Introduction
Materials and Methods
Plant Materials and MAB Population Construction
Selection of the SNP Background Marker Set
trsSCAR Marker Screening and Foreground Selection
SNP Genotyping and Background Selection
Phenotyping Analysis of the Chinese Cabbage Backcross Population
Results
The Selection of Polymorphic SNP Markers and the Genotyping Result of the MAB Population
The Selection Efficiency of Backcrossing Using the MAB Marker Set
Discussion
Introduction
Turnip mosaic virus (TuMV; genus: Potyvirus) infects a wide range of cultivated plants, commonly causing a distinct mosaic of light- and dark-green coloration in the leaves. Other symptoms such as necrotic streaks, flecks,or ring spots may also appear, depending on the virus strain and the crop species. The major hosts of TuMV are the brassicaceous species, in which it causes severe economic losses in terms of crop production in Asia, North America, and Europe (Edwardson and Christie 1991; Shattuck et al., 1992). TuMV is transmitted by aphids in a non-persistent manner (Shattuck et al., 1992). One of its target species, Chinese cabbage (Brassica rapa subsp. pekinensis) is cultivated worldwide, but it most commonly grown in Asia and Europe. The serious loss of yield caused by TuMV means that the development of broad-spectrum resistance to this virus is still an important target for Chinese cabbage breeders.
Dominant TuMV resistance genes have been identified in the Brassicaceae, including TuMV Resistance in Brassica 01 (TuRB01), TuRB03, TuRB04, TuRB05, TuRB07, TuRB01b, Conditional TuMV resistance 01 (ConTR01), and TuRBCS01 (Robbins et al., 1994; Walsh et al., 1999; Jenner et al., 2002; Hughes et al., 2003; Rusholme et al., 2007; Qian et al., 2013; Jin et al., 2014; Liz et al., 2015). Recessive resistance genes, Recessive TuMV resistance 01 (retr01), retr02 and trs (TuMV resistance discovered at Seoul National University), have also been identified in Brassica rapa (Walsh et al., 2002; Kim et al., 2013). Among these resistance genes, trs was found to provide broad-spectrum resistance against several TuMV strains (Kim et al., 2013).
The purpose of backcross breeding is to transfer a specific allele at the target locus from a donor line to a recipient line. The fundamental theory of the traditional backcrossing system is that the proportion of the recurrent parent genome (RPG) is recovered at a rate of 1 – (1/2)t+1 for each backcrossing generation (Babu et al., 2004); thus, the time taken to achieve the appropriate number of generations and the linkage drag observed are the two major issues to resolve in the traditional backcrossing system (Semagn et al., 2006). Marker-assisted backcrossing (MAB) is considered an innovative tool for the effective introgression of a single targeted locus in plant breeding, while retaining the important characteristics of the recurrent parent (Hospital and Charcosset 1997; Hospital 2001; Collard and Mackill 2008). MAB can remarkably accelerate the recovery speed of the recurrent parent genome. The use of molecular markers that permit the genetic dissection of the progeny at each generation increases the speed of the selection process, thus increasing the genetic gain per unit time (Tanksley et al., 1989; Hospital 2003).
The MAB system enables the recovery of the recurrent parent genotype in only two or three backcrosses. The efficiency of MAB is known to be dependent on the experimental design, most notably on the marker density and position, population size, and selection strategy (Frisch et al., 2000; Frisch et al., 2005). MAB is available only when there is a reference genome from which useful markers can be developed, however, meaning few vegetable crop breeders have attempted to use this method to date. Many of the studies introducing MAB systems have been those examining field crops, such as rice (Oryza sartiva), wheat (Triticum aestivum), barley (Hordeum vulgare), peanut (Arachis hypogaea), and millet (Pennisetum glaucum; Liu et al., 2000; Jefferies et al., 2003; Neeraja et al., 2007; Koilkonda et al., 2012; Soto-Cerda et al., 2013; Vishwakarma et al., 2014; Lee et al., 2015). Simple sequence repeat (SSR) markers were used in most of these cases. MAB systems for multiple gene introgressions and QTL introgressions have also been used in several crops as maize, sweet corn, bean, rice, wheat and chickpea (Frisch et al., 2001; Yousef et al., 2002; Miklas et al., 2007; Iftekharuddaula et al., 2011; Salameh et al., 2011; Cuc et al., 2012; Khanh et al., 2013; Taran et al., 2013; Hao et al., 2014; Varshney et al., 2014; Mallick et al., 2015).
In this study, a high-throughput (HT) marker system was used to develop a MAB system in Chinese cabbage. The 468 previously developed single-nucleotide polymorphism (SNP) probes (Brassica Marker Assisted Backcrossing SNP 1 [BraMAB1]), designed for the Fluidigm system, were applied in this study. We selected the broad-spectrum TuMV-resistant Chinese cabbage line SB22 as a donor plant and crossed it with the TuMV-susceptible 12mo-682-1 elite line, determining that the HT marker system is highly efficient in the B. rapa backcross breeding system. This is the first report of the application of a SNP marker set in the background selection of Chinese cabbages using HT SNP genotyping technology.
Materials and Methods
Plant Materials and MAB Population Construction
The Chinese cabbage (B. rapa subsp. pekinensis) backcross population BC1F1 was constructed by crossing the SB22 TuMV-resistant donor line (Kim et al., 2013) with the recurrent parent, 12mo-682-1, an elite cultivar provided by the Biobreeding Research Center, Ansung, Korea (Table 1). The 12mo-682-1 line produces a closed head with yellow inner leaves, and displays strong heterosis, late flowering, and early maturity, which are all suitable characteristics for kimchi production. This accession is susceptible to TuMV. The SB22 produces half-opened heads. The BC1F1 plant with the highest RPG recovery rate was backcrossed to 12mo-682-1 to construct the BC2F1 population. The MAB scheme is described in Figure 1.
Selection of the SNP Background Marker Set
To develop the SNP marker set for the background selection of the BC1 and BC2 populations, previously developed SNP markers, BraBSM (Brassica Background Selection Markers), were used to screen for polymorphisms between the donor and the recipient plant (Kim et al., 2016). The 468 BraBSM SNP markers are evenly distributed throughout all ten chromosomes (Kim et al., 2016). Among these markers, 119 SNPs which showed polymorphism between the donor and the recipient plant were selected for further analysis (Suppl. Table 1s). The markers showing no polymorphism or an inconclusive result were eliminated. Linkage maps of SNPs used in the backcross selection of this study were constructed using the results of the BC1F1 genotyping (Figure 3) using Carthagene software (de Givry et al., 2005).
trsSCAR Marker Screening and Foreground Selection
The total genomic DNA was isolated from the young green leaves of Chinese cabbage using the cetyl trimethylammonium bromide (CTAB) method described by Hwang et al. (2009), and its quality was estimated using a NanoDrop spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). To screen for TuMV resistance in the backcross population, the trsSCAR (trs sequence characterized amplified region) marker (three marker primers) was used based on the modified protocol of Kim et al. (2013). A PCR analysis was performed using a total reaction volume of 25 μL, containing 100 ng of template genomic DNA, 2.5 μL 10× rTaq Buffer, (Mg2+ plus, Takara Shuzo Co., Ltd., Shiga, Japan), 2.5 mM dNTP, 2.5 U of Takara rTaq (Takara Shuzo Co., Ltd.), 4 pmol of primer F1, 3 pmol of primer F2, and 5 pmol of primer R (Kim et al., 2013). The PCR conditions comprised 37 cycles at 94°C for 30 s, 63.5°C for 30 s, and 72°C for 1 min 40 s.
SNP Genotyping and Background Selection
The total genomic DNA of 75 BC1F1 plants and 169 BC2F1 plants was extracted and checked as described above. Genotyping analysis using the Fluidigm system (IFC controller MX and EP1 system; Fluidigm Corp., San Francisco, CA, USA) was performed by the Foundation of Agriculture Technology Commercialization and Transfer (FACT) in Korea.
Phenotyping Analysis of the Chinese Cabbage Backcross Population
The BC1F1 population was phenotyped 60 days after transplant. The total weight, head weight, head height, and maximum head width were quantified, while the head type, outer leaf color, and internal leaf color were visually estimated. These seven traits are typically bred for in Chinese cabbage crops produced in Korea. Photographs of the vertical aspect of each plant were taken during the phenotyping process.
Results
The Selection of Polymorphic SNP Markers and the Genotyping Result of the MAB Population
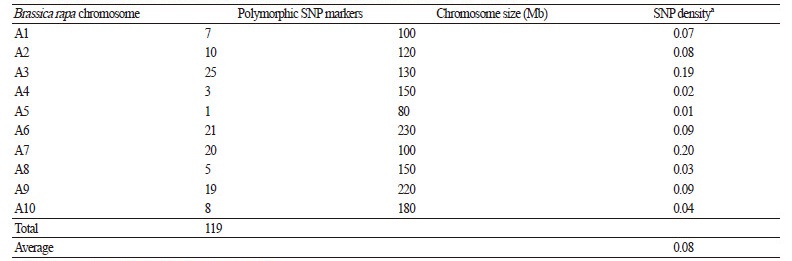
A total of 119 SNPs were identified from the BraBSM SNP markers between the donor and recurrent parents, and were used for the background selection of the BC1 and BC2 populations. The numbers of polymorphic SNPs per Chinese cabbage chromosome are shown in Table 2. The range of SNP densities was between 0.01 and 0.20 SNPs·Mbp-1, with an average SNP density of 0.08 SNPs·Mbp-1. The SNP density of chromosome A07 was the highest, and that of chromosome A05 was the lowest.
To preserve the TuMV resistance allele in the backcrossing procedure, foreground selection was performed using the trsSCAR marker, which was previously developed as an indicator of TuMV recessive resistance (Kim et al., 2013). The polymorphisms of the TuMV-resistant donor parent (SB22) and the TuMV-susceptible recurrent parent (12mo-682-1) plants were first confirmed using the trsSCAR marker. The 75 BC1 plants were then screened for their genotypic segregation of this trait (Figure 2), resulting in the identification of 33 plants with a heterozygote genotype and 42 plants with a homozygote genotype (Table 3). Because of the recessive inheritance pattern of TuMV resistance, heterozygous plants were selected for the background selection.
Fig. 2
Example of BC1 foreground selection using the trsSCAR marker. A total of 75 individuals were genotypically screened using the trsSCAR marker. ‘SB22’ and ‘12mo’ refer to the SB22 (donor parent) and 12mo-682-1 (recurrent parents) lines of Chinese cabbage, respectively. M: DNA marker; H: Heterotype genotype; S: susceptible genotype.
TThe RPG rates in the BC1 population ranged from 54.6% to 90.3% (Table 3). A total of 18.2% of the 75 BC1F1 plants showed an RPG rate above 80%, 68.8% had an RPG rate above 70%, and only one plant had an RPG rate above 90%. The BC1F1 plant with the highest recovery rate (90.3%) and with the heterozygous trs genotype was chosen to backcross to the recurrent parent to generate the next generation, BC2F1.
Fig. 3
Linkage map of SNPs constructed using the BC1F1 genotyping result. The SNP probes are named on the right side of the map. The genetic distance (cM) is shown on the left. The ten chromosome numbers shown in this map are based on the physical position (kb) in the Brassica rapa reference genome sequence. The linkage map was generated using Carthagene software (de Givry et al., 2005).
A total of 169 BC2F1 plants were screened for the genotypic segregation analysis. A foreground selection was also performed using the trsSCAR marker. Of these, 108 plants had a heterozygous genotype, and 61 plants were homozygotes (Table 3). According to the screening data, the result was slightly biased to the heterozygote genotype. Of the 169 BC1F1 plants, 39.1%
had an RPG rate above 80%, and 85.2% had an RPG rate above 70%. The percentage of the genome-recovered plants (≥70% RPG rate) in BC2F1 was clearly increased compared with the BC1F1 population.
These results show that the selected SNP marker set is useful for the efficient backcross breeding of Chinese cabbage. The RPG rates in the BC2 population ranged from 58.4% to 93.3%. Among the BC2F1 plants, four plants had an RPG rate above 90%. This study shows that using MAB breeding, over 90% of the background genome can be recovered in only two generations.
The Selection Efficiency of Backcrossing Using the MAB Marker Set
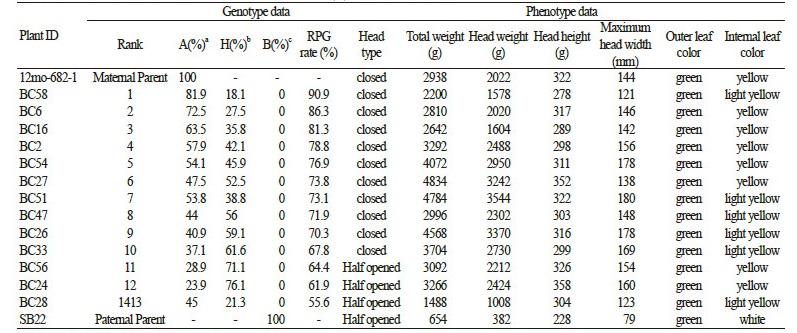
To analyze the efficiency of the MAB background marker set, the genotype data of BC1F1 plants were compared with their phenotypic data. A total of 13 plants were selected, representing RPG rates ranging from 55.6% (plant BC28) to 90.9% (plant BC58). The change in RPG rate between any two individuals was 0.7-6.3% (Table 4).
The phenotype data of the selected BC1F1plants are shown in Table 5 and Figure 4. Seven traits (head type, total weight, head weight, head height, maximum head width, outer leaf color and internal leaf color) were determined. According to the phenotype data, the head type trait showed a clear difference as the RPG rate changed, with the half-opened head type of the donor plant SB22 observed in the plants with an RPG rate < 65%. The density of the leaves, an indicator of maturity, also decreased in the plants with a lower RPG rate, similar to the SB22 donor. The quantitative analysis of total weight, head weight and width indicate that the plant (BC58) with the highest RPG rate has a strong phenotypic similarity to the recurrent parent 12mo-682-1. There was no difference in the head height and the outer leaf colors among the BC1F1 plants.
The tolerance of the various cultivars is presented in Table 3. Cultivars 1-5, 7, and 9-12 were tolerant to chilling stress at the germination and seedling stages, while cultivars 11 and 14 were tolerant to salinity stress, and cultivars 1-5, 7-10, 12, 13, and 15 were tolerant to the combined stress treatment. None of the cultivars were tolerant to all three stress conditions for both growth stages. Cultivar 11 did not exhibit tolerance to the combined stress treatment even though it was tolerant to the individual stress treatments (i.e., chilling and salinity stresses) at the germination and seedling stages.
Discussion
Although background selection has become a standard tool in plant breeding, the high cost of marker analysis still limits its use in practice and restricts the experimental design of gene introgression programs (Collard and Mackill, 2008). Here, we developed a Chinese cabbage MAB system using a HT genotyping method, the first time that this technique has been reported in this crop. We constructed the BC1 and BC2 Chinese cabbage populations to introduce TuMV broad-spectrum recessive resistance into the elite line, 12mo-682-1. The 119 polymorphic SNPs between the donor and recurrent parents identified from the previously developed BraMAB1 SNP probes were applied for the background selection, with foreground selection performed using the previously developed trsSCAR marker. We found that 68.8% of 75 BC1F1 plants had a high RPG rate of over 70%, while more than 80% of the BC2F1 plants showed this high RPG recovery rate. This study shows that the use of MAB breeding could lead to plants containing over 90% of the background genome in only two generations. We concluded that the developed MAB marker system is highly efficient in the B. rapa backcross breeding system.
Fig. 4
The phenotype of selected BC1F1 individuals with varying rates of recurrent parent genome (RPG) recovery. The photographs of the vertical aspect of each individual were taken 60 days after transplant. The recurrent parent is the ‘12mo- 682-1’ elite line, and the donor parent is the ‘SB22’ plant. Thirteen BC1F1 plants are shown in this picture in order of RPG rate, from the plant with the highest RPG rate (BC58) to the plant with the lowest (BC28).
We also find that the background genotyping result and the phenotypic similarity between the recurrent parent and BC1F1 showing a correlation in the traits related to head. While differences in the internal leaf color were observed between the SB22 donor and the other BC1F1 plants, this trait did not appear to be clearly associated with the background genotype. There was a report about the gene explaining yellow and orange color on the inner leaf of the Chinese cabbage (Feng et al., 2012). We concluded that the further analysis may be needed to study the inner leaf color variation.
To date, few studies have reported the use of SNP data in MAB systems for vegetable crops; tomato (Solanum lycopersicum), cucumber (Cucumis sativus), and pepper (Capsicum annuum) are the only cases in which a MAB system has been applied (Behera et al., 2011; Hwang et al., 2012; Jeong et al., 2015). In tomato, 108 SSR primer sets were selected from the tomato reference genetic maps for identifying introgressions of the recurrent parent genome (Hwang et al., 2012). In cucumber, to increase the genetic diversity of breeding materials, the marker-assisted strategy was performed using SNP, SSR, and SCAR markers through introgression backcrossing (Behera et al., 2011). Recently, MAB based on the high-throughput SNP genotyping method using the Fluidigm EP1 system was demonstrated to introduce a high level of capsinoid content in pepper (Jeong et al., 2015).
In our previous study, we generated a useful B. rapa SNP database by sequencing the transcriptomes of 20 Chinese cabbage accessions (Kim et al., 2016). Searching for expressed sequence tag (EST)-derived SNPs through transcriptome sequencing has been found to be much more attractive than performing whole-genome sequencing to discover genes based on a molecular marker source (Kim et al., 2008). The present study is the first report of the application of a HT marker system to Brassica rapa backcross breeding.
MAB systems can be divided into a single-marker (SM) and a HT marker systems (Herzog et al., 2011). The SM system primarily uses a marker type, such as the SSR marker, and restriction fragment length polymorphisms, while the HT marker system is based on SNPs, a cutting-edge technology in molecular breeding (Herzog et al., 2011). HT MAB is a much more useful tool due to its low cost and the ability to automate genotyping technologies. Another important advantage of the HT marker systems is that they can provide linkage maps with equally spaced markers (Herzog et al., 2011).
In this study, we only used 119 SNP probes showing polymorphism between two parents for the genotype back selection. It can be assumed that the genetic gap of the two parents is not large, despite the differences in their phenotype. In a previous study in C. annuum, the MAB system was successfully used with only ~200 SNP markers (Jeong et al., 2015). Our result demonstrates that a SNP marker set of less than 150 polymorphisms can also play an effective role in selecting suitable individuals for Chinese cabbage backcross breeding.
In this study, we only used 119 SNP probes showing polymorphism between two parents for the genotype back selection. It can be assumed that the genetic gap of the two parents is not large, despite the differences in their phenotype. In a previous study in C. annuum, the MAB system was successfully used with only ~200 SNP markers (Jeong et al., 2015). Our result demonstrates that a SNP marker set of less than 150 polymorphisms can also play an effective role in selecting suitable individuals for Chinese cabbage backcross breeding.
The highest RPG rate obtained in the present study after the two backcross generations was 93.3%, indicating that 6.7% of the genome still needs to be recovered. Constructing a BC3 generation may be necessary to achieve a higher RPG rate in Chinese cabbage; however, the number of backcrossing steps required for different studies may change depending on the breeding objectives and available genomic materials; for example, the efficiency of the MAB system could be influenced by the combination of different accessions. Further study into the application of this MAB system to the various Chinese cabbage backcross populations is necessary to develop a widely applicable marker system.