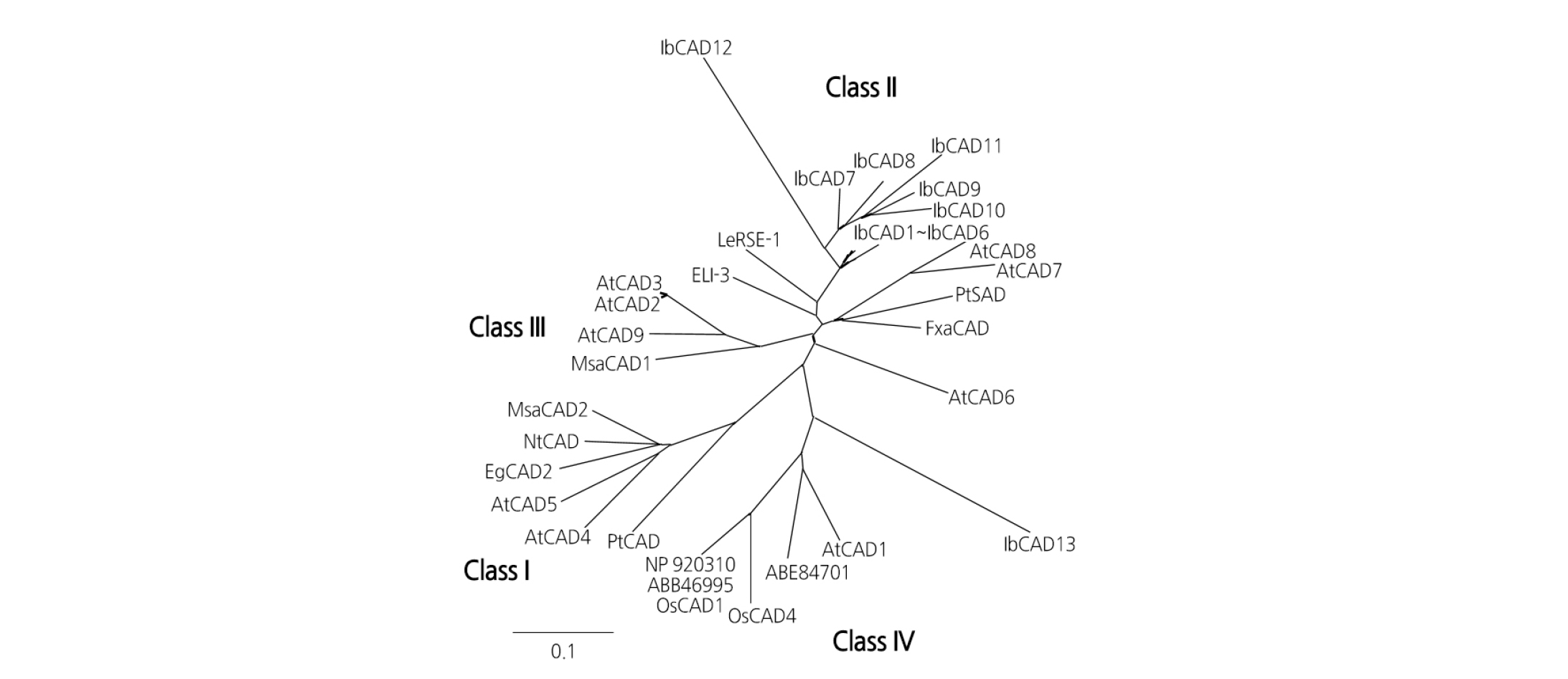
Fig. 1. Phylogenetic tree of cinnamyl alcohol dehydrogenases (CADs) in evolutionarily distant plants. The phylogenetic tree was built with the amino acid sequences indicated below. The following proteins, with their accession numbers in parentheses, were used to build the tree: NtCAD of Nicotiana tabacum (X62343), EgCAD2 of Eucalyptus gunnii (X88797), MsaCAD1 of Medicago sativa (AF083332), MsaCAD2 of M. sativa (AF083333), PtCAD of Populus tremuloides (AF217957), PtSAD of P. tremuloides (AF273256), FxaCAD of Fragaria x ananassa (AF320110), ELI-3 of Petroselinum crispum (X67817), AtCAD1- AtCAD9 of Arabidopsis thaliana (AY288079, AY302077, AY302078, AY302081, AY302082, AY302075, AY302079, AY302080, and AY302076), and OsCAD1 (NP_001064283.1) and OsCAD4 (NP_001068303.1) of Oryza sativa. ABB46995 and NP_920310 are the accession numbers of mannitol dehydrogenase and a putative CAD from O. sativa, respectively. ABE84701 is the accession number of the alcohol dehydrogenase superfamily from Medicago truncatula.
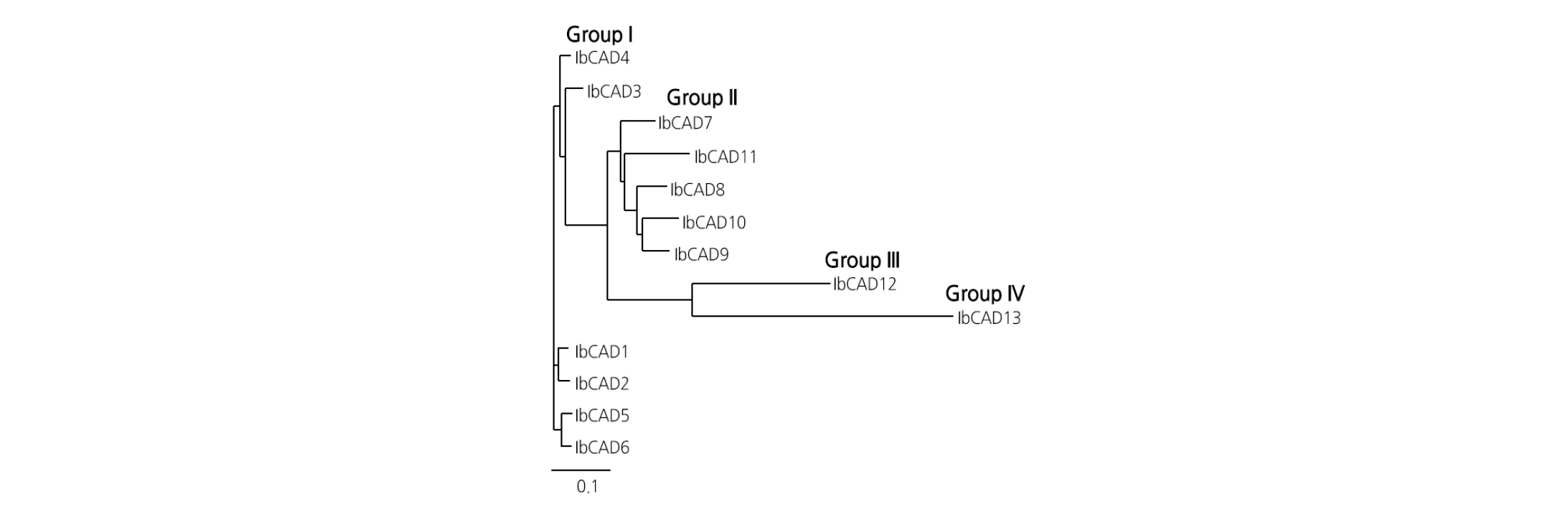
The amino acid sequence identity among the 13 IbCADs ranged from 28.3 - 99.4%. IbCADs were categorized into four major groups according to their amino acid sequence similarities (Fig. 2). In Group I, IbCAD1-IbCAD6 showed the highest amino acid sequence homology (92.2 - 99.4%) among themselves. In Group II, IbCAD7-IbCAD11 showed 78.6 - 88.7% amino acid sequence identity among themselves and 68.9 - 79.4% identity with Group I. In Group III, IbCAD12 showed a relatively low amino acid sequence homology with Group I (45.8 - 50.5%) and Group II (45.5 - 53.3%). In Group IV, IbCAD13 showed the lowest amino acid sequence homology with the other three groups, being 28.3 - 29.1% with Group I, 28.7 - 34.3% with Group II, and 37.2% with Group III.
Structural Characteristics of IbCADs
Amino acid sequence alignment of 13 IbCAD proteins was performed to characterize their structural features (Fig. 3). The amino-terminal amino acid sequences of the 12 IbCAD proteins belonging to all groups, except Group IV, shared high sequence homology, whereas the carboxyl-terminal amino acid sequences were highly similar within the same group, but different between groups. The amino acid sequences of IbCADs in all groups present the consensus sequences GHE(X)2G(X)5G(X)2V and GD(X)9,10C(X)2C(X)2C(X)7C for Zn1- and Zn2-binding motifs, respectively. Furthermore, the Cys residues at positions 104, 107, 110, and 118 have been identified as structural zinc ion-coordinating residues; thus, the IbCAD genes are a family of zinc-dependent alcohol dehydrogenases. The coenzyme (NADPH)-binding domain- containing glycine-rich repeat motif GXG(X)2G, which is a characteristic of a CAD enzyme, is present only in Groups I and II.

Fig. 3. Amino acid sequence alignment of 13 IbCADs and structural features of substrate binding in each group. Identical amino acids are dotted. Boxes indicate the Zn1- and Zn2-binding motifs; the line indicates the NADPH-binding motif; ● indicates the key bulky active sites; ▾indicates the structural Zn ion-coordinating residue; and ★ indicates the key determinant of substrate specificity.
In previous reports, several classifications of the CAD family have been proposed on the basis of substrate specificity, expression patterns, or gene structure (Sibout et al., 2003; Barakat et al., 2009; Guo et al., 2010). Fig. 4 shows a partial sequence alignment of the developmental and nonclassical CADs, indicating that their key residues were quite different. The amino acid sequence alignment showed that IbCAD1 in Group I had certain residues similar to those in nondevelopmental CADs. Nonclassical CADs shared the key bulky aromatic residues, including Trp62 and Phe286, at their active site. This Phe residue was replaced by Pro286 in developmental CADs. Gly299, the key determinant of substrate specificity, especially in sinapaldehyde substrate, also exhibited identity; it was substituted with Phe299 in developmental CADs. Although nonclassical CADs (PtSAD and FxaCAD) and developmental CADs (PtCAD and AtCAD4) catalyze the same reaction and share some sequence identity, their important active sites for specificity were quite different.
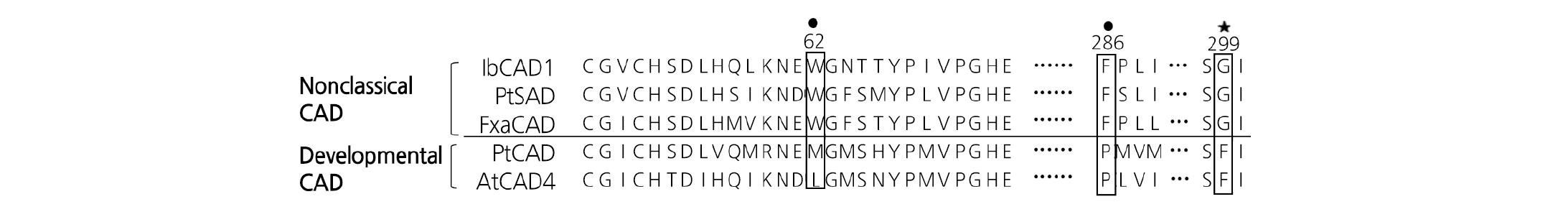
Fig. 4. Partial sequence alignment of nonclassical and development-related cinnamyl alcohol dehydrogenases (CADs). Alignment of the IbCAD1 key residues with PtSAD, FxaCAD, and development-related CADs, PtCAD, and AtCAD4. ● indicates the key bulky active sites, and ★ indicates the key determinant of substrate specificity.
Differential Expression of the IbCAD Family Under Environmental Stresses
The expression patterns of IbCAD family genes were investigated by RT-PCR analysis. The following cDNAs were chosen as a representative of each group: IbCAD3 for Group I, IbCAD7 for Group II, IbCAD12 for Group III, and IbCAD13 for Group IV. The expression of IbCAD family genes was investigated to determine whether their transcripts were regulated by abiotic stresses, e.g., cold, H2O2,andwounding (Fig. 5). Because sweet potato is grown in tropical regions, it is cold sensitive and could be damaged at chilling temperatures (15°C). The lowest safe temperature that sweet potato can withstand is 10°C (Lukatkin et al., 2012). Therefore, the sweet potato plants were exposed to 15 and 10°C for 24, 48, and 72 h to determine the transcriptional regulation of IbCADs in response to cold stress. The transcript levels of IbCADs varied depending on tissues and temperatures. The expression of Group I-III IbCAD genes was low in the leaves. However, Group IV IbCAD was strongly expressed in all tissues.
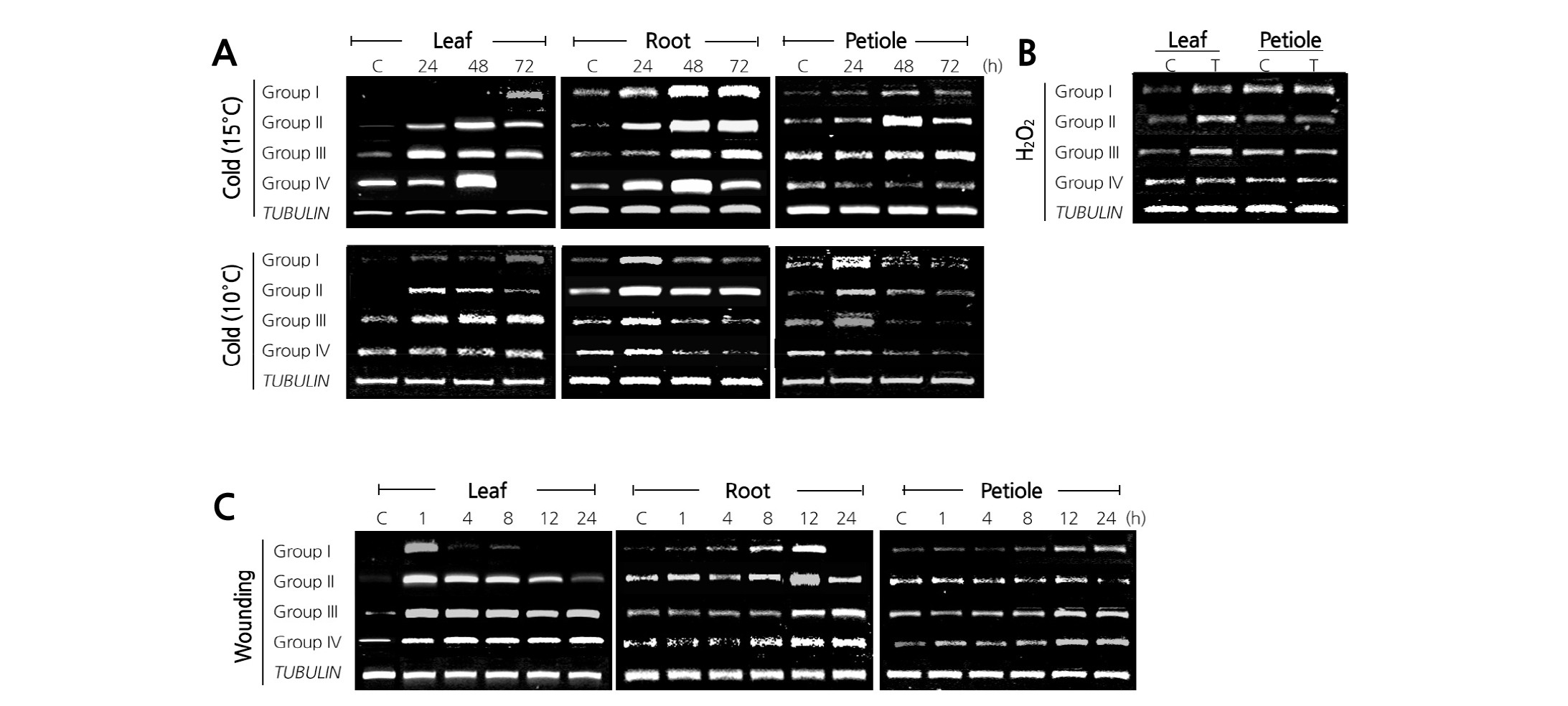
Fig. 5. Differential expression of the IbCAD family genes under environmental stresses. (A) For cold stress treatment, the sweet potato plants were exposed to temperatures of 10 and 15°C for 24, 48, and 72 h. (B) For hydrogen peroxide treatment, the second or third leaves from the apical tips were removed from a plant and incubated in 0.44 M H2O2 solution at 25°C for 24 h. (C) For wounding treatment, the sweet potato plants were cut into small pieces and incubated for 1, 4, 8, 12, and 24 h. Total RNA was isolated from the leaves, petioles, and roots, and an RT-PCR analysis was performed using all group-specific primers. The TUBULIN gene was used as an internal control.
At 15°C, the IbCAD family genes showed diverse expression patterns in leaves. The transcript levels of Group I IbCAD genes were induced at 72 h. In Group II, the transcripts were steadily increased and reached the maximum level at 48 h and then decreased at 72 h. In Group III, the transcripts were induced at 24 h and then decreased gradually. In Group IV, the highest level was observed at 48 h and then dramatically decreased at 72 h. In roots, the transcripts of IbCAD family were steadily increased and reached the maximum level at 48 h. The mRNA induction in roots was much higher than that in the other tissues. However, in petioles, the expression of IbCADs, except those in Group II, was not significantly responsive to cold stress. The overall expression of IbCADs steadily increased at 15°C and was highly induced with prolonged chilling exposure. Under cold treatment at 10°C, the IbCAD transcripts of Groups I-III in leaves were highly induced after 24 h and then their expression patterns varied depending on the groups. However, no change was observed in Group IV IbCAD transcripts. In roots, the IbCAD transcripts increased at 24 h and then continuously decreased at 10°C. The expression pattern in petioles was similar to that in roots (Fig. 5A).
The effect of ROS in the IbCAD gene was investigated by H2O2 treatment. The transcript levels in leaves in response to H2O2 were increased in Groups I-III but remained unchanged in Group IV. H2O2 had little effect on the expression of the IbCAD family in petioles (Fig. 5B).
The expression patterns of the IbCAD family were analyzed after wounding treatment. In leaves, the rapid induction of expression of all IbCADs was observed at 1 h. However, the IbCAD expression rapidly decreased in Group I and gradually decreased in Group II after 1 h. In Groups III and IV, the transcripts increased at 1 h, but they remained until 24 h after treatment. In roots, the IbCAD transcripts steadily increased and reached the maximum level at 12 or 24 h. In petioles, IbCADs were induced after 12-h treatment in Group I, but no clear accumulation of IbCAD transcripts was observed in the other three groups (Fig. 5C).
Discussion
In this study, 13 CAD cDNAs (IbCAD1-IbCAD13) were isolated from the EST library of sweet potato suspension- cultured cells. According to the phylogenetic analysis, the previously known plant CAD proteins were classified into four classes (Classes I-IV). Class I, known as bona fide CAD, proteins are involved in monolignol biosynthesis. The CAD proteins AtCAD4 and AtCAD5 (Arabidopsis), SbCAD2 (sorghum), and OsCAD2 (rice) have been characterized and play a significant role in developmental lignification (Kim et al., 2004; Tobias et al., 2005; Saballos et al., 2009). Twelve IbCADs, except IbCAD13, were grouped into Class II. It is known that several CAD genes in Class II are related to biotic and abiotic stresses as a defense mechanism in plants (Logemann et al., 1997; Kim et al., 2010; Cheng et al., 2013; Choi et al., 2016; Eom et al., 2016). PtSAD catalyzes sinapyl alcohol biosynthesis and is responsible for syringyl lignin deposition in poplar (Li et al., 2001). ELI3 from celery is strongly activated in elicitor-treated cells and in fungal infection sites (Schmelzeret al., 1989). FxaCAD from strawberry displays high activity for sinapaldehyde and possibly deposition of syringyl lignin (Blanco-Portales et al., 2001). AtCAD7 and AtCAD8 from Arabidopsis were also induced after pathogen infection (Kiedrowski et al., 1992). Class IV includes IbCAD13, CADs from rice, an alcohol dehydrogenase superfamily from Medicago truncatula, and AtCAD1 from Arabidopsis (Tobias et al., 2005; Eudes et al., 2006). AtCAD1 is less similar to the closest group and the most distant protein among Arabidopsis CADs. The structural characteristics of CAD proteins are important for their substrate specificity. Although they have an important amino acid domain implicated in CAD function, they showed sequence divergence in the residues, implicating substrate specificity. Lauvergeat et al. (1995) suggested that CAD isoforms might exhibit different levels of substrate specificity, necessitating the quantitative evaluation of kinetic parameters for the family members with several putative substrates. Thirteen IbCADs were regrouped into four groups (Groups I-IV) based on amino acid similarity and residues for substrate specificity.
The presence of a consensus motif conserved in the CAD proteins is in agreement with a CAD function. All IbCAD groups have a Zn-binding motif, catalytic Zn ion-coordinating residue, and structural Zn ion-coordinating residue; thus, the IbCAD family is a family of zinc-dependent alcohol dehydrogenases. A NADP+-binding domain exists as a coenzyme for CAD enzymatic activity only in Groups I and II. In Group III, IbCAD lacks the key residues representing substrate specificity; thus, it was not expected to specifically attracted substrate. An IbCAD with markedly different residues was found in Group IV.
It was shown that the key residues of developmental and nonclassical CADs were quite different. Group I IbCADs, which have a sinapaldehyde-specific residue, are structurally similar to PtSAD. Gly299 is a key determinant of substrate specificity in Group I IbCADs. The specificity and efficiency of the enzyme reaction in Group I IbCADs might be different from the other IbCAD groups. Bomati and Noel(2005) demonstrated that major changes in specificities and catalytic efficiencies could occur as a result of minimal changes in active-site residues. Although the function of PtSAD was not verified, they suggested that PtSAD was structurally most closely related to the dehydrogenases functioning in plant defense. From these structural characteristics, IbCAD Group I probably is more closely related to sinapyl alcohol dehydrogenase (SAD) enzyme clustering with defense-related enzymes likely exhibiting sinapaldehyde specificity.
The expression profiling of IbCADs under different conditions could provide an insight into the physiological function of monolignols associated with plant defense. The IbCAD genes in each group showed diverse and complex expression patterns in different tissues exposed to cold, H2O2,and wounding stresses. The expression of IbCADs was induced and delicately regulated by low temperature (10 and 15°C). It seemed that the cold-responsive enzyme could compensate for the metabolic reaction of plant growth and contributed to protection from cold-induced oxidative stress. The cold stress increases the production of phenolics, which are incorporated subsequently into the cell wall either as suberin or lignin (Logemann et al., 1997). The lignin deposition in the cell wall could maintain the water status by acting as a water barrier. Flax CADs (LuCADs) are also induced by cold stress; LuCAD1B, especially, is probably involved in low-temperature- induced lignin synthesis (Preisner et al., 2018). H2O2 is directly involved in certain defense reactions, such as lignification and oxidative cross-linking of cell wall proteins, thereby creating a physical barrier against pathogens (Solecka, 1997). Although H2O2 had little effect on the expression of IbCADs in petioles, the transcript level was increased in leaves. A similar expression pattern was reported in kenaf HcCAD, indicating the important role of H2O2 in lignin polymerization (Choi et al., 2016). Many phenylpropanoid compounds were induced by wounding stress. Expression of IbCADs was predominantly induced in leaves by wounding. Like IbCADs, the induction of the CAD transcript by wounding has been reported in other plants, such as alfalfa, ryegrass, and kenaf (Brill et al., 1999; Lynch et al., 2002; Choi et al., 2016). The responses to mechanical wounding and pathogen attack in plants are often similar, including the rapid accumulation of phenolic compounds, lignin, and suberin at the site of injury (Levine et al., 1994).
An increase in the expression of sweet potato IbCADs under environmental stresses suggests an increased carbon flux into the lignin pathway. The differential response of the IbCAD family genes under different stresses might indicate a more complex nature of the CAD gene expression in sweet potato. The substrate specificity and key active enzyme site of IbCADs were different for each group according to amino acid sequence analysis. These results indicate a difference in the expression profile of the IbCADs. Different stresses might trigger distinct signal transduction pathways and activate the transcription of each group of IbCAD genes. These diverse expression patterns represent an important function of each IbCAD, mostly classified as Class II, for tolerance to environmental stresses.