Introduction
Materials and Methods
Plant Materials
Primer Design for the Fluidigm SNP Type Assays
DNA Extraction
Specific Target Amplification
SNP Type Assay
Scoring of SNPs
Analysis of HRM Markers
Results and Discussion
Development of the SNP Type Assays
Application of SNP Type Assays to Pepper Cultivars and Breeding Lines
서언
Chili pepper (Capsicum annuum L.) is one of the most important vegetable crops in Korea (Lee et al., 2004). However, annual production and cultivation has gradually declined due, in part, to pepper diseases that usually occur in the summer season.
Major pepper diseases include Phytophthora root rot (Phytophthora capsici; Liu et al., 2014), anthracnose (Colletotrichum scovillei and C. truncatum, formerly C. acutatum and C. capsici, respectively;Mahasuk et al., 2016), powdery mildew (Leveillula taurica; Lefebvre et al., 2003), bacterial wilt (Ralstonia solanacearum; Mimura et al., 2009), bacterial spot (Xanthomonas campestris pv. vesicatora; Truong et al., 2011), Cucumber mosaic virus (CMV; Eun et al., 2016), Pepper mild mottle virus (PMMoV; Yang et al., 2012), Tomato spotted wilt virus (TSWV; Kim et al., 2017), and Pepper mottle virus (PepMoV; Kim et al., 2011, 2017). These diseases are difficult to control, even with the use of agrichemicals. Therefore, pepper varieties with multiple resistance are highly desired. In addition, higher levels of pungency are also desired by the pepper processing industry (Lee et al., 2016a). The pungency of chili pepper is related to the content of capsaicinoids such as capsaicin and dihydrocapsaicin (Aza-González et al., 2011).
Molecular markers are widely used to improve the efficiency of plant breeding programs, to construct genetic linkage maps, and to detect genes or quantitative trait loci (QTL) controlling specific traits (Collard et al., 2005; Collard and Mackill, 2008; Xu and Crouch, 2008). Marker-assisted selection (MAS) and marker-assisted backcrossing (MABC) are commonly used to improve the efficiency of selection in plant breeding. MAS involves foreground selection of a target trait or traits of interest, while MABC involves background selection of overall genomic regions in BC generations (Collard and Mackill, 2008). MAS and MABC can reduce the breeding period and the number of generations needed in a breeding program compared to conventional phenotypic selection because codominant markers allow breeders to distinguish between homozygotes and heterozygotes and to detect early the desired traits in seeds or seedlings without having to grow the plants to maturity or inoculate them.
Several molecular markers have been developed in pepper for disease resistance and capsaicinoids content. Two dominant markers, OpD04-717-SCAR and P5-SNAP, for the detection of a major QTL, Phyto.5.2, for resistance to P. capsici have been reported (Quirin et al., 2005), and codominant markers M3-CAPS and Phyto5NBS1-HRM for the trait have also been developed (Lee et al., 2012; Liu et al., 2014). The markers CaR12.2M1-CAPS and CcR9M1-SCAR have been linked to the major QTLs for resistance to C. scovillei and C. truncatum, respectively (Lee et al., 2010, 2011). Bacterial spot resistance genes (Bs2 and Bs3) have been cloned in pepper (Tai et al., 1999b; Römer et al., 2007), and gene-based codominant markers 14F/14R and 25-1 for Bs2 and PR-Bs3 for Bs3 were subsequently developed (Römer et al., 2010; Truong et al., 2011). Three SNP markers, CaTmint3HRM, CaT1616BAC, and 240H02sp6, were identified and linked to a single dominant gene, Cmr1, that controls CMV resistance (Kang et al., 2010). Two CAPS markers, pvr1-R1 and pvr1-R2, were developed to detect pvr1 and pvr12 alleles for potyvirus resistance in C. chinense accessions (Yeam et al., 2005). Pvr4 and Tsw genes have also been cloned: Pvr4 is a potyvirus resistance gene originating from C. annuum ‘CM334’ and Tsw is a TSWV resistance gene from C. chinense accessions ‘PI159236’ and ‘PI152225’(Kim et al., 2017). Markers 61786 and NB575m were found to cosegregate with the Pvr4 and Tsw genes, respectively. In addition, high-resolution DNA melting (HRM) markers linked to the QTLs responsible for high capsaicin and dihydrocapsaicin content in C. chinense ‘Bhut Jolokia’ have been developed (Lee et al., 2016a). Many other trait-linked markers have been reported in pepper. Therefore, high-throughput screening methods are needed for the simplified and costeffective analysis of multiple molecular markers in a single reaction.
Remarkable technological achievements have occurred over the past few decades in the field of DNA sequencing and SNP genotyping, including next-generation sequencing (NGS) and high-throughput SNP genotyping (Varshney et al., 2009; Kumar et al., 2012; Poland and Rife, 2012; Thomson, 2014). High-throughput SNP genotyping is particularly useful in crop breeding (Thomson, 2014). Molecular markers can be rapidly developed for SNPs, which are the most abundant polymorphisms with unlimited nucleotide variations between individual organisms, even within the same species (Rafalski, 2002). Moreover, analysis of SNP markers is accurate, rapid, and inexpensive.
Several high-throughput SNP genotyping platforms have been reported, including: Illumina Infinium iSelect HD array, Affymetrix Axiom array, Douglas Array Tape, Fluidigm dynamic arrays, restriction-enzyme-based genotyping-by-sequencing (GBS), and amplicon sequencing (Thomson, 2014). Among them, Fluidigm dynamic arrays adopt a flexible, PCR-based SNP platform using a nanofluidic integrated fluid circuit (IFC; Wang et al., 2009). There are three different types of Fluidigm dynamic arrays: a 48.48 dynamic array, which yields 2,304 data points with 48 samples and 48 markers, as well as 96.96 and 192.24 dynamic arrays, which yield 9,216 and 4,608 data points, respectively (Wang et al., 2009; Thomson, 2014). This system can save both resources and time by reducing the reaction volumes to 7-10 nL and producing 2,304-9,216 data points within 2-4 hours (Thomson, 2014), while 384- and 96-well PCR systems require 5-20 μL of the reaction volumes per well and it takes 4-8 days to produce the same number of data points with the PCR systems.
In this study, we developed SNP type assays linked to disease resistance or capsaicinoids content for use in Fluidigm 192.24 dynamic arrays that could simultaneously analyze 24 SNP markers. The CAPS, SCAR, and SNP markers previously developed in chili pepper were successfully converted into the Fluidigm SNP type assays.
Materials and Methods
Plant Materials
Three populations (PG, CHB-F3, and JN-F5) were used to develop the SNP type assays. The PG population consisted of 51 Capsicum accessions including six species (C. annuum, C. baccatum, C. chinense, C. frutescens, C. pubescens, and C. chacoense), which were obtained from the National Agrobiodiversity Center, Rural Development Administration, Republic of Korea (Table 2; http://genebank.rda.go.kr/). The CHB-F3 population was made up of 48 individuals originating from a cross of C. annuum ‘A1’ × C. annuum ‘2602’, which was used for QTL analysis of CMVP1 resistance (Table 3; Eun et al., 2016). The JN-F5 population consisted of 50 individuals of an F5 single seed descent (SSD) population derived from a cross of C. annuum ‘NB1’ × C. chinense ‘Bhut Jolokia’, which was used for QTL analysis of capsaicinoids content (Table 4; Lee et al., 2016a). In addition, 33 commercial cultivars, 30 breeding lines, and 7 genetic sources for pepper were used for validation of the newly developed SNP type assays (Tables 5 and 6).
Primer Design for the Fluidigm SNP Type Assays
The following target sequence criteria were employed to design primers for the SNP type assays. Length of the target sequences: a minimum of 60 bp (including both upstream and downstream of the target SNP site) and a maximum of 250 bp. For SNPs, only one SNP was present in the target sequence. For insertions/deletions (In/Dels), the length of the In/Del was shorter than 10 bp. The G/C content of the target sequence was < 65%. A total of 43 primers were designed using D3 Assay Design (https://d3.fluidigm.com/; Fluidigm, South San Francisco, CA, USA). Primer information is listed in Table 1. Each assay consisted of three types of primers: a specific target amplification (STA) primer, a locus-specific (LS) primer, and an allelespecific (AS) primer (Wang et al., 2009).
DNA Extraction
Genomic DNA was prepared from fresh leaves using the miniprep method described by Eun et al. (2016). The DNA concentration was measured using a BioDrop μLITE (BioDrop UK Ltd., Cambridge, UK) and adjusted to 50 ng·μL-1. The DNA was used for the SNP type assays and HRM analysis.
Specific Target Amplification
Before performing the SNP type assay, specific target amplification (STA), which is used to enrich the amplicon including the targeted SNP sequences, was performed to increase the probability of success of the SNP type assay (Wang et al., 2009). First, a 10× STA primer pool was prepared comprising a mixture of 2 μL of STA primer for each of the 24 markers, 2 μL of LS primer for each of the 24 markers, and 304 μL of DNA suspension buffer (Teknova, Holister, CA, USA). For each of the 191 samples, STA was executed using a LightCycler 96 Real-Time PCR (Roche, Basel, Switzerland) in a total volume of 5 μL per reaction, which contained 2.5 μL of master mix (Qiagen, Hilden, Germany), 0.5 μL of the 10× STA primer pool, 0.75 μL of PCR-certified water, and 1.25 μL of genomic DNA with the following PCR profile: pre-denaturation for 900 s at 95°C followed by 14 cycles of a 2-step amplification of 15 s at 95°C and 240 s at 60°C. Then, 3 μL of amplified product was diluted in 97 μL of PCR-certified water and then used for the SNP type assay.
SNP Type Assay
To perform the SNP type assays using the 192.24 IFC, the assay mix and sample mix were prepared. The assay mix contained 1.2 μL of PCR-certified water, 2 μL of 2× assay loading reagent, and 0.8 μL of the assay pre-mix, which was comprised of 3 μL of each AS primer, 8 μL of each LS primer, and 29 μL of DNA suspension buffer (Teknova, Holister, CA, USA). The sample pre-mix contained 540 μL of 2× Fast Probe Master Mix (Biotium, Fremont, CA, USA), 54 μL of SNP type 20× sample loading reagent, 18 μL of SNP type 60× reagent, 6.48 μL of 50× ROX dye (Invitrogen, Waltham, MA, USA), and 11.52 μL of PCRcertified water. Subsequently, the sample mix was prepared by mixing 1.9 μL of each STA product and 2.6 μL of the sample pre-mix in each well of two 96-well plates. Finally, 3 μL of each sample mix and 3 μL of each assay mix were loaded into 192 sample inlets and 24 assay inlets of the 192.24 IFC, respectively.
The SNP type assays were performed in series using three machines, the IFC controller RX (Fluidigm, South San Francisco, CA, USA), the IFC cycler (Fluidigm, South San Francisco, CA, USA), and the EP1 system (Fluidigm, South San Francisco, CA, USA) according to the manufacturer’s instructions (Wang et al., 2009).
Scoring of SNPs
In each SNP type assay, two types of fluorescence, FAM (red, Y axis) and HEX (green, X axis), were analyzed and each fluorescence was linked to each SNP (Table 1). Using Fluidigm SNP genotyping analysis version 4.1.3 (Fluidigm, South San Francisco, CA, USA), three different genotypes (A, H, and B) were identified: A and B refer to a specific homozygous SNP; H refers to a heterozygous SNP (Fig. 1).
Analysis of HRM Markers
HRM analysis was performed using a LightCycler 96 Real-Time PCR machine (Roche, Basel, Switzerland). The reaction solution for HRM analysis was prepared and PCR reactions were performed according to Lee et al. (2016b). High-Resolution Melt software version 1.1 (Roche, Basel, Switzerland) was used to analyze the marker types of HRM. HRM marker information was derived from the references in Table 1.
Results and Discussion
Development of the SNP Type Assays
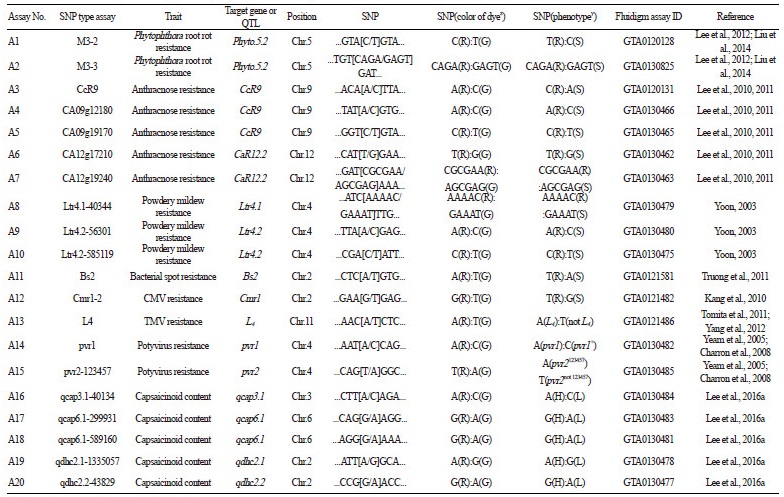
A total of 43 primer sets were designed for the SNP type assays and 20 assays were clearly analyzed via classification into three or two groups (Fig. 1). In Fig. 1, red, green, and blue points indicate XX (fluorescence of only FAM dye), YY (only HEX dye), and XY (both FAM and HEX dyes) marker types, respectively. The marker type of each SNP type assay was divided into resistant (R), heterozygous (H), or susceptible (S) types for disease resistance, and high (H), heterozygous (M), or low (L) types for capsaicinoids content (Fig. 1and Table 1). The 20 successful SNP type assays are listed in Table 1, which includes the name of the original genes or QTLs for capsaicinoids content (qcap3.1, qcap6.1, qdhc2.1, and qdhc2.2) and for resistance to diseases such as Phytophthora root rot (Phyto.5.2), anthracnose (CcR9 and CaR12.2), powdery mildew (Ltr4.1 and Ltr4.2), bacterial spot (Bs2), CMV (Cmr1), TMV (L4), and potyvirus (pvr1 and pvr2). A total of 5 192.24 ICFs were analyzed using the 43 primer sets and the three segregating populations (Tables 2, 3, and 4). The 20 successful SNP type assays showed clear results that were polymorphic in at least one population (Fig. 1and Table 1). In addition, 11 assays were compared with the corresponding original HRM markers to determine whether the marker types cosegregated (Tables 2, 3, and 4).
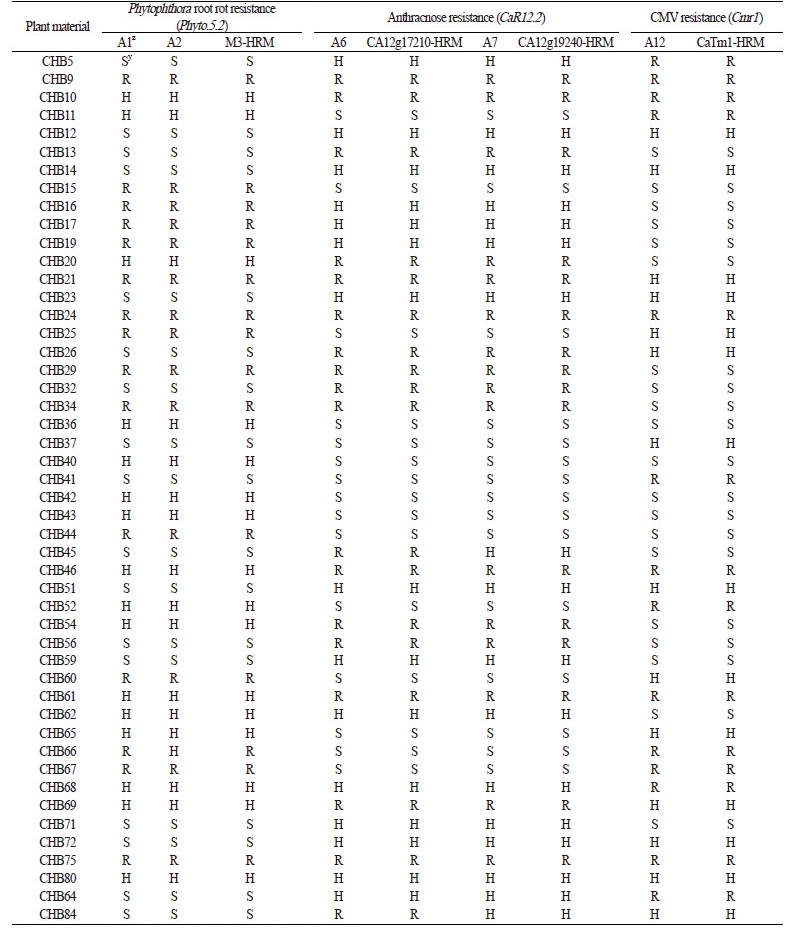
Two SNP type assays, M3-2 (Fig. 1A) and M3-3 (Fig. 1B), were converted from the M3-CAPS marker tightly linked to Phyto.5.2, a major QTL for resistance to root rot caused by P. capsici (Table 1, A1, and A2; Quirin et al., 2005; Lee et al., 2012). These assays revealed that the resistance allele was widely distributed among domesticated Capsicum species except for C. annuum (Table 2, A1 and A2). Thus, the resistance of the five domesticated species should be analyzed via inoculation with P. capsici because the resistance allele originated from C. annuum accessions, including ‘CM334’, ‘PI201232’, ‘PI201234’, and ‘AC2258’ (Liu et al., 2014). The marker types for the M3-2 assay were perfectly matched with those of the previously developed M3-HRM marker in two segregating populations, CHB-F3 and JN-F5 (Tables 3 and 4, A1), while the M3-3 assay revealed only one recombinant in the CHB-F3 population (Tables 3 and 4, A2). Therefore, the M3-2 assay can be used to select Phytophthora root rot resistance because it cosegregated perfectly with the M3-HRM marker.
Three assays, CcR9 (Fig. 1C), CA09g12180 (Fig. 1D), and CA09g19170 (Fig. 1E), were derived from the CcR9M1-SCAR marker to detect CcR9, a major QTL for resistance to anthracnose caused by C. truncatum (Table 1, A3, A4, and A5), while two assays, CA12g17210 (Fig. 1F) and CA12g19240 (Fig. 1G), originated from the CaR12.2M1-CAPS marker to select CaR12.2, a major QTL for resistance to anthracnose caused by C. scovillei (Table 1, A6 and A7; Lee et al., 2011). These assays showed that resistance alleles only appeared in C. baccatum and C. chacoense (Table 2, A3-A5). Resistance was already known to be present in C. baccatum, including ‘PBC80’, ‘PBC81’, ‘Cbp’, and ‘PI594137’(Park et al., 2009). However, the resistance of C. chacoense has not been analyzed with inoculation assays, and thus needs to be examined. The marker types of the CA12g17210 and CA12g19240 assays perfectly coincided with those of the CA12g17210-HRM and CA12g19240-HRM markers, respectively, in the CHB-F3 and JN-F5 populations (Table 2, A6; Table 3, A6 and A7).
Fig. 1.
Scatter plots of 20 SNP type assays. R, resistant; S, susceptible; H, heterozygous for A-O, high capsaicinoid content allele for P-T; L, low capsaicinoid content allele.
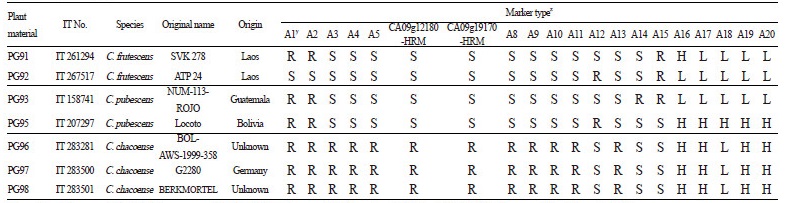
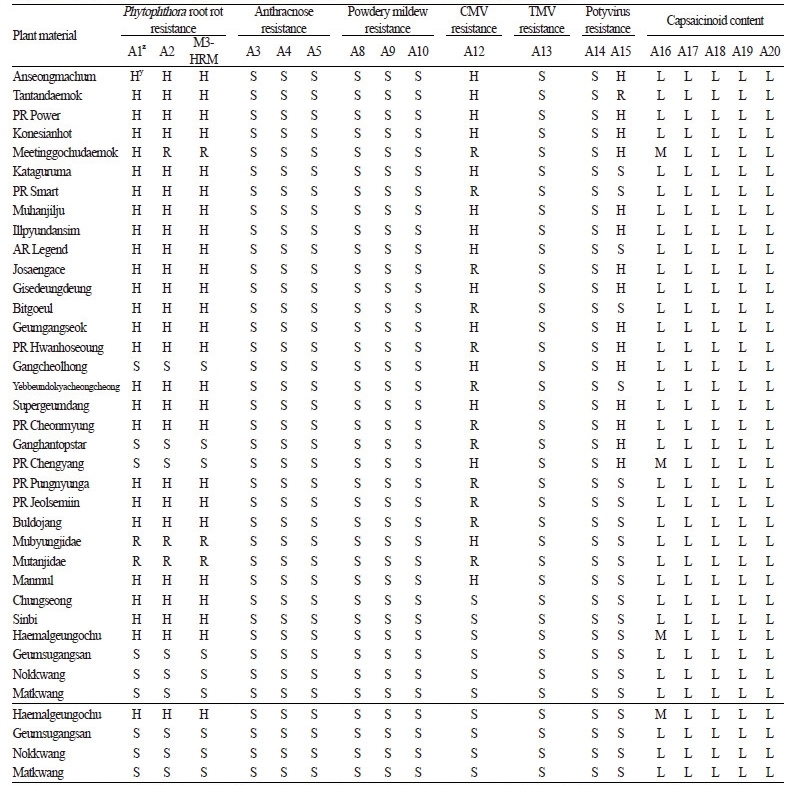
Table 2. Marker types of 18 SNP type assays and two HRM markers in 51 pepper accessions including six Capsicum species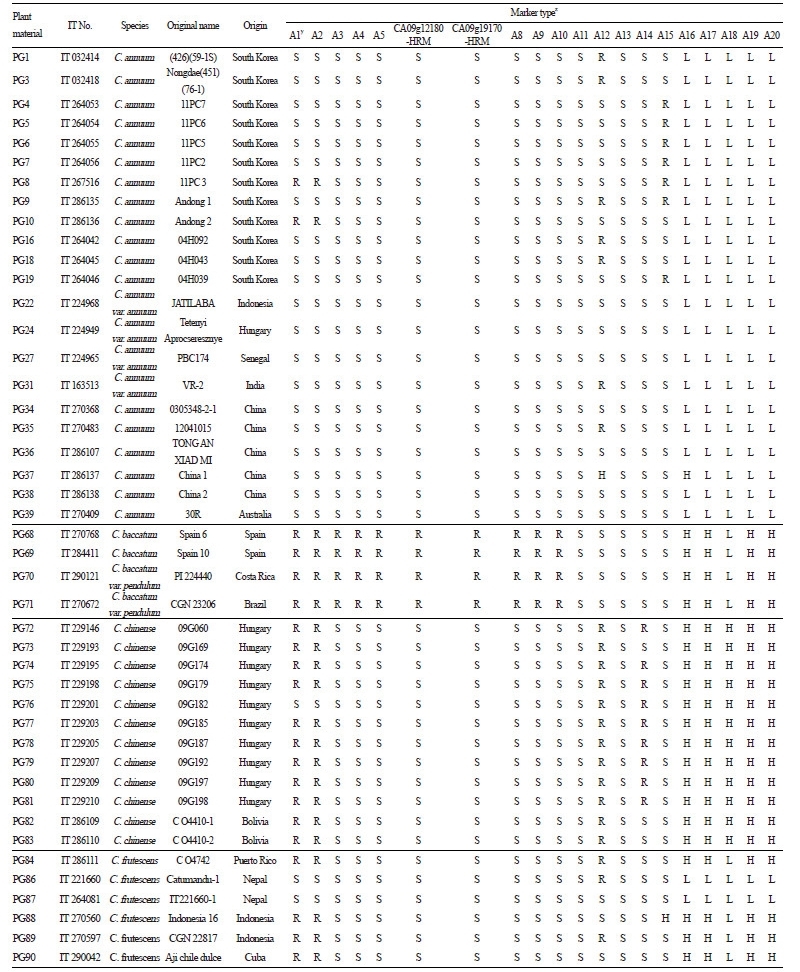 |
Three assays, Ltr4.1-40344 (Fig. 1H), Ltr4.2-56301 (Fig. 1I), and Ltr4.2-585119 (Fig. 1J), were developed to identify two QTLs, Ltr4.1 and Ltr4.2, for powdery mildew resistance, which were derived from C. baccatum ‘PBC81’(Table 1, A8, A9, and A10; Yoon, 2003). Like the anthracnose resistance assays, the resistance marker types of these assays were also specific to accessions of C. baccatum and C. chacoense (Table 2, A8-A10).
The Bs2 (Fig. 1K) assay was based on the Bs2 gene conferring resistance to most Xanthomonas campestris pv vesicatora races including 0, 1, 2, 3, 7, and 8 (Table 1, A11; Stall et al., 2009; Truong et al., 2011). The resistance marker type of this assay was found only in C. chacoense accessions (Table 2, A11). This result was supported by Tai et al. (1999a) who described Bs2 resistance from a wild species of pepper, C. chacoense ‘PI260435’.
The Cmr1-2 (Fig. 1L) assay was derived from the CaTm-int3-HRM marker closely linked to Cmr1, a dominant resistance gene against CMVKorean and CMVFNY strains (Table 1, A12; Kang et al., 2010). The resistance marker type of this assay was widely distributed throughout Capsicum spp. except for C. baccatum and C. chacoense (Table 2, A12), and its segregation pattern was the same as that of the CaTm1-HRM marker in the CHB-F3 and JN-F5populations (Tables 3 and 4, A12). Therefore, this assay can be used to select the Cmr1 gene.
The L4 (Fig. 1M) assay was developed to select L4 alleles resistant to most pathotypes including ToMV (P0), PaMMV (P0) and P1), and PMMoV (P1.2 and P1.2,3) (Table 1, A13; Tomita et al., 2011). Similar to the Bs2 gene, the resistance marker type of the L4 assay was present only in C. chacoense accessions (Table 2, A13). This result was consistent with the findings of Boukema (1984) who reported that L4 resistance was found in C. chacoense accessions ‘PI260429’ and ‘SA185’.
Two assays, pvr1 (Fig. 1N) and pvr2-123457 (Fig. 1O), were created to detect pvr1 and pvr2 genes, respectively, which were allelic and encoded an eIF4E protein, but originated from different genetic resources: pvr1 from C. chinense and pvr2 from C. annuum and C. frutescens (Table 1, A14 and A15; Kang et al., 2005; Charron et al., 2008). A resistant marker type of the pvr1 assay was mainly distributed in C. chinense accessions (Table 2, A14), while that of the pvr2-123457 assay was found in C. annuum and C. frutescens accessions (Table 2, A15). The pvr2-123457 assay was based on a specific SNP of potyvirus-resistance genes, pvr21, pvr22, pvr23, pvr24, pvr25, and pvr27 (Charron et al., 2008).
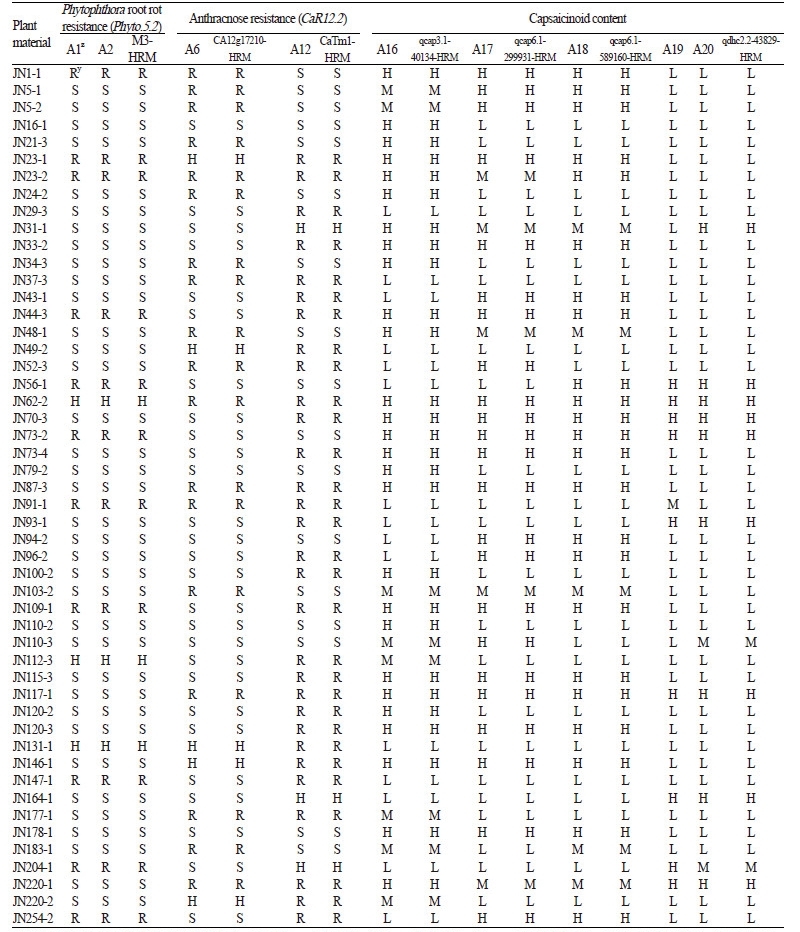
Five assays, qcap3.1-40134 (Fig. 1P), qcap6.1-299931 (Fig. 1Q), qcap6.1-589160 (Fig. 1R), qdhc2.1-1335057 (Fig. 1S), and qdhc2.2-43829 (Fig. 1T), were linked to four QTLs, qcap3.1 and qcap6.1 for capsaicin content and qdhc2.1 and qdhc2.2 for dihydrocapsaicin content, which were identified in C. chinense ‘Bhut Jolokia’(Table 1, A16-A20; Lee et al., 2016a). The qcap6.1- 589160 assay was specific to only C. chinense accessions, while the other four assays were widely polymorphic in five Capsicum species, with the exception of C. annuum (Table 2, A16-A20). This result implied that the QTL qcap6.1 might have a greater effect on capsaicinoid content than the other three QTLs. In addition, linkage analysis in the JN-F5 population showed that the four assays cosegregated with corresponding HRM markers (Table 4, A16, A17, A18, and A20). This result suggests that the assays can be used to detect corresponding QTLs.
Application of SNP Type Assays to Pepper Cultivars and Breeding Lines
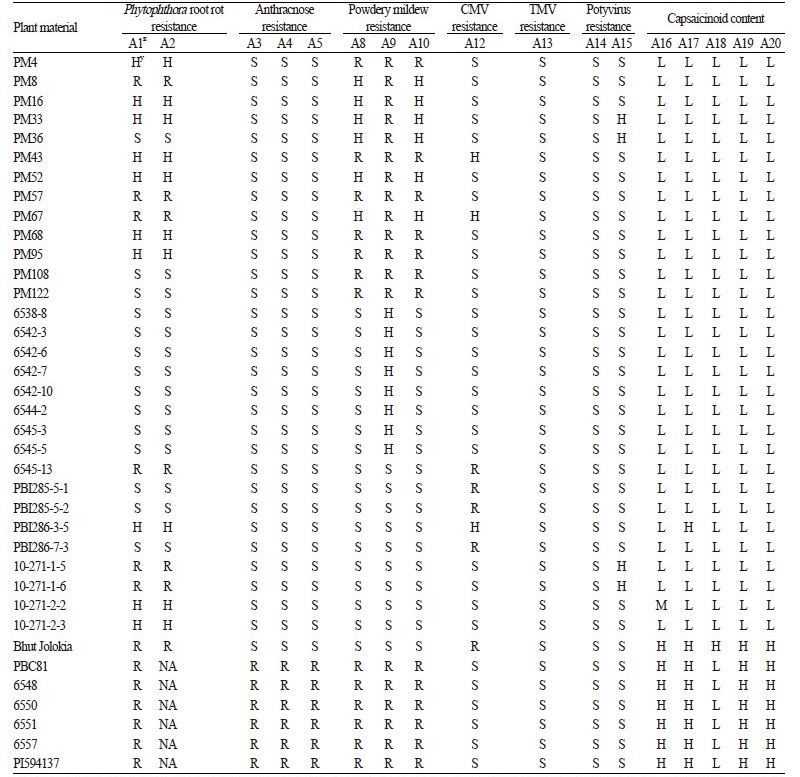
Successful SNP type assays were performed on 33 commercial cultivars, 30 breeding lines, and 7 genetic sources of chili pepper (Tables 5 and 6). Only five assays, M3-2 (A1), M3-3 (A2), Cmr1-2 (A12), pvr2-123457 (A15), and qcap3.1-40134 (A16), exhibited polymorphic results for the 33 commercial cultivars (Table 5). All cultivars except for ‘Geumsugangsan’,‘Nokkwang’, and ‘Matkwang’ were Phytophthora-resistant varieties. The results of M3-2 and M3-3 assays were consistent with the resistance phenotypes except for those of ‘Gangcheolhong’, ‘Ganghantopstar’, and ‘PR Chengyang’, which might have developed with another resistance gene(s) unrelated to the Phyto.5.2 gene because the M3-CAPS marker was perfectly matched with the resistance in the C. annuum resources including ‘AC2258’, ‘CM331’, ‘CM334-INRA’, ‘CM334-KBU’, ‘PBC602’, ‘YCM334’, ‘PI201234’, ‘PBC280’, and ‘PBC495’(Table 5, A1 and A2; Lee et al. 2012). A comparison of the assays with M3-HRM markers showed that the M3-3 assay was more accurate than the M3-2 assay, which had one recombinant (‘Meetinggochudaemok’). The other three polymorphic assays, Cmr1-2, pvr1-123457, and qcap3.1-40134, indicated that cultivars with the resistance marker type might have CMV and potyvirus resistance and that cultivars having the high capsaicinoid content marker type might be more pungent (Table 5, A12, A15, and A16). In an analysis of pepper breeding lines, 9 assays demonstrated polymorphic results among the cultivars (Table 6). Resistance marker types of three assays, Ltr4.1-40344 (A8), Ltr4.2-56301 (A9), and Ltr4.2-585119 (A10), were present in PM breeding lines that were resistant to powdery mildew (Table 6, A8, A9, and A10). In the analysis of the 7 genetic sources, a C. chinense accession, ‘Bhut Jolokia’, one of the world’s hottest peppers, had all high capsaicinoid content marker types on the five capsaicinoid content assays (Table 6, A16-A20; Lee et al., 2016a). Six anthracnose-resistant C. baccatum accessions, including ‘PBC81’ and ‘PI594137’, had all of the resistance marker types for the three anthracnose resistance assays (Table 6, A3, A4, and A5; Park et al., 2009).
Fluidigm dynamic arrays, a high-throughput SNP genotyping method, include three formats for IFCs: 96 samples × 96 SNPs, 48 samples × 48 SNPs, or 192 samples × 24 SNPs. They can be used with three types of assays: TaqMan, KASP, or SNP type assays (Wang et al., 2009; Thomson, 2014). In this study, we used Fluidigm dynamic arrays for the first time for foreground selection of targeted genes or QTLs in pepper by combining 192.24 IFCs with SNP type assays to analyze 24 SNP markers at a time. This system can save both resources and time by reducing the reaction volume and producing 4,608 data points at a time (Wang et al., 2009). Of the 43 primer sets designed, 20 SNP type assays were successfully developed (Fig. 1 and Table 1). The accuracy of 11 assays was confirmed by comparing with the corresponding original HRM markers (Tables 2, 3, 4, and 5). These results suggest that the newly developed SNP type assays can be used in place of the original markers. The other 9 assays also need to be compared to original markers using polymorphic populations to confirm their accuracy. These SNP type assays will be useful in molecular breeding programs for developing new pepper varieties resistant to multiple diseases and with increased pungency.