Introduction
Material and Methods
Plant Materials
Genomic DNA Isolation
S-haplotype Identification
Assessment of Genetic Diversity
Results and Discussion
S-allele Distribution of ‘Danji’ Radish
Genetic Diversity by RAPD
Introduction
‘Danji’ radish (Raphanus sativus L. var. hortensis Baker f. gigantissimus Makino, 2n = 2x = 18) is an annual plant in the Brassicaceae family, and is native to Jeju Island in Korea. ‘Danji’ radish had been used as vegetable before F1 radish varieties became commercialized, but it is no longer cultivated on a large scale by farmers. This plant, however, shows several advantageous characteristics. The original form of ‘Danji’ radish had unique root characters, such as a round shape and big size (as much as 10-15 kg), and was crispy enough to use in Kimchi. Different from F1 radish cultivars, it is extremely late flowering, with a strong vigor and a thick, dark green mesophyll. ‘Danji’ radish cultivars have been maintained by home seed pro-duc-tion by farmers. However, the present ‘Danji’ radish lost its own characteristics by pollination with the other pollens of F1 varieties. Thus, restoring the original character of ‘Danji’ radish has required breeding for the development of commercially useful varieties.
Like other species of Cruciferous vegetables (e.g. cabbage, broccoli, turnip), the ‘Danji’ radish is a cross-pollinating crop. The prevention of self-fertilization and promotion of outcrossing are mainly controlled by a genetic mechanism called self-incompatibility (SI) (Bateman, 1955). SI acts by inhibiting the germination of pollen on stigma, or by inhibiting the elongation of the pollen tube in the styles based on protein-protein interactions (Nasrallah et al., 1970). This mechanism is controlled by multiple interactions between many different alleles at a single locus (the S locus) (Nasrallah et al., 1991).
In breeding, SI is widely used for the production of F1 hybrids. The genetic mechanism of SI has been studied in several rapeseed cultivars. SI is controlled sporophytically by a single multi-allelic locus (the S locus). The S-locus consists of two linked genes expressed in the pistiland anther, the S locus glycoprotein (SLG) gene and the S receptor kinase (SRK), that are female determinants expressed in stigma papillae (Nasrallah et al., 1985; Rundle et al., 1993). The S locus cysteine-rich (SCR) gene andS locus protein 11 (SP11) are male determinants expressed in the anthers (Kakayama et al., 2001; Watanabe et al., 2000). In a SI reaction, the S domain of the membrane-associated SRK forms a complex with the pollen-born SCR/SP11 protein. Formation of this complex leads to activation of the kinase domain of SRK, inducing a signal transduction cascade involving phosphorylation events, that finally lead to rejection of the pollen and preventing fertilization. Based on sequence polymorphisms, SLG alleles are divided into two classes (Nasrallah et al., 1991). The class-I haplotypes are dominant and have strong self-incompatibility, whereas the class-II haplotypes are considered recessive and display a weak self-incompatible phenotypic effect (Nasrallah and Nasrallah, 1993). However, when a female determinant interacts with a male determinant of a different haplotype, no SI is created.
For the identification and classification of S haplotypes, the PCR-RFLP method was developed (based on DNA sequence variation in the SLG and SKR gene) and successfully used in Brassica and Raphanus (Lim et al., 2002). Later, these S-allele-specific PCR-RFLPs were converted to Sequence characterized amplified region (SCAR) markers that provide easier assessment of various haplotypes of the SLG and SKR genes. Random amplified polymorphic DNAs (RAPDs) can be extremely useful for marker-assisted selection (MAS) of important traits and DNA fingerprinting for genetic diversity assessment. DNA fingerprinting has been widely used to group germplasm with similar genetic constitution and identify parental combinations that could potentially create maximal genotypic variability in their progenies to achieve advances in selection. Genotypic differences can also be used for cultivar identification, germplasm management, and protection of the plant breeder’s intellectual property rights (Lombard et al., 2000).
Despite its potential for revealing new sources of radish cultivar and breeding needs, no study has reported the identification and allele frequency of S-haplotypes in the ‘Danji’ radish population of Jeju Island. In the present study, we identify S-haplotypes of ‘Danji’ radish conserved on Jeju Island through germplasm collection and characterize its genetic diversity by using S-allele specific SCAR and RAPD markers.
Materials and Methods
Plant Materials
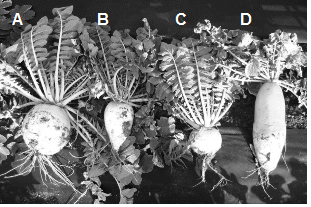
Plant materials include 47 accessions of ‘Danji’ radish collected on Jeju Island (and used as breeding materials) and one commercial radish cultivar, ‘Gwandong’ (Fig. 1).
|
Fig. 1. Phenotypes of different accessions of the ‘Danji’ radish (A, B, and C; Raphanus sativus L. var. hortensis) and ‘Gwandong’ radish radish (D; Raphanus sativus L. var. sativus). |
Genomic DNA Isolation
Leaf tissue was ground in a 1.5 mL micro-centrifuge tube using steel beads and 600 µL of DNA extraction buffer with a Tissue Lyser (QIAGEN, Venlo, Netherlands). The tube was incubated at 65°C for approximately 45 min, 200 µL of 7.5 M ammonium acetate added, and then the tube was placed on ice for 15-20 min. The lysate was centrifuged at 14,240 ×g for 10 min, and the supernatant transferred to a new 1.5 mL tube containing 5 µL of glycogen solution (5 mg・mL-1) and 600 µL of isopropanol. After centrifugation at 14,240 ×g for 10 min, the liquid was carefully decanted, and the DNA pellet washed with 300 µL of 70% ethanol. The washed pellet was dried and resuspended in 200 µL of 0.1 M Tris. DNA quality and concentration were evaluated with a Nanodrop 1000 Spectro-photometer (Thermo Scientific, Pittsburgh, PA, U.S.A.).
S-haplotype Identification
For the identification of S-haplotypes, PCR primer sequence information was obtained from the National Center for Biotechnology Information (NCBI) database and primers synthesized by Bioneer Inc. (Daejeon, Korea). PCR was performed to amplify S allele-specific genes with the following cycling profile: 1 cycle of 15 min at 95°C for preheating, 35 cycles of 20 s at 95°C for denaturation, 40 s at an appropriate temperature for each primer (Table 1) for annealing, 1 min at 72°C for extension, and 1 cycle of 5 min at 72°C for an final extension step. PCR products were separated on a 1.5% agarose gel containing TBE at 120 V for 2 h, and visualized under UV light after staining with ethidium bromide.
For DNA fingerprinting of ‘Danji’ radish cultivars, 40 10-mer RAPD primers were randomly chosen and tested for RAPD. PCR amplification for RAPD evaluation was performed in a total volume of 20 μL containing 40 ng of genomic DNA, each 10-mer primer at 1.25 μM, 2.5 mM MgCl2, 1X PCR buffer, 0.2 mM dNTPs, and 1U of Taq polymerase (Solgent, Daejeon, Korea) with the following cycling profile: 1 cycle of 5 min at 95°C, 35 cycles of 30 s at 95°C, 30 s at 40°C, and 2 min at 72°C, and 1 cycle of 3 min at 72°C for an extension step. PCR products were separated on a 1.0% agarose gel containing TBE at 80 V for 1 h., and visualized under UV light after staining with ethidium bromide.
Assessment of Genetic Diversity
The polymorphic and monomorphic fragments were counted across all 48 accessions for each SCAR and RAPD primer. Only clearly readable bands with strong intensity were manually scored and included in the binary data matrix (i.e., 1 denoted a band was present, 0 denoted absence). All subsequent data analysis was performed using NTSYS-pc version 2.02k. Pair-wise genetic similarity coefficients among the cultivars were calculated based on Nei-Li (Nei and Li 1979) and the Nei-Li similarity coefficient was calculated as 2a/(n-d), where a is the number of loci for which the bands is present, d is the number of loci for which the band is absent, and n is the total number of loci. Dendrograms were constructed using the unweighted pair-group method with arithmetic averaging (UPGMA) clustering analysis of the genetic similarity coefficient matrices.
Results and Discussion
S-allele Distribution of ‘Danji’ Radish
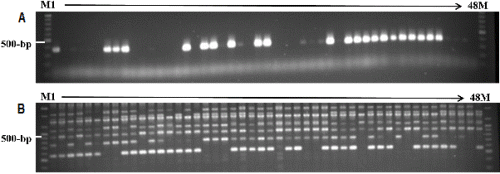
The S-allele classification and distribution of ‘Danji’ radish accessions originating from Jeju Island were investigated with SCAR markers (Table 2 and Fig. 2). Among 22 S- haplotype-specific SCARs, only six primer sets amplified locus-specific PCR fragments from at least one accession. These primer sets were specific to S1, S2, S5, S10, and S21 haplotypes for the SLG locus and S23 for the SRK locus. All other primer sets failed PCR amplification, possibly due to nucleotide sequence variation in primer binding sites or the lack of any corresponding S allele in the genome.
For the SLG locus, the frequency of the class-II haplotype was 72%, higher than the frequency of class-I. S5 and S21 were the most frequently distributed S-haplotypes and detected from 87.5% and 64.6% of the accessions investi-gated, respectively. Accessions containing both of these alleles accounted for 64.5%. S23 for SRK was detected from only seven cultivars and all cultivars also contained SLG alleles.
Grouping of the accessions based on the S-allele com-position revealed three major groups (Group A-1, A-2, and A-3) consisting of 29 (Group A-1), 13 (Group A-2), and 6 (Group A-3) cultivars. Eight cultivars showed unique allelic compositions and could be distinguished from other cultivars.
Lim et al. (2006) reported the classification of S-haplotypes in commercial F1 hybrid radish cultivars. Among 44 cultivars, almost half of the S-haplotypes in F1 hybrids contain the class-II haplotype. A high frequency of class-II haplotypes was also detected in ‘Danji’ radishes, indicating a close genetic relationship between these two species. It also indicates gene introduction by natural cross-pollination with radish F1 cultivars in which the class-II haplotypes of inbreds were frequently used as the parents of F1 hybrid cultivars.
Although conventional test crosses need to be performed for the determination of S haplotypes in ‘Danji’ radishes, the results of this study can be useful for predicting the possibility of self-incompatibility and determining proper cross combinations for improving genetic diversity.
Genetic Diversity by RAPD
The genetic diversity of 47 ‘Danji ‘radishes and 1 ‘Gwandong’ radish were evaluated by using 38 RAPD primers (Table 3 and Fig. 2). Forty-eight accessions were genotyped with 38 randomly chosen RAPD primers. A total of 312 bands were scored, with 138 bands (44.2%) mono-morphic among the 60 cultivars, whereas 174 (55.8%) polymorphic bands were identified. For each primer, an average of 8.2 bands was scored and an average of 45.7 polymorphic bands was detected. Polymorphism rates ranged from 0.2 to 1.0, indicating significant variations in detecting polymorphisms across the RAPD primers (Table 3).
Differences among the accessions at the DNA level were determined by comparing the genetic similarity coefficients from the pair-wise comparisons. The genetic similarity coefficients among all pairs of 48accessions varied from 0.62 to 0.93, and 42% of the comparisons exhibited values higher than 0.85. All cultivars could be distinguished based on the DNA fingerprints revealed by RAPD, indicating that RAPD profiles revealed in this study can be useful for cultivar identification.
In the study of three radish varieties (R. sativus var. sativus, var. hortenisis Becker, var. niger Kerner) in China using the amplified fragment length polymorphisms (AFLPs) and RAPDs, the most abundant genetic diversity was revealed among var. hortenisis Becker (Kong et al., 2011). Substantial levels of genetic variability were also detected in the germplasm of cultivated radish by analyzing AFLPs and inter-simple sequence repeats (ISSRs) (Muminovic et al., 2005). The wide genetic background harbored in ‘Danji’ radish can be exploited for genetic improvements of the radish germplasm. In addition, the data for S-haplotypes and genetic relationships assessed in this study will be greatly useful for deciding cross combinations to recover the unique traits of ‘Danji’ radish and extend its genetic diversity.
To evaluate how DNA variations at the RAPD level exist within the accessions carrying the same S-haplotype, den-drograms constructed based on RAPD and S-haplotype primers were compared (Fig. 3). When the accessions carrying S5 and S21 (Group A-1) were considered, these accessions tended to separate into two major groups (Group B-1and Group B-3). A majority of accessions harboring S2 and S5 (Group B- 2), tended to be closely related and located in GroupB-2. Four out of seven accessions harboring S23 of the SRK gene belonged to Group B-1. Accessions harboring S-haplotype with a low frequency (47, 48, and 50) were distantly related with other accessions as well in the RAPD dendrogram. In general, the comparison between the two dendrograms indicates that despite the detection of unrelated and sporadic distribution for some accessions, there was a tendency for the accessions carrying the same S-allelic composition to group into the same cluster. This suggests that the accessions of the same S-haplotype may share a similar genetic background and a common parentage.