Introduction
Watermelon [Citrullus lanatus (Thunb.) Matsum. et Nakai] is an important fruit vegetable, which provides nutritious juice that is beneficial for human health (Ren et al., 2012; Inthichack et al., 2014; Soteriou et al., 2014). Global watermelon production contributes to approximately 9.5% of overall vegetable production (http://faostat.fao.org). In Korea, the production of watermelon reached 672,000 tons in 2014, and yield increased 3.64% from 2003 to 2014. Powdery mildew, one of the major diseases of watermelon, reduces yields and quality of watermelon fruits worldwide. In watermelon, powdery mildew could be caused by Podosphaera xanthii (Castagne) U. Braun & N. Shishkoff (syn. Sphaerotheca fuliginea, formerly Sphaerotheca fusca Blumer) and Golovinomyces cichoracearum (syn. Erysiphe cichoracearum DC ex Merat.) (Sakata et al., 2006). These fungi belong to the same family (Erysiphaceae) and are often difficult to distinguish from each other without performing morphological and molecular analyses. Common cucurbit plants including watermelon are adapted to temperate conditions (Halliwell et al., 1986), and powdery mildew disease is a common problem in greenhouses kept at dry and low-temperature conditions (Keinath et al., 2010; Aguiar et al., 2012). Various fungicides are used intensively to control powdery mildew disease (Keinath and DuBose, 2004), resulting in increasing public health issues due to chemical residues on fruits, as well as the emergence of fungicide-resistant strains (Derbalah and Elkot, 2011). The development of resistant cultivars represents the most suitable alternative to fungicides to control the disease (Yuste-Lisbona et al., 2010).
Powdery mildew disease in watermelon is mainly caused by P. xanthii (Davis et al., 2007), of which race 1 was conventionally determined based on the susceptibility of melon (C. melo L.) differentials: susceptible for ‘Top Mark’ and resistant for ‘PMR 45’, ‘PMR 1’, and ‘PMR 6’ (McCreight, 2006). P. xanthii races that are infectious to melon can also infect watermelon, as indicated for the race 1W pathotype, which includes races derived from melon and infective to watermelon (Davis et al., 2007). Subsequently, pathotypes of P. xanthii that infect watermelon have been reported, including 2WU.S.(Tetteh et al., 2007) from the United States and 2WF from China (Zhang et al., 2011); ‘PMR 6’ is resistant to these pathotypes, while ‘Top Mark’ and ‘PMR 45’ are susceptible.
Resistance to powdery mildew has long been studied in watermelon, which is commonly considered to have the highest resistance among cucurbits to powdery mildew (Kristkova et al., 2009). In the United States, eight of 1573 accessions presented high resistance to P. xanthii race 1 (Davis et al., 2007), and Tetteh et al. (2010) reported that some wild-type lines exhibit intermediate resistance to the race 2WU.S. Furthermore, Zhang et al. (2011) discovered a potential source of resistance to the race 2WF, which is the predominant race of P. xanthii in China. In South Korea, 23 cultivars out of 120 watermelon accessions showed moderate resistance to powdery mildew (Lee et al., 2010). In another trial, 51 commercial cultivars of cucumber (Cucumis sativus), melon (Cucumis melo), and watermelon were screened for resistance to P. xanthii race 1, but this strain was not pathogenic on tested watermelon cultivars (Lee et al., 2014). The authors suggested that the tested P. xanthii race 1 that is infectious to cucumber and melon does not correspond to race 1W, which is infectious to watermelon. Kim et al. (2013) identified naturally occurring powdery mildew on watermelon in a greenhouse in South Korea as race 1 of P. xanthii using melon differentials during a study of inheritance of the resistance trait in watermelon. However, in general, both P. xanthii and G. cichoracearum are usually found solo or in mixed form around the greenhouse (McCreight, 2006; Chen et al., 2008; Horst, 2013). Thus, identification of the powdery mildew genus is a prerequisite for characterizing the causal fungi for powdery mildew in watermelon.
Although powdery mildew is a common disease in watermelon, few molecular markers associated with powdery mildewhave been developed (Kim et al., 2013; Gama et al., 2015). The limited development of resistance-linked markers has mainly been caused by the difficulty in collecting and maintaining the fungi to infect watermelon.
In this study, we performed the first characterization of the genus and pathotype of the causal fungi of powdery mildew on watermelon in South Korea based on morphological and molecular characteristics and differential reactions to the host plants for corresponding races. In addition, we attempted to develop molecular markers linked to the putative genomic region for resistance to the P. xanthii race 1W-AN using near-isogenic lines (NILs; BC4F6) derived from a cross between SBB (recurrent parent; Korean elite inbred line) and PI 254744 (donor parent; US Plant Introduction).
Materials and Methods
Plant Materials
For race identification, melon (Cucumis melo L.) plants used as differential hosts included ‘PMR 45’, ‘PMR 1’, ‘PMR 5’, ‘PMR 6’, ‘MR 1’, PI 124112, PI 313970, PI 414723, and ‘Top Mark’ were used. Eight NILs of BC4F6, from the Korean elite line (C. lanatus var. lanatus; ‘SBB’; recurrent plant) and the P. xanthii race 1W-resistant line (C. lanatus var. lanatus; US PI254744; donor plant), were developed by Partner Seed Company (Ansung, Korea) in order to develop powdery mildewresistant cultivars. To analyze resistance inheritance, the parental lines (‘SBA’: P1 and PI 254744: P2) and their F1, BC1P1, BC1P2, and F2 progeny were used. BC4F6 NILs, ‘SBB’, and PI 254744 were used to analyze random amplified polymorphic DNA (RAPD) and sequence characterized amplified regions (SCAR). To validate candidate linked markers, 107 of their segregated progeny (BC4F6 NIL-1 × ‘SBB’) and 138 F2:3 progeny derived from ‘SBA’ × PI 254744 were used. All tested plants were grown in plug-trays and transplanted in a mulched field in a greenhouse at Chung-Ang University.
Race Identification and Seedling Inoculation
Powdery mildew isolates originated from infected commercial watermelon plants at Chung-Ang University (Ansung, Korea) were maintained on ‘Top Mark’ melons and ‘SBB’ watermelons at 25?30°C/10?18°C (day/night). High humidity (>60%) during the night was provided by covering the benches with plastic film in the greenhouse; during the day, the film was removed. Morphological characteristics of powdery mildew were observed under a light microscope and the shape and number of conidiophores were investigated and measured before inoculation.
The genera and race of harvested powdery mildew were identified by following the protocol described by Chen et al. (2008). A small amount of the propagated fungi was collected with a toothpick and transferred to a 1.5-mL microfuge tube. The harvested mycelia were suspended in 800 μL extraction buffer [200 mM Tris-HCl (pH 8.0), 50 mM EDTA, 1.5 M NaCl, 0.5% PVP, 2% β-mercaptoethanol] and ground with a plastic pestle. The mycelia solution was added to 1 mg·mL-1 1ysozyme and 200 μg·mL-1 proteinase K. After incubation for 30 min at 37°C, 2.5% SDS was added and the tube was heated to 65°C for 20 min. Supernatant, obtained by the addition of one-tenth volume of 5M potassium acetate (pH 5.2) and incubation on ice for 10 min, was added to 500 μL isopropanol. The DNA was pelleted at 12,000 × g for 20 min. After airdrying, the pellet was dissolved in 100 μL TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0).
The internal transcribed spacer (ITS) of rDNA was amplified using PN23/PN24 (Bardin et al., 1999). To identify P. xanthii, G. cichoracearum, and Leveillula taurica, three primer sets were used for PCR as previously reported; S1/S2, G1/G2, and L1/L2, respectively (Chen et al., 2008). The PCR mixture contained 1× ExTaq buffer (Takara Co., Japan) including 2 mM MgCl2, 100 μM dNTP, 0.24 μM primer each, 1U ExTaq polymerase (Takara Co., Japan), and different concentrations of fungal DNA template. Samples of 0.5, 1, and 5 ng purified DNA, and 10 and 24 conidia diluted for direct PCR were prepared as templates for each reaction. PCR amplification was conducted with a thermal cycler (AB 2700, PerkinElmer Inc., USA) using the following steps: denaturation at 95°C for 2 min, followed by 35 cycles of 20 s at 94°C, 30 s at 62°C, and 30 s at 72°C, with a final extension at 72°C for 5 min. PCR products were separated using 1.2% agarose gel. The ITS sequences were determined by Sanger sequencing (Solgent Inc., Daejeon, Korea) after TA-cloning of those amplicons. Resulting ITS sequences were compared to previously reported sequences of four races of P. xanthii (GenBank Accession No. AB774155 for race 1; AB774156 for race 3; AB774157 for race 4; and AB774158 for race 2) and two isolates of G. cichoracearum (EU233820 and JQ010848) using multiple sequence alignment software, ClustalW (http://www.genome.jp/ tools/clustalw/).
Inoculum was prepared by rubbing infected leaves with a soft brush and rinsing with distilled water; this water was then collected and used as inoculum. Differential hosts for race identification and segregated progeny were mechanically inoculated by spraying suspensions with 2 × 105 conidia·mL-1 and 2 × 106 conidia·mL-1, respectively. Cotyledons and the first true leaves of seedlings were inoculated at 1-week intervals for 3 weeks.
Disease Assessment
The leaves of individual plants were rated on a 0 to 9 scale for disease severity 25 days after the first inoculation (Tetteh, 2008) in order to describe the degree of lesion and mycelia coverage of the leaf. The disease index (DI) was rated as follows: 0, no symptoms; 1, faint yellow specks on leaves; 2, chlorotic lesions on leaves; 3, chlorotic lesions covering 20% of leaves; 4, yellow chlorotic lesions on leaves turned to brown necrotic areas; 5, 2?3 healthy colonies of mycelium on leaves; 6, less than 20% mycelium coverage on leaves; 7, 20?50% mycelium coverage on leaves; 8, 50?70% mycelium coverage with large necrotic areas, 9, all leaves fully covered with powdery mycelium or the plant is dead. At 0?5 on the scale, the infection did not progress further (resistant plants), whereas at 6?9, the infection progressed (susceptible plants).
Genetic Analysis
Resistance and susceptibility to P. xanthii 1W-AN was analyzed using the P1, P2, BC1P1, BC1P2, and F2:3 populations. The goodness-of-fit of the observed and expected segregation ratios in the populations was evaluated using a Chi-square test.
Molecular Marker Development
Leaves were individually sampled and stored at -80°C in a deep freezer until DNA extraction. To develop molecular markers associated with resistance to P. xanthii 1W-AN, DNA samples from each of eight NILs were isolated and pooled in equal amounts. Genomic DNA was extracted from each plant sample using a modified CTAB method (Stewart and Via, 1993). The DNA of susceptible samples from the ‘SBB’ inbred line, resistant samples from PI 254744, and pooled DNA of resistant samples from the NILs were analyzed using the RAPD method. In total, 600 UBC primers (10-mer) were used for RAPD analysis by following previously described methods (Kim et al., 2013). PCR-amplified bands of the same size were eluted from an agarose gel using GeneClean® Turbo Kit (MP Biomedicals, Inc., USA) for cloning. Eluted DNA was clonedinto the pGEM-T Easy Vector (Promega, USA) and subjected to Sanger sequencing at Solgent Inc. (Daejeon, Korea). Based on the obtained DNA sequences, primers were designed by Primer3web (http://primer3.ut.ee/) and the sequences were aligned to the reference sequence of watermelon 97103 (http://www.icugi.org) using the BlastN procedure to index the chromosomal location of candidate regions showing the highest e-value. SCAR markers were produced by confirming the expected size and the markers were individually genotyped with each of eight NILs and segregated progeny. To develop a cleaved amplified polymorphic sequence (CAPS) marker representing codominance, restriction enzyme sites were investigated in the reference genome of watermelon, including the sequence of the SCAR marker.
A high resolution melt (HRM) marker was also developed by aligning the sequence of the developed SCAR marker with the reference genome of watermelon. PCR products ranged in size from 80 to 120 bp, and the annealing temperature was set at 60°C. Three primer pairs harboring SNPs were used in a PCR-coupled HRM assay on a LightCycler® 96 (Roche Life Science, Inc., USA). The HRM assays were performed using 10-μL reactions containing 1× LightCycler® 480 High Resolution Melting Master (Roche Life Science), 3 mM MgCl2 (Roche Life Science), 0.5 μM of each primer, and 1 ng of dsDNA template. The PCR conditions were as follows: preincubation at 95°C for 10 min, followed by 55 cycles of 30 s at 94°C, 30 s at 60°C, and 30 s at 72°C. A preconditioning step was performed for DNA melting of 60 s at 95°C, 60 s at 40°C, and 60 s at 65°C following the PCR stage, and HRM was then performed at 65?97°C, with a temperature increment of 0.07°C per second; dsDNA melting was detected using 15 readings per second. HRM data were analyzed using the LightCycler® 96 SW 1.1 program (Roche Life Science, Inc., USA), and the sizes of all amplified products were confirmed using 1.5% agarose gel electrophoresis.
Results
Identification of the Causal Agent of Powdery Mildew
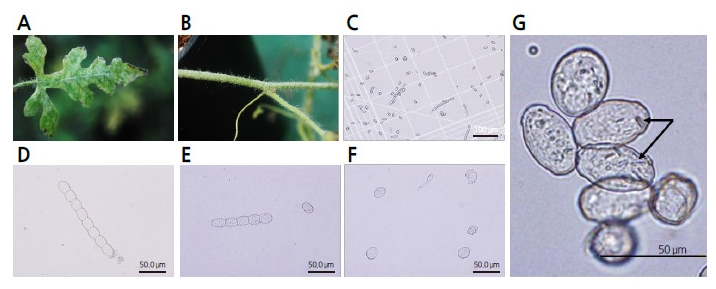
White colonies of fungi naturally occurring on the leaves and stems of the ‘SBB’ watermelon were collected in the greenhouse at Chung-Ang University and used to characterize the morphology of powdery mildew (Fig. 1 A, B) under a light microscope (Fig. 1 C, D, E). The samples of powdery mildew possessed hyphae consisting of appressoria and conidiophores in the euoidium type. Conidia were shaped like ellipsoidal-ovoids and presented fibrosin bodies, which are typically found in P. xanthii, and not in G cichoracearum. The diameter of the mature conidia ranged from approximately 30?45 × 18?25 μm, and the average index from the length/width ratio of conidia was close to 1.67?1.78 (Fig. 1 F, G).
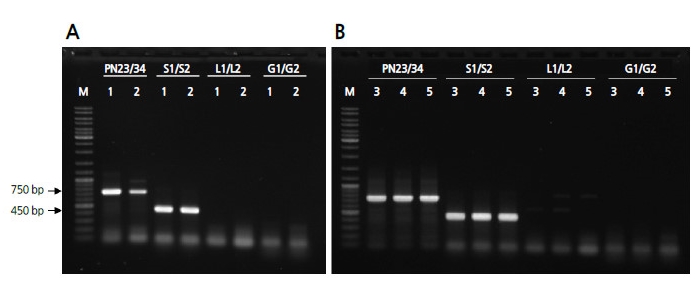
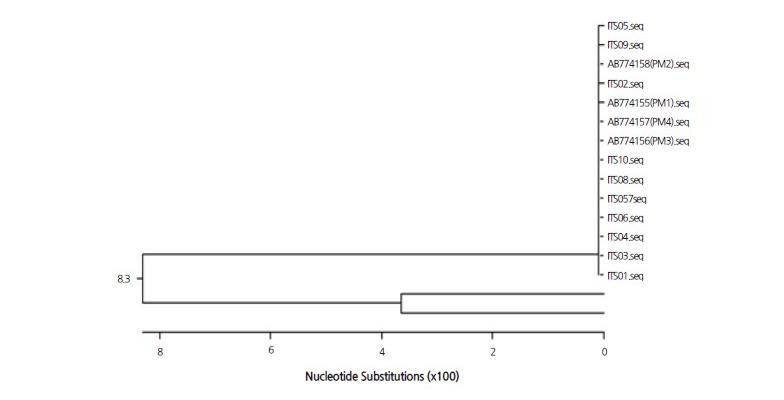
To identify the causal pathogen of powdery mildew on a molecular level, DNA extracted from powdery mildew was amplified and the ITS regions of representative cucurbit-infecting fungi were detected using PCR with previously reported primer sets as follows: PN23/PN34 specific for Erysiphales as a universal ITS-specific primer to detect the powdery mildew family (Bardin et al., 1999), S1/S2 for P. xanthii, L1/L2 for L. taurica (Chen et al., 2008), and G1/G2 for G. cichoracearum (Chen et al., 2008). PCR analysis revealed that a 750-bp amplicon for Erysiphales and a 454-bp amplicon for P. xanthii were present, but amplicons of 391 bp for L. taurica and 374 bp for G. cichoracearum were absent (Fig. 2). In total, 10 amplicons of P. xanthii from colonies on different leaves of ‘SBB’ watermelon were sequenced. When the identified sequences were compared to those previously reported for four races of P. xanthii (GenBank Accession No. AB774155 for race 1; AB774156 for race 3; AB774157 for race 4; and AB774158 for race 2) and two isolates of G. cichoracearum (EU233820 and JQ010848) using ClustalW analysis, the isolate from this study was determined to be almost identical to the four races of P. xanthii (Fig. 3 and electronic suppl. Fig. 1s).
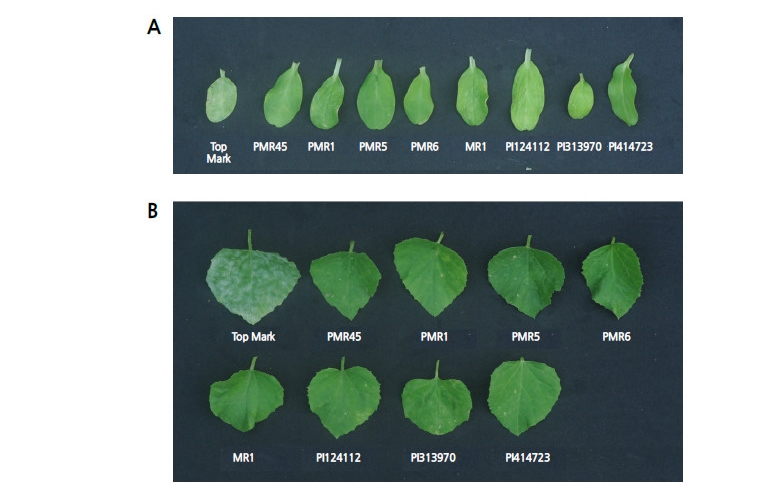
The results of morphological and molecular characterizations revealed the causal pathogen to be P. xanthii. Subsequently, the physiological race was identified using nine differential melon hosts. When ‘Top Mark’, PMR45, PMR1, PMR5, PMR6, MR1, PI 124112, PI 313970, and PI 414723 were artificially inoculated using inoculum derived from P. xanthii, only ‘Top Mark’ was infected in the cotyledon and first true leaf (Fig. 4).
Inheritance of the resistance to P. xanthii race 1W-AN
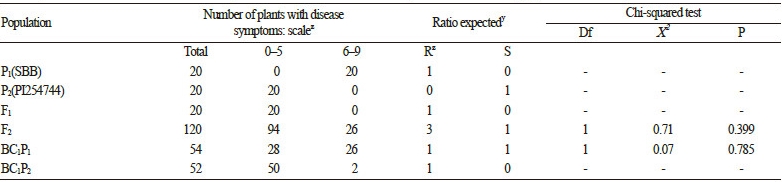
Two inbred lines showing susceptibility (‘SBA’; elite inbred line; P1) and resistance (PI 254744; P2) to the Ansung isolate (AN) of the P. xanthii race 1W were used for the production of F1, F2, and BC1F1 progeny for the study of inheritance. The P. xanthii race 1W-AN was artificially inoculated at the stage of fully expanded cotyledons. The severity of the disease in the cotyledons and the first true leaf was rated 1?3 weeks following inoculation. The parental lines showed disease indexes(DIs) consistent with resistance (DI 0?5) or susceptibility (DI 6?9) at both developmental stages (Table 1). In 120 F2 plants in the segregated population, 3 resistant (R):1 susceptible (S) were found fit a single dominant gene Mendelian model. Furthermore, the test cross of BC1P1 fit to the 1 IR:1 S ratio of the single gene model.
Development of Molecular Markers Linked to Resistance to P. xanthii race 1W-AN
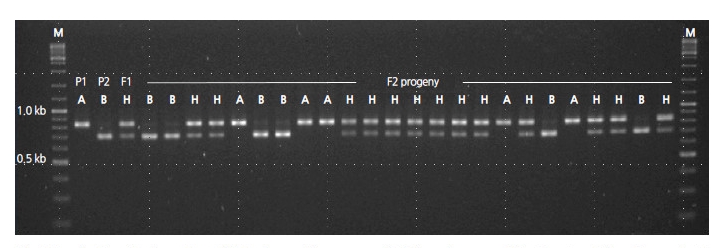
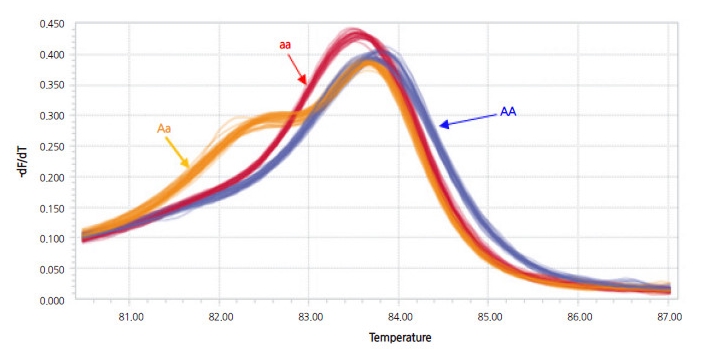
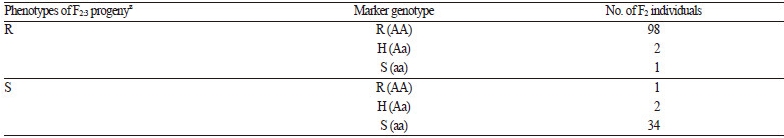
Pooled DNA derived from eight NILs of the BC4F6 generation harboring resistance to powdery mildew was compared to that of the donor (PI 254744) and the recurrent plant (‘SBB’) to develop resistance-linked markers using RAPD analysis. A total of 600 arbitrary decamers were used to identify polymorphisms. Twenty-three polymorphic amplicons absent from susceptible ‘SBB’ plants and present in PI 254744 plants and the bulked NILs of resistant plants were selected (data not shown). Each of the polymorphic bands was eluted from the agarose gel and sequenced in order to convert to SCAR markers. One SCAR marker (PM-SCAR12) produced a single band of 1,042 bp, indicating its presence in all eight resistant plants of NILs and PI 254744. In total, 107 F2 plants were tested with the developed SCAR marker to evaluate consistency with the resistant and susceptible phenotypes. There were discrepancies in the results from four F2 plants (No. 8, 12, 57, and 61 of F2 progeny) in the reaction to P. xanthii 1W-AN in cotyledons and the genotype of the SCAR marker (electronic suppl. Fig. 2s). Two F2 individuals (No. 12 and 57) exhibiting a resistant genotype were strongly infected by the fungi at both developmental stages of the cotyledon and the first true leaf, and the other two F2 individuals (No. 8 and 61) showed the opposite result. Since the dominant SCAR marker could not discriminate hybrid genotypes, codominant CAPS and HRM markers were developed. No appropriate sequences were found to convert CAPS or HRM markers within the amplicon of the SCAR marker; the sequence of the amplicon was therefore aligned to the reference genome of 97103 watermelon (http://www.icugi.org). The candidate regions of the reference genome sequences for CAPS and HRM markers linked to resistance of PI 254744 were scanned up- and down-stream of the PM-SCAR12 locus, ranging from -100 to +100 kbp. In addition, candidate regions that can discriminate parental genotypes were screened for the development of a CAPS markerby digesting many PCR amplicons harboring restriction enzyme sites. Once the population of BC4F6 NIL-1 × ‘SBB’ was genotyped for cosegregation analysis, one of the CAPS markers (254PMR-Nco) containing a Nco I site was selected (Fig. 5). Since HRM markers are relatively cheap and easy to genotype in comparison to other high-throughput (HT) genotyping platforms, we converted 254PMR-Nco to HRM markers. Based on the sequences of the PCR amplicon for the CAPS marker, primer pairs for HRM analysis, which can discriminate C/T SNP (254PMR-HRM3), were constructed and assayed (Fig. 6). To confirm the applicability of the 254PMR-HRM3 marker in another population originated from PI254744, 138 progeny of the F2:3 segregation population (‘SBA’ × PI254744) were tested to confirm cosegregation of the resistant locus of PI254744. Only six of 138 progeny showed a discrepancy between phenotype and genotype with a 4.3-cM linkagedistance observed in the tested F2:3 progeny. The genomic location of the 254PMR-HRM3 marker developed in this study was shown to be close to the locus harboring the resistance (Pmr1) of PI254744 to P. xanthii 1W-AN.
Discussion
Powdery mildew is one of the most predominant foliar diseases of watermelon in South Korea, especially in plants under plastic greenhouse cultivation. Many germplasms from Plant Introductions of the United States have been investigated, and used by breeders as wild-type lines or cultigens harboring resistance to powdery mildew. Since resistance to powdery mildew is also race-specific (Tetteh et al., 2013), resistant lines or cultivars must be developed based on pathogen identification. Two predominant genera of fungi (P. xanthii and G. cichoracearum) are thought to infect cucurbits, including watermelon (Davis et al., 2001; del Pino et al., 2002; Zhang et al., 2011; Aguiar et al., 2012).
Although a few studies have identified the causal fungi of powdery mildew in watermelon in Korea (Lee et al., 2010; Kim et al., 2013; Lee et al., 2014), we attempted to characterize the genera and races of the pathogen using previously reported methods for other crops (Bardin et al., 1999; Chen et al., 2008) based on morphological and molecular characters. All morphological characters of the conidia, including the shape, size, and the presence of fibrosin bodies, matched those of P. xanthii from previous reports (Chen et al., 2008; Cosme et al., 2012; Lee, 2012) (Fig. 1). PCR using previously reported primer sets that discriminate between genera of the Erisiphe family revealed the amplicons of P. xanthii only (Fig. 2). In addition, in a sequence comparison of the amplicons from 10 different fungal colonies derived from different plants, revealed that the ITS sequences contained few SNPs and clustered together with previously reported P. xanthii race 1 (AB774155), race 2 (AB774158), race 3 (AB774156), and race 4 (AB774157), which were determined in melon differentials (Fig. 3). Those sequences were not grouped with two isolates of G. cichoracearum (EU233820 and JQ010848). Based on these results, we concluded that the causative fungi of powdery mildew collected from watermelon are P. xanthii, and not G. cichoracearum, at least in the samples from the greenhouse at Chung-Ang University in Ansung City, South Korea.
Watermelon presumably has the highest resistance to powdery mildew among Cucurbitaceae (Kristkova et al., 2009), and most research on the physiological race identification studies of powdery mildew have been accomplished using melon differentials. To identify the race of the P. xanthii, we used the fungi to inoculate nine melon differentials and three inbred watermelons at the cotyledon expansion stage (data not shown). Only ‘Top Mark’ of the melon differential and other watermelons showed powdery mildew on cotyledons and leaves at the first true leaf stage, whereas the other tested differential hosts, including PI 254744, did not show symptoms of infection (Fig. 4). The pathotype of powdery mildew infectious to watermelons (based on the response to P. xanthii) includes three races (Tetteh et al., 2013), including race 1W (Davis et al., 2007), 2WU.S. (Tetteh et al., 2010), and 2WF (Zhang et al., 2011). Based on those criteria, we observed the reactive ‘Top Mark’ and non-reactive PI 414723 (Fig. 4) hosts and demonstrated this race is neither 2WU.S. nor 2WF. As a result, the causal pathogen of powdery mildew infecting watermelon at Chung-Ang University corresponds to the P. xanthii race 1W, which was also reported by other researchers (Davis et al., 2001; McCreight, 2006; Tetteh et al., 2010). This result corresponds to a report showing that the causal fungus of powdery mildew in watermelon is thought to be P. xanthii race 1 (Kim et al., 2013; Kim et al., 2015). In the present study, we used morphological and molecular methods with data to discriminate between P. xanthii and G. cichoracearum, which are largely predominant in South Korea. To date, we have not detected any genera or races of powdery mildew other than race 1W in Ansung Province of Korea (data not shown).
With the critical identification of P. xanthii race 1, we attempted to develop a molecular marker linked to the resistant region of PI 254744 (C. lanatus var. lanatus), a well-known cultigen in Korea from the US PI watermelon germplasm harboring high resistance to P. xanthii race 1W (Davis et al., 2001; Kim et al., 2013). In the present study, results of the Chisquare test in the BC1 and F2 populations revealed that the resistance gene from PI 254744 was inherited by the progeny as a single dominant trait via Mendelian manner. ‘Arka Manic’, a watermelon germplasm harboring resistance to race 1W, showed incomplete resistance to the pathogen (Kim et al., 2013), and PI 254744 had intermediate resistance to race 2WU.S. (Tetteh, 2008). In an inheritance study of resistance to race 2WU.S. in F2, BC1P1, and BC1P2 populations derived from the cross between ‘Charleston Gray’ and PI 189225, the segregation patterns of leaf resistance trait fit the model for single gene inheritance (Tetteh et al., 2013). In addition, PI 189225 did not show resistance to race 2WF, the predominant race in the watermelon fields in China (Zhang et al., 2008). Thus, the inheritance of powdery mildew resistance in watermelon would depend on the races of powdery mildew and resistant sources of germplasm. More studies on inheritance and alleles between germplasm should be performed to elucidate the resistance genes and to aid the development of markers closely linked markers for resistance against the three races of P. xanthii, which would facilitate gene pyramiding of the resistance trait into elite inbred lines in the future.
Although powdery mildew is known to be an emerging foliar disease of watermelon in the United States (Davis et al., 2001), only two reports have focused on the developing molecular markers linked to powdery mildew resistance in watermelon (Kim et al., 2013; Gama et al., 2015). Kim et al. (2013) demonstrated the incomplete resistance in ‘Arka Manic’ (Rai et al., 2008), which is known to harbor resistance to race 1 of P. xanthii. The authors developed a molecular marker (OP-483) corresponding to a single partially dominant gene (Pm1.1) from ‘Arka Manic’ (resistant cultivar) × HS3355 (susceptible line) via bulk segregant analysis (BSA) of the F2 generation using RAPD. The O-483 was converted to codominant CAPS and HRM markers based on the presence of an SNP. Gama et al. (2015) developed microsatellite markers, MCPI_11, CYSTSIN, and BVWS02441 using the BSA-microsatellite method in an F2 population of ‘BRS Opara’ × ‘Perola’. These markers are presumably linked to a powdery mildew resistance gene at a distance of 2.6 cM.
In summary, we identified the genus and physiological race of a causal pathogen of powdery mildew and clarified that PI254744 carries resistance to P. xanthii 1W-AN. In addition, we successfully developed CAPS and HRM-type molecular markers, 254PMR-Nco and 254PMR-HRM3, for resistance to P. xanthii 1W-AN. The results of this study should facilitate the use of PI254744 for breeding via marker-assisted selection.