Introduction
Material and Methods
Plant Materials and DNA Extraction
Virus Materials and Inoculation
Detection of the Presence of CMV by ELISA
RAPD Analysis and Linkage Group Construction
Results and Discussion
Resistance Responses to CMV P1 of C. annuum
Inheritance of Resistance to CMV P1 Strain in C. annuum ‘I7339’
Linkage Analysis of the CMV P1 Resistance Loci
Introduction
Various plant viruses infect peppers (Capsicum annuum L.) cause inhibition of growth, decrease of fruit quality and death. Cucumber mosaic virus (CMV) is one of the important viral diseases in Korea and worldwide, infecting many different crop species including peppers (Choi et al., 2005; Palukaitis et al., 1992). Besides its importance, no complete resistances were found capable of restriction of CMV replication or long-distance movement in pepper. In some cucurbits, resistance to CMV was controlled by one recessive gene (Kang et al., 2005), tomato and Arabidopsis have single dominant gene for resistance (Stamova and Chetelat, 2000; Takahashi et al., 2001, 2002). Several poly-genes for resistance were reported in Capsicum species including ‘Perennial’ (Caranta et al., 1997; Chaim et al., 2001; Grube et al., 2000; Lapidot et al., 1997), ‘Vania’ (Caranta et al., 2002), C. baccatum ‘PI439381-1-3’ (Suzuki et al., 2003) and C. frutescens ‘BG2814-6’ (Grube et al., 2000). In Arabidopsis and tomato, single dominant gene RCY and Cmr were suggested for resistance (Stamova and Chetelat, 2000; Takahashi et al., 2001, 2002). Up to date, three different steps of inhibition of the virus cycles were reported in distinct resistant Capsicum accessions: restriction of host-cell infection (Caranta et al., 1997), suppression of virus multiplication in the whole plants (Nono-Wondim et al., 1993) and restraint of long- distance movement (Dufour et al., 1989).
In Korea, CMV was reported all province cultivating peppers (Choi et al., 2005). Some commercial cultivars, such as ‘Bukang’ have been used for the resistance sources to CMV in seed companies. From progeny tests, it was revealed that ‘Bukang’ have a single dominant gene Cmr1 resistant to CMV Fny and CMV P0 (Kang et al., 2010). For the single dominant resistant gene, three closely linked molecular markers were developed and applied for breeding. Recently, a new strain CMV P1 was reported, which could infect cultivars resistant to both CMV Fny and CMV Korean (Kang et al., 2010; Lee et al., 2006). Korean pepper breeders have been developed many cultivars resistant to both CMV and Phytophthora root rot for open field cultivation, however occurrence of CMV infected plants did not decreased and strength of resistance to CMV was different in many CMV resistant cultivars in the repeated annual survey. Therefore, breeding of more robust CMV resistant cultivars will be important goal of seed companies. In this study, we surveyed inheritance of CMV P1 resistance, and described the develop-ment of markers linked to the resistance loci.
Materials and Methods
Plant Materials and DNA Extraction
A cayenne-pepper inbred line ‘I7339 (I)’ was used as a CMV P1resistant parent and another inbred line ‘E1338 (E)’ was used as a susceptible parent, both were obtained from DongbuFarm Hannong (Anseong, Korea). A CMV Fny resistant cultivar ‘Bu-hong’ (DongbuFarm Hannong, Korea) and one CMV P1 resistant accession ICPN-18-8 (provided from Dr. Myung-Cheorl Cho, National Institute of Horticultural & Herbal Science, Korea) were included. Yolo Wonder was used as susceptible controls in each inoculation. The F1 hybrid (E1338 × I7339) was self-pollinated to produce 189 F2 plants, and they were analyzed for both CMV resistance and molecular markers. Total genomic DNA was isolated using the protocol described by Kim and Kim (2005).
Virus Materials and Inoculation
CMV P1 was provided by Dr. Byoung-Cheorl Kang (Seoul National University, Seoul, Korea). The seedlings were inoculated with CMV P1 at two leaves stage. The inoculums were prepared from the infected leaves of Nicotiana benthamiana. One gram of infected leaves was ground with 10 mL of 0.1 M phosphate buffer pH 7.0 and carborundum #400 (Junsei Pure Chemical Ind., Japan) and inoculated by rubbing the virus onto the two cotyledons. Control plants were mock-inoculated with 0.1 M phosphate buffer pH 7.0. After inoculation, the inoculated plants were kept in a growth chamber at 25°C and symptoms were observed two times, 15 days and 45 days post inoculation (dpi).
Detection of the Presence of CMV by ELISA
To test systemic infection, ELISA was used to detect CMV P1 according to the manufacturer’s protocol (Agdia, USA). Samples for ELISA test were collected and the upper leaves from inoculated cotyledons at 15 dpi and upmost leaves from top branches at 45 dpi, two times. Samples were considered to be positive for the presence of CMV P1 when the absorbance value (405 nm) of each sample was greater than the mean of two healthy control ‘Yolo Wonder’ plants.
RAPD Analysis and Linkage Group Construction
RAPD analyses were performed on each of DNA from resistant and susceptible bulks and 189 individual plants of F2 population. A total of 1,200 random primers from OPA to OPBH primer sets (Operon Technologies, Alameda, CA) and 800 primers from UBC001 to 800 primer sets (University of British Columbia, Vancouver, BC, Canada) were screened. RAPD reactions were performed as described by Zhang et al. (2000). The genomic DNA of the parent cultivars were screened with simple sequence repeat (SSR) and sequence tagged sites (STS) markers (Lee et al., 2009; Mimura et al., 2012; Minamiyama et al., 2006). RAPD and other marker genotyping data were analyzed using CarthaGene software (Schiex and Gaspin., 1997).
Results and Discussion
Resistance Responses to CMV P1 of C. annuum
Cucumber mosaic virus is a major viral disease in Korea, their sequence variations, virulence and survey of resistant germplasm were rarely studied. A new strain CMV P1 infecting resistant cultivars to CMV Fny and CMV P0 was reported, and belonged to subgroup IB different to CMV Fny (subgroup IA) from movement protein (MP) and coat protein (CP) sequence analysis (Lee et al., 2006).
From CMV P1 inoculation, all of inoculated 'Yolo Wonder' plants were susceptible to CMV P1 strain, indicated that the CMV P1 screening method is reliable and consistent. Both vein clearing mosaic and leaf distortion symptoms were observed at 15 dpi and expanded to systemically. One accession of CMV P1 resistant accession ICPN18-8 showed no visual symptoms at 15 dpi, only detected by ELISA test, finally showed delayed symptoms after 45 dpi. All ‘Bu-hong’ plants were susceptible to the CMV P1 strain, indicated that the CMV P1 strain can overcome the CMV resistance in ‘Bu-hong’ plants. CMV P1 symptoms were appeared on the leaves of ‘E1338’ plants at 15 dpi, but no visible symptoms were observed in ‘I7339’ plants. The result of ELISA test, CMV was detected only in the upper leaves of ‘E1338’ plants, not in the upper leaves of ‘I7339’ plants at 15 dpi (Table 1).
Times for virus resistance test of plant breeding materials will be different for each virus types, but usually started at cotyledon stages or one pair of true leaves completed 15 to 20 dpi.
Changes in resistance through growth stages of host plant had been rarely reported. In C. annuum, Tobacco Mosaic Virus (TMV)-induced gene (CaTin2) encodes a mature 23 kDa protein that shares little sequence similarity with other cell wall proteins, fluorescently-tagged CaTin2 appeared to localize and in common with other cell wall proteins (Shin et al., 2003). CaTin2 was usually expressed in leaves and roots, constitutive expression of CaTin2 in transgenic tobacco resulted in some resistance against TMV or CMV, between 7 and 14 dpi after inoculation, but the resistance was lost one month after inoculation, the expression of the CaTin2 cell wall protein probably restricted virus movement and accumulation (Shin et al., 2003).
Inheritance of Resistance to CMV P1 Strain in C. annuum ‘I7339’
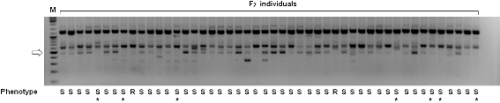
In the F2 population, a total of 189 plants were examined for CMV P1 resistance in two times, first at 15 dpi and second at 45 dpi. Of these, 42 individuals showed resistance, 147 plants showed susceptible from ELISA test at the first test (15 dpi). The phenotype segregation ratio was closed to the 1:3 ratio (resistance : susceptible plants), indicating single recessive gene inheritance pattern (χ2 = 0.002, P = 0.967). In the lateral ELISA test at 45 dpi, 11 plants showed resistance, 178 plants showed susceptible, the phenotype segregation ratio was close to the 1:15 (resistance: susceptibility) (χ2 = 0.060, P = 0.807) (Table 1). A total of 31 resistant plants at 15 dpi became susceptible at 45 dpi. These results indicated that the resistance in C. annuum ‘I7339’ is controlled by two different genes with a recessive manner. We named these resistance genes ‘cmr3E’ and ‘cmr3L’ respectively.
Studies for the resistance to CMV in C. annuum have been reported many research groups, most of them were incomplete dominant and quantitatively inherited (Caranta et al., 1997, 2002; Grube et al., 2000), except for Cmr1 (Kang et al., 2010). Cmr1 on the LG2 is the only dominant resistance gene known in pepper, but resistance is broken down by CMV P1 strain, requiring more robust CMV resistant germplasm (Kang et al., 2010). Recently, single recessive gene ‘cmr2’ resistant to CMV P1 was reported in pepper, located on chromosome 1 (Park et al., 2011). The resistance was controlled at the level of virus replication, different to ‘Cmr1’ which involves inhibition of the systemic movement of CMV (Kang et al., 2010).
In this study, virus infection was tested two times, 15 dpi and 45 dpi. Firstly, 47 plants showed no visible symptoms and negative in ELISA test at 15 dpi, 36 individuals become susceptible at 45 dpi. These results indicated that two different mechanisms might be involved for the resistance to CMV P1 - one gene for suppression in cellular level of virus replication (cmr3E), the other for inhibition of long distance movement of CMV (cmr3L). Because CMV inoculated plants of EI F2 population were transplanted to vinyl green-house at 30 dpi, gene for suppressing CMV replication (cmr3E) could be affected by environmental conditions such as high temperature.
Linkage Analysis of the CMV P1 Resistance Loci
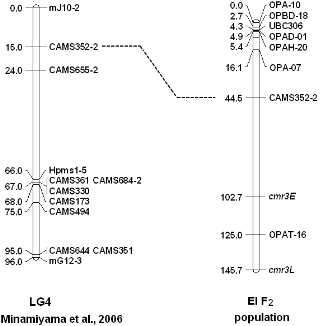
To locate cmr3E and cmr3L loci in pepper linkage map, total 2,000 RAPD primers, 115 SSR markers and 37 STS markers were screened using two parental lines, then polymorphic markers were test in the resistant and susceptible bulks of EI F2 population, only 9 primers were showed polymorphisms between bulked plants and mapped (3.6%). Because two parents of EI F2 population were breeding lines they shared low polymorphisms compared to wild types. One linkage group was constructed containing 9 markers (Fig. 2). It shared one common marker (CAMS352-2) with LG4 in the map of Minamiyama et al. (2006), other SSR markers were monomorphic between parents used int this study. Only one RAPD primer OPAT16 (5’-CTCTCCGTAG-3') was distantly linked with cmr3E (22.3 cM) and cmr3L (20.7 cM). Two loci for resistance to CMV P1 were located on the LG4 in this study, different to the cmr2 on chromosome 1 in previous study (Park et al., 2011), due to differ in genetic resources. Therefore, it is supposed that the gene required for inhibition of the systemic movement of CMV are absent in 36 plants, resulting in susceptibility. In the RNA3 sequence analysis for P0 resistant-breaking isolate (Ca-P1-CMV), the same CMV isolate used in this study, revealed Ca-P1-CMV belongs to CMV subgroup IB whereas the ordinary strain (Fny-CMV) group to subgroup IA from movement protein (MP) and coat protein (CP) comparison (Lee et al., 2006). They also suggested the possibility of changes in the MP and CP sequences resulting in different responses between pepper and virus. Similar response was found in the ‘Bu-hong’ cultivar (resistant to CMV Fny), it showed susceptible symptoms after inoculated with CMV P1. Further research remains for the comparison of genes for resistance to different CMV isolates in different pepper germplasms.
One distantly linked marker OPAT16 is insufficient for marker assisted selection of breeding lines or germplasms, requiring more closely linked markers. To narrow down and delimit loci, AFLP analysis and diverse approaches using pepper genome sequences being performed, further experiments will be needed to clarify genetic and inheritance of genes for resistance to diverse CMV strains including CMV P1.