Introduction
Downy Mildew
Factors Influencing DM Outbreaks
DM Detection
DM Control
Physical control
Chemical control
Biological control
Cultural control
Genetic Patterns of Resistance to Cucumber DM
Molecular Markers for Cucumber DM
QTL Mapping for DM Resistance
DM Resistance Genes
Breeding for DM Resistance
Discussion
Introduction
Cucumber (Cucumis sativus L.), originally native to India, was introduced into Central Asia and China. It ranks among the most widely cultivated vegetables globally and holds significant agricultural, economic, and nutritional importance (Xu et al. 2022). Global cucumber production totals 91.25 million tons, and China is the leading producer of cucumbers, contributing 72.83 million tons and thereby accounting for 79.8% of the total world output (Wang et al. 2018a).
Downy Mildew
Cucumbers are susceptible to numerous diseases, resulting in substantial yield losses. Among these, downy mildew (DM) is one of the most destructive threats to cucurbit crops (Savory et al. 2011). More than 80 countries face severe DM outbreaks, with most experiencing considerable economic effects (Lebeda and Urban 2004; Colucci et al. 2006). In 2004, DM was classified as a major disease affecting cucumbers in the United States (Holmes et al. 2004; Holmes and Thomas 2009).
The oomycete pathogen Pseudoperonospora cubensis, the causal agent of DM, severely damages the leaves of cucurbit crops (Wang et al. 2016). Being airborne, it infects through the leaf stomata and poses a highly destructive threat to cucumber production (Savory et al. 2011). Throughout the growing season, under favorable environmental conditions, DM spores repeatedly infect cucumber plants through a cyclical process involving sporangium germination, zoospore release, and host penetration, resulting in recurrent disease outbreaks (Fig. 1). DM exhibits a broad distribution, with over 80 countries reporting its presence (Palti and Cohen 1980; Lebeda and Urban 2007). Its host range is similarly wide, encompassing species from at least 20 genera within the Cucurbitaceae family (Palti and Cohen 1980; Lebeda and Urban 2007). The disease initially appears as chlorotic lesions along leaf veins on the adaxial leaf surface; as infection progresses, the centers of these lesions become necrotic. Under high humidity, black sporangia develop on the abaxial leaf surface. In advanced stages, the necrotic areas expand further, eventually leading to complete yellowing and collapse of the leaf. The disease spreads rapidly, and if unmanaged, localized infection can lead to the entire cucumber field being affected within a few days.
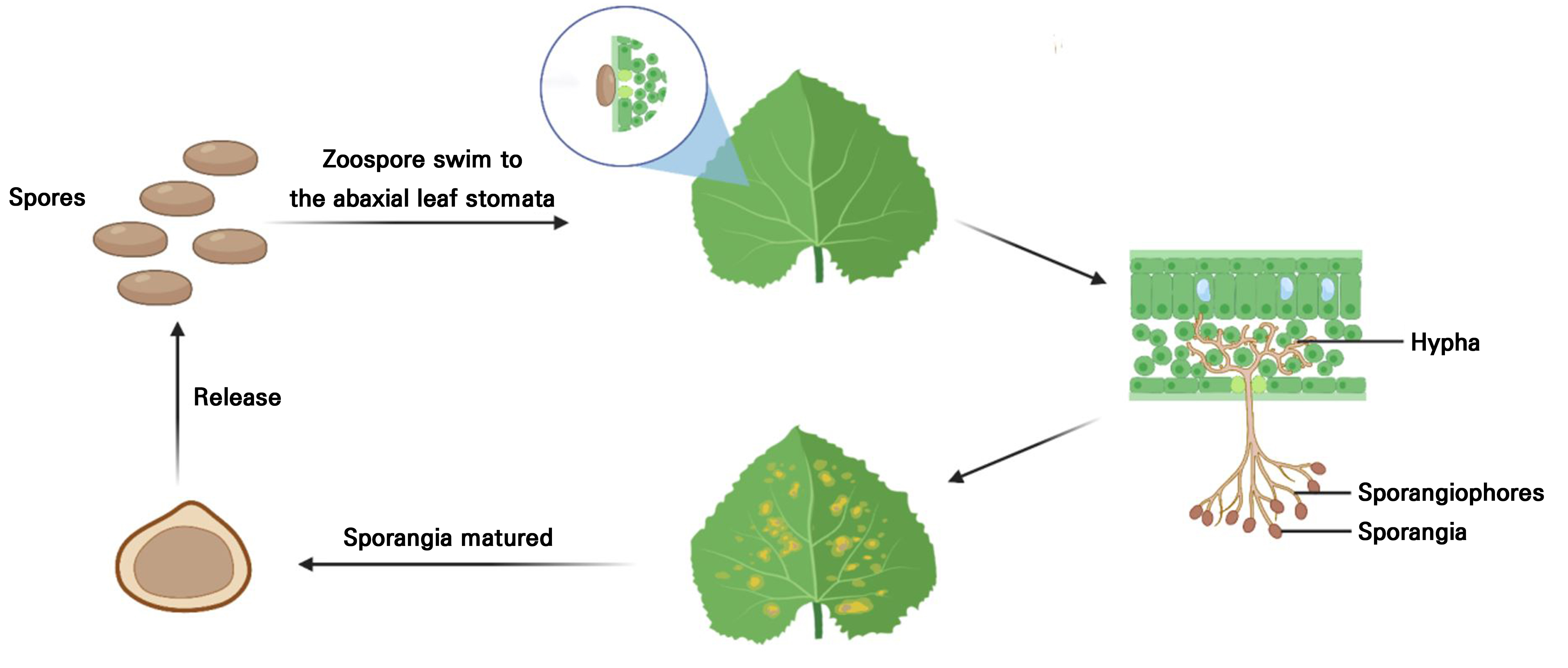
Fig. 1.
Infection cycle of cucumber DM. Zoospore swim to the external surface of the abaxial leaf stomata. Spore germination is followed by hyphal invasion through the stomata. Subsequently, a large number of sporangiophores and sporangia are formed in the infected tissues. Under favorable environmental conditions, the sporangia germinate and release zoospores, initiating a new round of infection.
Factors Influencing DM Outbreaks
DM development is influenced by several environmental conditions, among which temperature and humidity are the primary drivers of disease progression. DM infects plants in the temperature range of 5°C to 25°C. The optimum temperature range for spore release from the sporangia of cucumber DM is 5°C to 10°C, and the optimum temperature for DM sporangial germination is between 15°C and 20°C (Li et al. 2024). Therefore, this infection tends to spread in cold weather. In dry and hot weather, the development of the infection slows down. When internal greenhouse temperatures are in the range of 12–30°C and the relative humidity exceeds 83%, sporangia are produced in large quantities, with sporulation rates increasing as the humidity rises (Zhang 2020). A water film on the abaxial leaf surface, combined with humidity above 85%, results in the highest incidence of cucumber DM; in contrast, at 60–70% humidity, sporangial germination is limited and disease development in cucumber plants is hindered (Guo and Niu 2012). Therefore, outbreaks of cucumber DM are prevalent during periods characterized by heavy dew, fog, and frequent rainfall.
In addition to temperature and humidity, multiple other factors influence DM outbreaks. During cucumber disease outbreaks, proximity to the inoculum source is more critical than temperature (Cohen 1971). Planting frequency also influences the incidence of cucumber DM, with a positive correlation observed between successive plantings (first, second, and third) and disease occurrence (Ali et al. 2023). Long-term monoculture of cucumbers can lead to soil degradation, reduced organic matter, and disruption of the rhizosphere microbial community, thereby increasing the incidence of DM (Ali et al. 2023). Maintaining long-term soil health and managing plant diseases using integrated biological methods in monoculture-dominated Chinese production systems are essential to achieve sustainable agricultural goals. Cucumber plants are susceptible to infection during the cotyledon stage (Cohen and Eyal 1977), with disease severity increasing in later growth phases. Low-lying terrain, dense planting, and uneven fertilization (e.g., excessive nitrogen application) favor disease development (Wu 2024). Balanced nutrient management, including proper fertilization and foliar top-dressing, can enhance resistance to DM (Li and Xie 2024). However, the application of fertilizer must be carefully managed, as nutritional imbalances, especially excess nitrogen, elevate plant susceptibility to DM. Solar radiation also influences DM incidence; sunlight reduces sporangial viability such that stronger and longer exposure correlates with lower disease occurrence (Kanetis et al. 2010). In addition, host genetic traits and co-infection by bacterial or fungal pathogens may increase DM prevalence.
DM Detection
Early symptoms of DM are subtle, and current diagnostic practices depend on growers’ visual assessments based on experience. However, by the time visible symptoms appear, the optimal window for intervention has typically passed (Xu 2024). Therefore, early detection or even the prediction of pre-symptomatic DM infection using advanced technologies is critical for effective agricultural management.
Thermography (monitoring shifts in maximum temperature differences), Fourier transform infrared spectroscopy (targeting biomarker peaks at 2977, 1544, and 1050 cm-1), hyperspectral imaging (using Dis-CARS-SPA feature extraction), and optimized YOLO models (e.g., Faster-NAM-YOLO and improved YOLOv5s) have demonstrated efficacy in laboratory settings for detecting DM prior to symptom onset (Wen et al. 2019; Zhang 2022; Qiao et al. 2023; Qiao et al. 2024). Despite this potential, real-world implementation remains limited by three key challenges: (1) operational complexity in field deployment (e.g., hyperspectral systems requiring specialized hardware); (2) model fragmentation, where predictive algorithms such as long short-term memory–back propagation networks operate independently of diagnostic tools (Xu 2024); and (3) insufficient validation, with fewer than 10% of studies tested under actual greenhouse conditions (Li and Ge 2022). This gap highlights an urgent translational need: transitioning from isolated detection methods to integrated decision-support platforms that unify thermal, spectral, and meteorological data streams, enabling timely and actionable disease management in capital-intensive cucumber production systems where early intervention is economically essential. Dual quantitative polymerase chain reaction and high-resolution melting analyses are also used to detect DM, offering high sensitivity, rapid processing, and efficiency. However, these techniques entail complex protocols and high costs.
DM Control
Physical control
Improved greenhouse ventilation enhances control of DM, primarily by modulating relative humidity—a key factor influencing the spread of cucumber DM. In experiments conducted by Khudhair and Aljarah (2023), conventional non-ventilated greenhouses exhibited DM infection rates of 31.69%, 31.40%, and 29.35% in three cultivars. In contrast, structures equipped with three roof-mounted circular air vents (40 cm diameter each) arranged in an isosceles triangular configuration at 2.8 m spacing reduced infection to 1.47%, 1.17%, and 1.21%, respectively. These air vents promoted convective airflow that removed sporangia-laden humid air while maintaining microenvironmental conditions below the optimal range for P. cubensis. Consequently, the three openings in the greenhouse plastic covering reduced the infection rate and disease severity by 95.8% and 70%, respectively (Khudhair and Aljarah 2023). Higher radiation transmittance also suppresses DM development, as evidenced by lower disease incidence in greenhouses using films with greater light transmission. Ávalos-Sánchez et al. (2022) demonstrated that increased radiation transmittance of plastic mulch correlates with reduced DM incidence.
Chemical control
New chemical controls include the oxysterol-binding protein inhibitor Y18501, which inhibits multiple developmental stages of cucumber DM. The co-application of Y18501 with chlorothalonil significantly suppresses cucumber DM (Wang et al. 2023). Barnea et al. (2022) investigated the effects of mineral nutrition and salt fog on cucumber DM, reporting that elevated nitrogen, potassium, or calcium concentrations in irrigation water exacerbate disease severity. However, under conditions involving chloride ions, phosphorus and magnesium fertilization, and exposure to salt fog, disease incidence is markedly reduced. Yang et al. (2021) demonstrated that diallyl disulfide, a component of allicin, induces resistance to DM in cucumbers. In addition, Atallah et al. (2021) reported that soybean β-sitosterol and catfish skin mucus p22 glycoprotein exhibit significant antifungal activity against cucumber DM. These two substances disrupt the integrity of the sporangial cell walls of P. cubensis, thereby suppressing disease development.
Biological control
Biological control primarily relies on microbial competition, antagonism, or the production of secondary metabolites to suppress DM (Sun et al. 2022). Trichoderma harzianum can be effective when applied, and a single application in the seventh week reduces infection rates to 11.0%, compared with 99.7% in the control group in a study by Al-Aswad et al. (2021). This treatment also enhances plant length, leaf weight, leaf area, the total chlorophyll content, fruit number, and the overall yield (Elsharkawy et al. 2014). Fan et al. (2019) showed that fungichromin, a polyene macrolide antibiotic produced by Streptomyces PMS-702, significantly reduces DM severity. At a concentration of 10 µg·mL-1, fungichromin inhibits sporangial germination by 90%; however, at higher concentrations, it induces phytotoxicity, and 50 µg·mL-1 causes slight necrosis in cucumber leaves. Schuster and Schmitt (2018) reported that bio-surfactants containing spores of Bacillus subtilis S and Aneurinibacillus migulanus white bacterial preparations help reduce cucumber DM incidence. Szczech et al. (2017) found that Trichoderma asperellum TRS25 also effectively controls cucumber DM. When applied to soil using an organic carrier, TRS25 reduced plant infection by approximately 20% relative to the control; when used as a seed coating, infection was reduced by about 40%.
Cultural control
Cultural control is an important measure for preventing downy mildew. The growth vigor of the plants also determines the disease resistance of cucumbers. If cultivation management measures are inappropriate, the plant's resistance will be weakened. Therefore, proper fertilization should be carried out to enhance the growth of the cucumbers and reduce the probability of downy mildew occurrence on the plants (Li 2025). Planting should adhere to scientific density management protocols, with proper attention to ventilation and drainage, as excessively high humidity is a major factor contributing to severe disease outbreaks (Wang and Jin 2025). In addition, it is necessary to ensure the cleanliness of the greenhouse. Before transplanting, plant residue, weeds and production waste must be thoroughly removed from the facility. During the cultivation process, old leaves and yellow leaves must be promptly removed and taken outside the greenhouse for centralized destruction. These measures help to reduce the number of disease sources remaining in the field (Wang et al., 2024a).
Managing cucumber DM requires an integrated, multi-faceted approach. Optimizing physical conditions such as greenhouse ventilation and lighting, combined with the judicious use of chemical agents, the expanded application of biological control strategies, and strengthened cultivation management, can significantly reduce both disease incidence and improve cucumber yields and quality levels. Environmental sustainability is also carefully considered in these integrated management practices. In addition to the prevention and control measures mentioned above, researchers have also focused on screening resistant genes and cultivating resistant varieties to prevent and control downy mildew.
Genetic Patterns of Resistance to Cucumber DM
Research on the genetic basis of resistance to cucumber DM has yielded diverse models. Previous studies proposed varying mechanisms: Jenkins (1942) suggested polygenic inheritance, whereas Shimizu et al. (1963) reported monogenic recessive control. Van Vliet and Meijsing (1977) identified the recessive resistance gene p in ‘Poinsett’, and Fanourakis et al. (1987) supported recessive single-gene control. Conversely, some researchers proposed dominant gene-mediated resistance. Pershin et al. (1988) indicated that at least three dominant resistance genes exist in ‘Sadao Rischu’.
Recent research increasingly supports quantitative resistance governed by multiple genes. Zhang et al. (2013) classified resistance into two types: quantitative (polygenic) and qualitative (single-gene), with their experimental line K8 exhibiting polygenic quantitative resistance. Innark et al. (2020) confirmed that multiple recessive quantitative trait loci (QTLs) exert additive effects, aligning with Kelly and Vallejo (2006), who noted that quantitative resistance tends to be more durable.
The genetic mechanism underlying cucumber DM resistance is complex and is not yet fully resolved. Multiple modes such as recessive, dominant, monogenic, and polygenic inheritance may coexist. Future studies should use multi-omics approaches to dissect the synergistic interactions among the different resistance mechanisms, thereby advancing precision breeding for disease resistance. Numerous investigations of the genetic structure of cucumber DM resistance have been conducted thus far.
Molecular Markers for Cucumber DM
Advances in molecular biology and DNA sequencing have enabled the precise localization of target genes using techniques such as simple sequence repeat (SSR) and competitive allele-specific PCR. DNA fragments closely associated with target traits are developed into molecular markers, which have become essential tools in molecular studies (Li 2024), accelerating breeding programs by enhancing their speed, accuracy, and efficiency (Cortés et al. 2022). Horejsi et al. (2000) employed random amplified polymorphic DNA markers in marker-assisted selection, using the DM-susceptible (DMS) variety ‘Straight 8’ and DM-resistant (DMR) varieties ‘WI 1983G’ and ‘Zudm1’ as parental lines. They identified five markers linked to DM resistance (Table 1). Zhang et al. (2013) analyzed the genetic basis of DM resistance using inbred lines K8 (DMR) and K18 (DMS), identifying six SSR markers associated with five QTLs (dm1.1, dm5.1, dm5.2, dm5.3, and dm6.1) (Table 1). Also, Pang et al. (2013) crossed the introgression line IL52 (DMR) with ‘Changchunmici’ (DMS) to construct an F2 population, identifying four DM-linked markers: SSR20648 and SSR13639 on chromosome 5 (at genetic distances of 11.9 cM and 18.7 cM, respectively), and SSR21885 and SSR00233 on chromosome 6 (68.2 cM and 76.0 cM, respectively). Wang et al. (2016) used WI7120 and 9930 as parents to generate F2 and F2:3 populations, identifying four linked markers corresponding to four QTLs (dm2.1, dm4.1, dm5.1, and dm6.1) (Table 1). Win et al. (2017) constructed an F2 population from crosses between TH118FLM (DMR) and WMEJ (DMS), demonstrating that SSR21734 had the strongest linkage to dm2.2 and SSR11167 to dm5.1. Wang et al. (2018b) studied recombinant inbred line (RIL) populations derived from plant introduction (PI) 197088 and ‘Coolgreen’, identifying 11 QTLs (dm1.1, dm2.1, dm2.2, dm3.1, dm3.2, dm4.1, dm5.1, dm5.2, dm5.3, dm6.1, and dm6.2), each associated with a linked marker (Table 1). Wang et al. (2024b) investigated the RIL-37 population from the cross WIge7088D × ‘Coolgreen’, locating the dm5.3 locus between two indel markers, DM5.3-21 (30.8 cM) and DM5.3-25 (30.9 cM).
Table 1.
Molecular markers linked to downy-mildew-resistant genes in cucumber
| Code | Marker name | Genetic distance (cM) | Marker type | References |
| 1 |
m-G14800, X151100, AS5800, BC5191100, BC5261000 |
In the ‘WI 1983G’ × ‘Straight 8’ population, (G14800) 16.5 cM, (AS5800) 32.8 cM, (BC5191100) 9.9 cM, (BC5261000) 19.2 cM In the ‘Zudm1’ × ‘Straight 8’ population, (G14800) 20.9cM, (X151100) 14.8cM, (AS5800) 24.8cM, (BC5261000) 32.9cM | RAPD | Horejsi et al. 2000 |
| 2 |
SSR31116, SSR20705, SSR00772, SSR11012, SSR16882, SSR16110 |
5.0 cM, 10.5 cM, 5.0 cM, 15.7 cM, None, 13.8 cM | SSR | Zhang et al. 2013 |
| 3 |
SSR20648, SSR13639, SSR21885, SSR00233 |
11.9 cM, 18.7 cM, 68.2 cM,76.0 cM | SSR | Pang et al. 2013 |
| 4 |
CHXSSR00134, UW05916, SSR15321, SSR18651 |
37.6 cM, 22.9 cM, 51.6 cM,46.9 cM | SSR | Wang et al. 2016 |
| 5 | SSR21734, SSR11167 | - | SSR | Win et al. 2017 |
| 6 |
Marker1_16905088, Marker2_8521946, Marker2_20890971, Marker3_9882197, Marker3_37674467, SSR17911, Marker5_64722, Marker5_16754619, Marker5_25566302, Marker6_5079443, Marker6 26132968 | - | SNP, SSR | Wang et al. 2018b |
| 7 | DM5.3-21, DM5.3-25 | 30.8 cM, 30.9 cM | Indel | Wang et al. 2024b |
QTL Mapping for DM Resistance
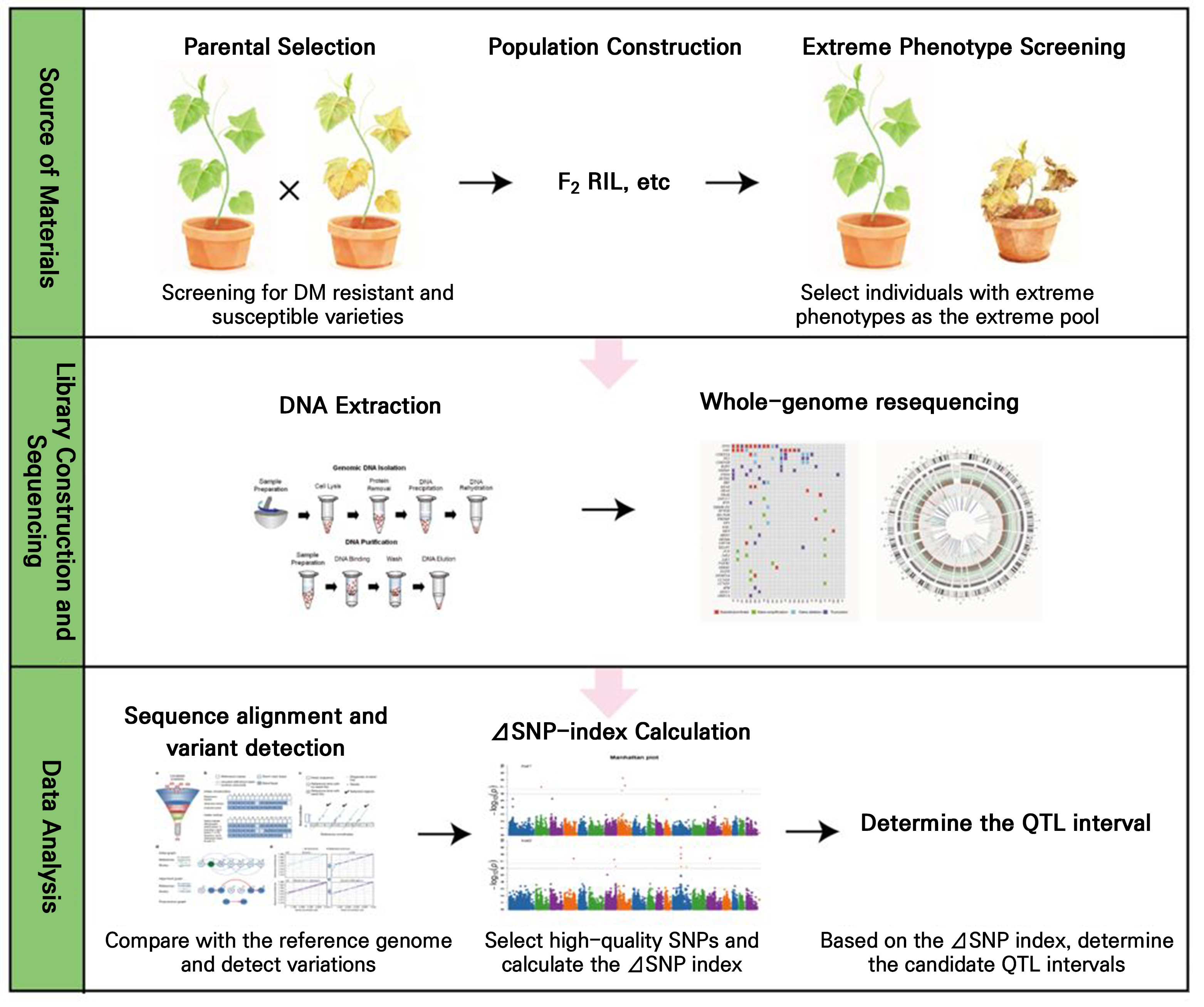
QTL mapping involves constructing appropriate genetic populations, such as F2 or RIL populations, and performing phenotypic assessments and genotypic analyses on individual plants (Fig. 2). By evaluating associations between marker genotypes and trait variation, QTLs that significantly influence the target trait can be identified, providing a foundation for map-based cloning and the marker-assisted selection of candidate genes in crop breeding. Bai et al. (2008) identified three DM-associated QTLs on linkage groups 1 and 6 using an RIL population derived from a cross between S94 and S06. Subsequently, Zhang et al. (2013) developed another RIL population using inbred lines K8 and K18 as parental lines; K8 carries cucumber DM resistance inherited from the Chinese hybrid ‘Yuanfeng’, which exhibits natural resistance, whereas K18 is susceptible. Their study detected five QTLs associated with DM resistance: dm1.1, dm5.1, dm5.2, dm5.3, and dm6.1. In the same year, Pang et al. (2013) generated an F2 population from a cross between IL52 (DMR) and ‘Changchunmici’ (DMS), identifying three QTL regions on chromosomes 5 and 6. Interval mapping revealed two significant QTLs on chromosome 5—dm5.1 and dm5.2—accounting for 17.9% and 14.2% of the phenotypic variance, respectively. Yoshioka et al. (2014) crossed CS-PMR1, a highly resistant line derived from PI 197088, with ‘Santou’, a moderately resistant cultivar, and detected ten QTLs by interval mapping. Among these, dm1.1, dm6.1, and dm7.2 significantly enhanced resistance, with dm1.1 showing the strongest effect. Caldwell et al. (2011) constructed a genetic population using PI 197088 (DMR) and identified one QTL each on chromosomes 2, 4, and 5. VandenLangenberg (2015) developed an RIL population derived from PI 197088 and ‘Coolgreen’, detecting three candidate QTLs for DM resistance on chromosomes 4 and 5. Wang et al. (2016) identified four QTLs for DM resistance—dm2.1, dm4.1, dm5.1, and dm6.1—using F2 and F3 lines from a cross between WI7120 (PI 330628) and ‘9930’, explaining 62–76% of the phenotypic variance. The major-effect QTLs were dm4.1 and dm5.1, while dm2.1 and dm6.1 exhibited moderate and minor effects, respectively; overdominance effects were observed for both dm2.1 and dm6.1 relative to dm4.1 and dm5.1. Win et al. (2017) analyzed two inbred lines, TH118FLM (DMR) and WMEJ (DMS), detecting five QTLs—dm2.2, dm4.1, dm5.1, dm5.2, and dm6.1—with dm2.2 having the greatest impact on resistance. A routine QTL analysis of the F2 population confirmed dm2.2 and dm5.2 as major QTLs and dm4.1 as a minor QTL. Further analysis using the F3 population revealed a new QTL in the dm2.1 region on chromosome 2. Wang et al. (2018b) reported that host resistance to DM and powdery mildew in PI 197088 are governed by 11 QTLs (three with major effects) and four QTLs (one with a major effect), respectively, with three loci co-localized. Of the 11 DM-resistance QTLs, dm5.1, dm5.2, and dm5.3 contributed major effects, whereas dm1.1, dm2.1, and dm6.2 were associated with susceptibility. Also, Li et al. (2018) constructed a population from a cross between PI 197088 (DMR) and ‘Changchunmici’ (DMS), developing a linkage map using 141 SSR markers. Five QTLs were detected on chromosomes 1, 3, 4, and 5, with the major-effect QTL dm4.1 (R² = 27%) consistently identified on chromosome 4. In addition, dm1.1 and dm5.2 showed moderate effects, while dm3.1 and dm5.1 were minor-effect QTLs. Liu et al. (2020) used 97 cucumber core germplasm accessions provided by the Cucumber Research Group at the Vegetable and Flower Research Institute of the Chinese Academy of Agricultural Sciences, applying genome-wide association studies to detect genetic loci linked to DM resistance across origins and ecotypes. Six stable loci—dmG1.4, dmG4.1, dmG4.3, dmG5.2, dmG7.1, and dmG7.2—were identified. Similarly, Innark et al. (2020) evaluated 14 cucumber accessions from the germplasm library at Rajamangala University of Technology Lanna, Thailand, detecting 14 QTLs accounting for 5.0–12.5% of phenotypic variance, including two major QTLs and two QTL hotspots. Collectively, QTLs conferring resistance to cucumber DM are distributed across all seven chromosomes. Although their exact positions vary among studies, chromosomes 5 and 6 harbor main-effect QTLs involved in key resistance pathways. Notably, chromosome 5 (particularly dm5.1, dm5.2, and dm5.3) emerges as a critical region for DM resistance, frequently detected in multiple investigations and potentially containing major resistance genes. Chromosome 6 (dm6.1) may interact epistatically with QTLs on chromosome 5. These findings indicate that resistance to cucumber DM is governed by a complex polygenic system.
DM Resistance Genes
Cucumber DM resistance genes have been reported in many studies (Table 2). The identification and development of DM resistance genes in cucumber are complicated by the polygenic nature of resistance. Liu et al. (2017) used cucumber varieties ‘D9320’ (DMR) and ‘D0401’ (DMS) to identify the resistance-associated gene CsERF004 by means of transcriptome sequencing. CsERF004 encodes a nuclear-localized protein; its overexpression in ‘D0401’ significantly upregulated defense-related genes CsPR1 and CsPR4, elevated salicylic acid (SA) and ethylene levels, and enhanced resistance to DM and other pathogens. Zhang et al. (2018) crossed ‘Changchunmici’ (DMS) with IL52 (DMR) and applied bulk segregant analysis (BSA)-sequencing to identify seven candidate DM resistance genes within the dm5.2 locus. Among these, Csa5M622800.1, Csa5M62230.1, and Csa5M623490.1 were also implicated in powdery mildew resistance. Csa5M622830.1, encoding a GATA transcription factor involved in carbon and nitrogen metabolism (Bi et al. 2005), may restrict nutrient availability to the obligate biotrophic DM pathogen (Evangelisti et al. 2014), thereby contributing to resistance. WRKY transcription factors have been widely studied in plant responses to biotic stress. Luan et al. (2019) reported that CsWRKY50, a subgroup IIc WRKY transcription factor localized in the nucleus, enhances cucumber resistance to DM by regulating reactive oxygen species and proline accumulation; modulating peroxidase, superoxide dismutase, and catalase activities; and influencing SA- and jasmonate (JA)-responsive and SA biosynthesis genes. Zheng et al. (2019) identified CsTCP14 as a key regulatory gene whose expression was markedly induced upon DM infection. Conversely, Csa6M344280.1 exhibited a negative correlation with CsTCP14 expression and was potentially targeted by the CsNBS-LRR gene product; transcriptional repression by this factor may increase susceptibility to foliar diseases. Wang et al. (2018a) used QTL mapping and map-based cloning with the resistant line ‘Gy14’ for causal gene isolation. They demonstrated that DM resistance in ‘Gy14’ is controlled by the cucumber STAYGREEN gene CsSGR, where a single-nucleotide polymorphism in the coding region results in a non-synonymous amino acid substitution conferring resistance. Liu et al. (2020) conducted genome-wide association studies of a core germplasm of diverse cucumber accessions to detect loci associated with DM resistance. Stable and resistance loci were linked to five potential causative genes: Csa1G575030 (dmG1.4), Csa2G060360 (dmG2.1), Csa4G064680 (dmG4.1), Csa5G606470 (dmG5.2), and Csa7G004020 (dmG7.1). Berg (2019) analyzed PI 197088 and identified a receptor-like kinase gene (CsLRK10L2) in sub-QTL dm4.1.2, which may play a primary role in detecting DM spores and sensing cell wall damage. In addition, a mutation in the amino acid permease 2A gene (CsAAP2A) was found in sub-QTL dm4.1.3. Cucumber lines carrying a functional allele deletion in CsAAP2A accumulated lower amino acid levels post-inoculation, suggesting reduced transport of nutrients to infected tissues. Tan et al. (2022) developed near-isogenic lines (NILs) from PI 197088 to model the dm5.3 resistance QTL dm5.3 and fine-mapped the locus using NIL-derived segregating populations. The sigma factor binding protein 1 gene CsSIB1 (CsGy5G027140) was identified as a candidate. Elevated CsSIB1 expression promoted SA signaling antagonistically while suppressing JA biosynthesis and signal transduction pathways. Recently, Wang et al. (2024b) identified ten CsZIP genes from whole-genome sequences of ‘China Long 9930’ and transcriptome data, all of which responded to DM infection and were designated CsZIP1 through CsZIP10. Collectively, these findings demonstrate substantial progress in characterizing cucumber DM resistance gene loci, with promising implications for molecular breeding applications.
Table 2.
QTLs and candidate genes for cucumber DM resistance
| Origin | QTL | Chromosome | Candidate gene | Ref. |
| S94 | - | Bai et al. 2008 | ||
| K8 | dm1.1, dm5.1, dm5.2, dm5.3, dm6.1 | Chr1, Chr5, Chr6 | - | Zhang et al. 2013 |
| IL52 | dm5.1, dm5.2 | Chr5 | - | Pang et al. 2013 |
| CS-PMR1 | dm1.1, dm6.1, dm7.2 | Chr1, Chr6, Chr7 | - | Yoshioka et al. 2014 |
| PI 197088 | Chr2, Chr4, Chr5 | - | Caldwell et al. 2011 | |
| WI7120(PI 330628) | dm2.1, dm4.1, dm5.1, dm6.1 | Chr2, Chr4, Chr5, Chr6 | - | Wang et al. 2016 |
| TH118FLM | dm2.2, dm4.1, dm5.1, dm5.2, dm6.1 | Chr2, Chr4, Chr5, Chr6 | - | Win et al. 2017 |
| D9320 | - | - | CsERF004 | Liu et al. 2017 |
| PI 197088 | dm5.1, dm5.2, dm5.3 | Chr5 | - | Wang et al. 2018b |
| PI 197088 | dm1.1, dm3.1, dm4.1, dm5.1, dm5.2 | Chr1, Chr3, Chr4,Chr5 | - | Li et al. 2018 |
| IL52 | dm5.2 | Chr5 | Csa5M606470.1, Csa5M608020.1, Csa5M608030.1, Csa5M615270.1, Csa5M622800.1, Csa5M62230.1, Csa5M623490.1 | Zhang et al. 2018 |
| 9930 | - | - | CsTCP14, Csa6M344280.1 | Zheng et al. 2019 |
| Gy14 | dm5.2 | Chr5 | CsSGR | Wang et al. 2018a |
| 97 cucumber CG populations | dmG1.4, dmG4.1, dmG4.3, dmG5.2, dmG7.1, dmG7.2 | - | Csa1G575030, Csa2G060360, Csa4G064680, Csa5G60647, Csa7G004020 | Liu et al. 2020 |
| PI 197088 | dm4.1.1, dm4.1.2 | Chr4 | CsLRK10L2 | Berg 2019 |
| PI 197088 | dm5.3 | Chr5 | CsSIB1 (CsGy5G027140) | Tan et al. 2022 |
| 9930 | - | - | CsZIP1, CsZIP2, CsZIP3, CsZIP4, CsZIP5, CsZIP6, CsZIP7, CsZIP8, CsZIP9, CsZIP10 | Wang et al. 2024b |
Breeding for DM Resistance
Relentless efforts have long been made by researchers in the race against DM through the breeding of resistant varieties (Table 3). Jenkins (1942) was the first to systematically screen cucumber lines for resistance to DM. Among the 12 tested lines, he identified ‘Puerto Rico 37’ and ‘China Long’ as exhibiting higher resistance, with non-segregating resistance observed in hybrid offspring. Barnes and Epps (1954) developed ‘Palmetto’ from ‘China Long’, but its resistance was lost after two years of cultivation. Barnes and Epps (1954) also identified a new resistant accession, PI 197087, from India. The dm-1 gene derived from PI 197087 effectively suppressed DM development, providing robust resistance that eliminated the need for fungicide applications (Holmes et al. 2004). This dm-1-mediated resistance became widely incorporated into commercial cucumber cultivars in the United States, including ‘Gy14’, ‘Poinsett 76’, ‘M21’, and PI 234517 (SC50) selected by Wehner and Shetty in 1997 (Call 2010), as well as varieties in the ‘Polaris’, ‘Clinton’, ‘Chipper’, ‘Poinsett’, and ‘Marketmore’ series (Wang et al. 2018a). The dm-1 gene confers recessive resistance to DM and is a mutated allele resulting from the deletion susceptibility gene at the CsSGR (STAYGREEN) locus on chromosome 5 (Wang et al. 2018a). For decades, most resistant lines carried monogenic or oligogenic resistance originating from either ‘China Long’ or PI 197087 (Barnes and Epps 1954; Cavatorta et al. 2007; Holdsworth et al. 2014). For nearly half a century, dm-1-based resistance successfully prevented severe DM damage without reliance on chemical control methods. However, this situation was upended in 2004 by the emergence of new DM strains. In 2004, a new strain of P. cubensis appeared in the southern United States (Holmes et al. 2006), overcoming the resistance in the country’s commercial cultivars. In affected production systems, yield losses reached 95–100% (Colucci et al. 2006; Savory et al. 2011), with substantial economic effects (Colucci et al. 2006). In many regions in the northern United States, organic cucumber production and late-season cropping were discontinued due to DM outbreaks (Holdsworth et al. 2014). Following extensive screening efforts, researchers identified several accessions with high resistance to these new strains, including PI 197085, PI 197088, PI 330628 (WI7120), and PI 605996 (Call et al. 2012). Among these, PI 197088 and PI 330628 showed the strongest resistance in the field to the strains of DM prevalent in the U.S. since 2004 (Tan et al. 2022). Studies conducted in the decade following 2004 revealed that the resistance levels of commercially available cultivars and USDA repository accessions generally fell below those of pre-2004 commercial standards (Call and Wehner 2010; Call et al. 2012). Consequently, breeding DM-resistant cucumber cultivars has remained a primary focus for public and private breeders and researchers in the United States (Weng 2008). Holdsworth et al. (2014) used pedigree-based breeding methods to develop a series of DM-resistant cucumber lines using ‘Ivory Queen’, ‘Marketmore 97’, ‘Platinum’, and ‘Salt & Pepper’ as parental genotypes. From crosses involving ‘Salt & Pepper’, researchers developed multiple breeding lines resistant to DM, including varieties with green skins (e.g., NY12263, DMR-NY264) and white skins (e.g., NY12257, NY12-258, NY12-260, NY12-261, and NY12-262). The most resistant line DMR-NY264 recorded an honestly significant difference (HSD) value of 817, whereas the highly susceptible cultivar ‘Picolino’ had an HSD of 3618. The four most resistant ‘Cornell’ lines (DMR-NY264, NY12-257, NY12-260, and NY12-261) demonstrated significantly stronger resistance than all evaluated commercial cultivars and PI accessions, although their performance remained inferior to that of PI 197088 and PI 330628. PI 432870, PI 390266, PI 418964, PI 390258, and G12 were identified in non-Sikkim cucumbers, which exhibit high resistance to DM (Hammer and Cohen, 2025). In summary, DM remains a serious disease affecting cucumbers, and ongoing efforts continue to focus on breeding new resistant varieties. Several promising lines have been developed, offering significant potential for sustainable cucumber production.
Table 3.
Cucumber varieties exhibiting DM resistance
| Code | Resistant variety | Researcher | Year |
| 1 | Puerto Rico 37 (PR37), China Long | Jenkins | 1942 |
| 2 | Palmetto | Barnes | 1954 |
| 3 | PI 197087 | Barnes and Epps | 1954 |
| 4 | Gy14, Poinsett 76, M21, PI 234517 (SC50) | Wehner and Shetty | 1997 |
| 5 | Polaris, Clinton, Chipper, Poinsett, and Marketmore series | - | - |
| 6 | PI 197085, PI 197088, PI 330628 (WI7120), and PI 605996, | - | - |
| 7 | NY12263, DMR-NY264, NY12257, NY12-258, NY12-260, NY12-261, and NY12-262 | Holdsworth | 2014 |
| 8 | PI 432870, PI 390266, PI 418964, PI 390258, and G12 | Hammer and Cohen | 2025 |
Discussion
Cucumber DM is one of the most serious diseases threatening cucumber production, with a broad distribution that poses a significant threat to cultivation. Current research explores various prevention and control strategies, including physical, chemical, biological, and resistance breeding methods. Biological control agents and resistance-inducing compounds have shown potential, while the impact of climate change on pathogen‒host dynamics warrants further investigation. Artificial intelligence and big data analytics are increasingly used to optimize assisted selection and genetic improvements for durable resistance. Substantial progress has been achieved in identifying candidate resistance genes and conducting genomic analyses. With ongoing advances in biotechnology, techniques such as gene silencing and gene editing offer powerful tools for elucidating disease resistance gene functions and mechanisms. However, resistance to cucumber DM is governed by a complex genetic system, and there remains no consensus with regard to the inheritance of resistance. Thus, further research is required to identify effective resistance genes and understand their modes of action.
Research on the pathogenic mechanisms of cucumber DM also merits attention. There is no unified classification of physiological races, and studies characterizing the pathogen itself remain limited. Elucidating its pathogenic mechanisms could enable effective and sustainable protection against DM, while understanding the molecular mechanisms of host defense strategies are important research directions.
Before 2004, the dm-1 gene from PI 197087 conferred effective resistance to DM, eliminating the need for fungicides (Holmes et al. 2004). However, the emergence of new DM strains in the United States in 2004 rendered the dm-1 gene ineffective. The pathogen gradually overcame dm-1-mediated single-gene resistance through frequent mutations or via an avirulence gene recombination route. Prolonged reliance on dm-1 imposed strong selection pressure, accelerating the evolution of virulent pathogen populations. At present, effective control measures still rely on the use of new resistant varieties, combined with the use of fungicides, to control DM and prevent yield losses. However, current commercial resistant varieties do not achieve the same level of resistance previously exhibited by PI 197087. Researchers continue with extensive efforts to develop improved resistant cultivars. Recent approaches integrate genomic studies, quantitative resistance, and gene editing as complementary strategies. Pyramiding can combine dm-1 with other major genes or minor-effect QTLs by backcrossing or transgenic technology. Marker-assisted selection frameworks provide efficient breeding solutions. The continued exploration of elite DM resistance germplasm is essential, using advanced technologies such as QTL sequencing, transcriptome sequencing, and next-generation sequencing-assisted BSA to discover molecular markers and screen candidate genes. As more resistance genes are identified, their introgression into elite breeding lines and cultivated varieties will become increasingly feasible. Breeding new, high-quality, and durably resistant varieties remains a key objective in ongoing research.