Introduction
Materials and Methods
Plant Materials and DNA Isolation
S-RNase PCR-CAPS Analysis
PpSFBB-γ PCR-CAPS Analysis
SSR Analysis
Results and Discussion
Analysis of S-genotypes using S-RNase from the Style and SFBB-γ from Pollen
SSR Analysis
Introduction
Pear (Pyrus spp.) is an important fruit tree worldwide and has been cultivated in more than 50 countries. In Korea, pear breeding began in the late 1920s by the National Institute of Horticultural and Herbal Science (NIHHS) of the Rural Development Administration (RDA). The goals of this breeding program include improving fruit quality in terms of fruit size, acidity, sugar content, flesh firmness and core ratio; developing disease and pest resistance for black spot, scab and mites; and developing laborsaving varieties by inducing self-compatibility (Shin et al., 2002). Korean pear cultivars ‘Chuwhangbae’, ‘Hanareum’, ‘Josengwhangkeum’, ‘Shincheon’ and ‘Sooyoung’ were bred using ‘Niitaka’ as a parent because ‘Niitaka’ ripens in mid-October at Suwon and has a high yield potential, producing large fruit with an attractive appearance and good storability (Kim et al., 1986; Hwang et al., 2002, 2005a, 2005b; Shin et al., 2007). The ‘Niitaka’ cultivar was developed in 1915 by a breeding program in Japan (Kikuchi, 1927). Identifying with an attractive appearance and good storability (Kim et al., 1986; Hwang et al., 2002, 2005a, 2005b; Shin et al., 2007). The ‘Niitaka’ cultivar was developed in 1915 by a breeding program in Japan (Kikuchi, 1927). Identifying the parent-offspring relationships of pear cultivars is important for current efforts to improve breeding efficiencies; however, some Japanese pear cultivars, including ‘Housui’ (Ishimizu et al., 1998), ‘Kisui’ (Hiratsuka et al., 1998), ‘Tanzawa’ (Castillo et al., 2001), ‘Oharabeni’ (Kim et al., 2007) and ‘Niitaka’ (Takasaki et al., 2004), exhibit discrepancies from their reported pedigrees in terms of skin type and self-incompatibility traits.
‘Housui’ (syn. ‘Hosui’) is a hybrid cultivar that was reported to be developed through a cross between ‘Ri-14’ and ‘Yakumo’ in 1954 (Kajiura et al., 1974). However, ‘Housui’ has russet skin and an S3S5 genotype for self-incompatibility. The reported parents, ‘Ri-14’ (S1S2 ) and ‘Yakumo’ (S1S4 and/or S2S4 ), have smooth skin and a different genetic makeup for self-incompatibility (Ishimizu et al., 1998). Kimura et al. (2003) used 20 SSR markers to confirm that ‘Ri-14’ and ‘Yakumo’ were not the parents of ‘Housui’. The parents of ‘Housui’ were found to be ‘Kousui’ (female parent) and ‘Hiratsuka 1 gou’ (syn. ‘I-33’) (male parent) by Sawamura et al. (2004) using 61 SSR markers.
Similarly, ‘Niitaka’ was reportedly derived from a cross between the pear cultivars ‘Amanogawa’ and ‘Imamuraaki’ (Kikuchi, 1927). The S-genotypes of ‘Amanogawa’ and ‘Imamuraaki’ are S1S9 and S1S6, respectively (Kim et al., 2002; Takasaki et al., 2004), and the S-genotypes of any offspring of a cross between these cultivars would have to be S1S6 or S6S9. However, the S-genotype of ‘Niitaka’ is S3S9, leading Takasaki et al. (2004) to suggest that ‘Amanogawa’ and ‘Imamuraaki’ may not be the parents of ‘Niitaka’. Two other cultivars, ‘Chojuro’ and ‘Shinchu’, have been suggested as candidate parents of ‘Niitaka’, because they were used as genetic resources during the breeding of ‘Niitaka’ in 1915 and have S3-RNase (Kajiura and Sato, 1990; Washio et al., 2006). Here, to determine the parents of ‘Niitaka’, we analyze the S-genotypes of seven pear cultivars based on markers specific for S-RNase and SFBB-γ and determine their SSR genotypes using 65 apple (Malus spp.) SSR markers and 42 pear SSR markers.
Materials and Methods
Plant Materials and DNA Isolation
Young leaves of six pear cultivars, ‘Amanogawa’, ‘Imamuraaki’, ‘Chojuro’, ‘Shinchu’, ‘Niitaka’ and ‘Housui’, and the pear line ‘126-29’ (an offspring from a cross between ‘Niitaka’ and ‘Housui’), were collected at the National Institute of Fruit Tree Science (Ibaraki, Japan). The leaves were frozen in liquid nitrogen and stored at -80ºC. Genomic DNA was extracted from the leaves by the method of Yamamoto et al. (2006) and used for genotyping with SSR markers and PCRCAPS analysis of S-genes.
S-RNase PCR-CAPS Analysis
PCR was performed using the S-RNase-specific primers FTQQYQ and PSprI, as described by Kim et al. (2007). The amplified S-RNase fragments were digested with the haplotype-specific restriction enzymes SfcI (S1 ), XbaI (S2 ), PpuMI(S3 and S5 ), AlwNI (S5 ), MluI (S6 and S7 ) and BstBI (S9 ) and analyzed by agarose gel electrophoresis as described by Kim et al. (2002) and Takasaki et al. (2004).
PpSFBB-γ PCR-CAPS Analysis
PpSFBB-γ fragments were amplified using the primers PpFBXf7 and PpFBXr3 and subjected to CAPS (cleaved amplified polymorphic sequence) analysis (Kakui et al., 2007). Derived CAPS (dCAPS) analysis was performed to detect PpSFBB-γ(Neff et al. 1998). PpSFBB-γ fragments were amplified with the 1 bp-mutated primer GdCAPSS2g1-Rsa and reverse primer PpFBXr11. PCR products were digested with TaqI (S1 ), RasI (S2 ), HpyCH4IV (S3 ), AflII (S5 ), DdeI (S6 ) and HaeIII (S9). TaqI and SmaI digestions were incubated for 3 h at 65°C and 30°C, respectively, and the other endonucleases were incubated for 3 h at 37°C. The digested fragments were separated on 2% agarose gels in TAE buffer, stained with ethidium bromide and visualized under UV illumination.
SSR Analysis
Genotyping was conducted using 107 SSR markers, including 65 SSR markers derived from apple and 42 SSR markers from pear (Yamamoto et al., 2004). Genomic DNA was isolated using the DNeasy Plant Mini Kit (QIAGEN, Germany). PCR amplification was performed according to the method of Yamamoto et al. (2002) with one modification; the forward primers were labeled with a FAM fluorescent dye. PCR products were separated by capillary electrophoresis on a POP-4 polymer using a Genetic Analyzer 3100 (PE Applied Biosystems, USA). The size of the amplified bands was calculated using an internal DNA standard (GeneScan-400HD ROX, PE Applied Biosystems) and GeneScan software (PE Applied Biosystems).
Results and Discussion
Analysis of S-genotypes using S-RNase from the Style and SFBB-γ from Pollen
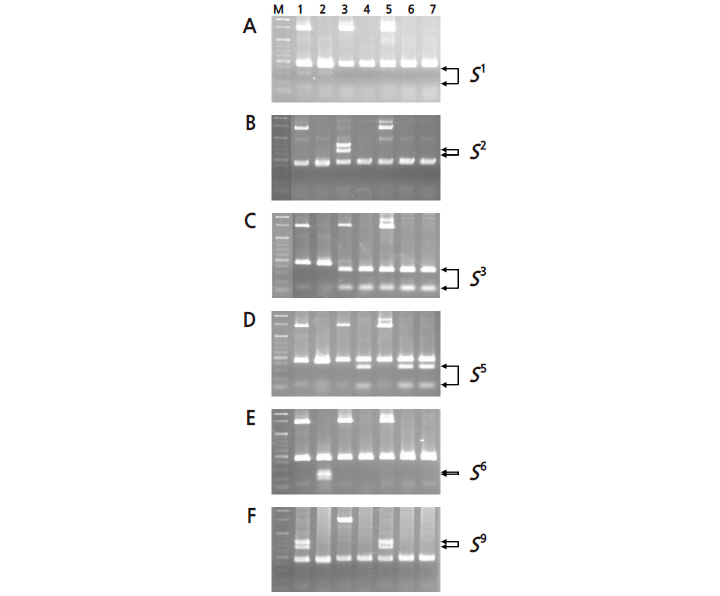
We identified the S-genotypes of the seven pear cultivars (Fig. 1) using a previously developed PCR-RFLP system (Fig. 1) (Kim et al., 2007). S-RNase PCR fragments (around 450 bp corresponding to S3 or S5RNase and 1,300 bp corresponding to S2 or S9RNase) were obtained from ‘Niitaka’ and the other pear cultivars. ‘Niitaka’ was digested with the S3 and S5 allele-specific restriction endonuclease, PpuMI, producing a 450-bp PCR fragment, but this was not sufficient to determine whether its corresponding S allele was S3 or S5. However, when the S5 allele-specific restriction endonuclease, AlwNI, was used, it was unable to digest the target allele, indicating that only the S3 allele was present in that locus in ‘Niitaka’ (lane 5; Fig. 1C, 1D, and 1F). The presence of the S3 allele was confirmed in ‘Chojuro’ (lane 3), ‘Shinchu’ (lane 4), ‘Housui’ (lane 6) and ‘162-29’ (lane 7) when digested with the S3 and S5 allele-specific restriction endonuclease, PpuMI (Castillo et al., 2001; Kim et al., 2002).
To determine the other S-allele, the 1,300-bp PCR fragments of ‘Niitaka’ were digested with the S2 and S9 allele-specific restriction endonucleases, XbaI and BstBI, respectively (Fig. 1B and 1F). The PCR fragments of ‘Niitaka’ (lane 5) were cleaved with the S9 allele-specific restriction endonuclease, but not with the S2 allele-specific enzyme, indicating that ‘Niitaka’ contained the S9 allele at that locus. The cultivar ‘Amanogawa’ (lane 1) was also confirmed to have the S9-RNase using the S9 allele-specific restriction endonuclease BstBI (Takasaki et al., 2004). Therefore, we concluded that the S-genotype of ‘Niitaka’ must be S3S9. The S-genotypes of the cultivars were determined as follows: ‘Amanogawa’ (S1S9 ), ‘Imamuraaki’ (S1S6 ), ‘Chojuro’ (S2S3 ), ‘Shinchu’ (S3S5 ), ‘Niitaka’ (S3S9 ), ‘Housui’ (S3S5 ) and ‘162-29’ (S3S5 ).
The S9 allele specific to the 1,300-bp fragment was detected in ‘Niitaka’ and ‘Amanogawa’, suggesting that the S9 allele in ‘Niitaka’ was derived from ‘Amanogawa’ (S1S9 ). The S3 allele was also detected in ‘Niitaka’; however, neither the S1 nor S6 allele of the previously reported male parent ‘Imamuraaki’ was found in ‘Niitaka’ (S3S9 ). Therefore, we concluded that ‘Imamuraaki’ is not a parent of ‘Niitaka’. Furthermore, our results suggest ‘Chojuro’ (S2S3 ) and ‘Shinchu’ (S3S5 ) as possible candidates for the parent that contributed the S3 allele of ‘Niitaka’.
The recent identification of the S locus F-box brothers (SFBB) in Japanese pear and apple suggested that these multiple F-box genes are pollen-specific candidate genes for the S haplotypes. These S haplotypes exhibit pollen-specific expression, polymorphisms and linkage to the S locus (Sassa et al., 2007). In Japanese pear, three SFBBs were identified from a single S haplotype and were found to be homologous to other haplotype genes of the same group (i.e., α-, β- and γ-groups; Kakui et al., 2007).
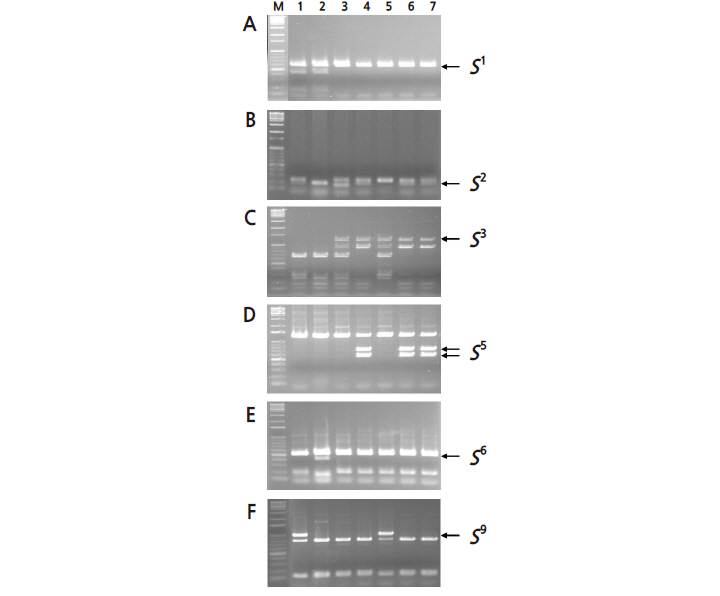
We also determined the S-genotypes of the seven pear cultivars using the PCR-CAPS/dCAPS method based on the PpSFBB-γ genes (Fig. 2A-2F). The 1,245-bp S3-PpSFBB-γ PCR fragments of ‘Niitaka’ (S3S9 ) (lane 5), ‘Shinchu’ (S3S5 ) (lane 4), ‘Housui’ (S3S5 ) (lane 6) and ‘162-29’ (S3S5 ) (lane 7) were not cleaved with HpyCH4IV (Fig. 2C). The PpSFBB-γ fragments of ‘Shinchu’ (lane 4), ‘Housui’ (lane 6) and ‘162-29’ (lane 7) were cleaved by the S5-specific restriction endonuclease AflII, but ‘Niitaka’ (lane 5) was not cleaved by this enzyme (Fig. 2D). The S9-PpSFBB-γ PCR fragments of ‘Niitaka’ (lane 5) and Amanogawa (S1S9 ) were cleaved by HaeIII (Fig. 2F). Based on these polymorphisms of the PpSFBB-γ genes, the S-genotype of ‘Niitaka’ was concluded to be S3S9 and the S-genotypes of ‘Shinchu’, ‘Housui’ and ‘162-29’ were concluded to be S3S5, which correlates to the results of the S-RNase analysis. Therefore, we suggest that PCR-CAPS/ dCAPS analysis of both PpSFBB-γ and S-RNase genes can be used to determine the S-genotypes of Japanese pear, identify cultivars with highly diverse S-genotypes and reveal obscure relationships between parents and progeny. However, the S-genotype alone cannot perfectly describe the parent-offspring relationships or be used to identify new potential parents.
SSR Analysis
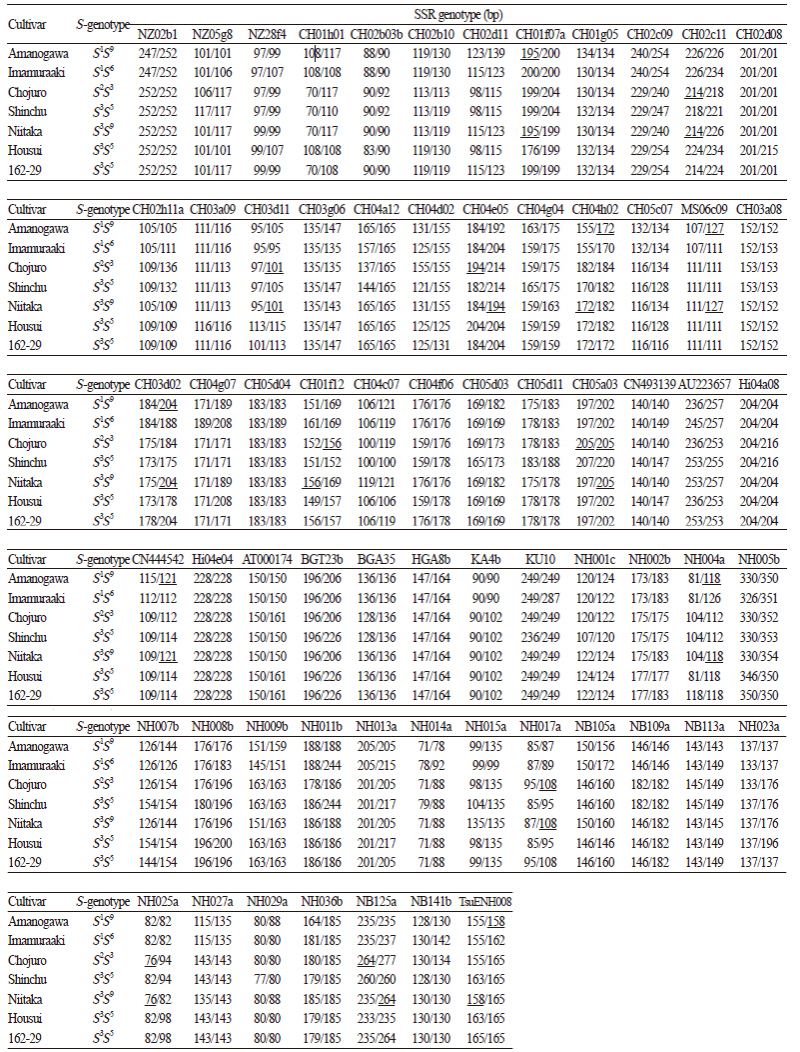
To determine the parentage of the pear cultivar ‘Niitaka’, 107 SSR markers were tested. Among them, 67 SSR markers contained one or two alleles represented by discrete fragments and 14 SSRs contained three or more alleles, whereaS26 SSRs did not amplify. The 67 polymorphic SSR markers were used to determine the parentage of ‘Niitaka’ (Table 1). Sixteen alleles from ten SSR markers, NZ28f4, CH01h01, CH01f07a, CH04h02, CH03d02, CN444542, NH004a, NH014a, NH015a and TsuENH008, amplified in the previously reported parent ‘Imamuraaki’ but were not found in ‘Niitaka’, indicating that ‘Imamauraaki’ is not the male parent of ‘Niitaka’. Based on similar discrepancies at seven SSR markers, the cultivar ‘Shinchu’ was also ruled out as a parent of ‘Niitaka’.
In humans, the probability of parentage is calculated based on the allele frequency (Hashiyada et al., 1997; Katsumata et al., 2001). According to the criteria for human parentage, an allele frequency of 0.9911 indicates an extremely high probability that an individual is a parent, whereas an allele frequency of 0.9999 shows a definitive parent-offspring relationship. In cultivated pears, it is very difficult to evaluate allele frequency, because it varies according to cultivar and species; however, the parent-offspring relationship of eight pear cultivars was determined with 15 to 20 SSR loci and the probability of parentage ranged from 0.9911 to 0.9999 (Kimura et al., 2003). Recently, Sawamura et al. (2008) analyzed the parent-offspring relationship of 55 Japanese pear cultivars using 18 SSR markers and determined that the reported parents of ten hybrid cultivars, including ‘Niitaka’, were incorrect due to discrepancies at three or more SSR loci.
In this study, we used 67 SSR markers, which should give a high level of confidence in parentage determination. Seven specific alleles of ‘Amanogawa’ were identified in ‘Niitaka’ by the SSR markers CH01f07a, CH04h02, MS06c09, CH03d02, CN444542, NH004a and TsuENH008. Eight specific alleles of ‘Chojuro’ were also found in ‘Niitaka’ using the SSR markers CH02c11, CH03d11, CH04e05, CH01f12, CH05a03, NH017a, NH025a and NB125a. Therefore, we conclude that ‘Niitaka’ was produced from a cross between ‘Amanogawa’ and ‘Chojuro’. Based on the SSR analysis, we also confirmed that ‘Niitaka’ and ‘Housui’ are the parents of ‘162-29’.