Introduction
Climate Change: An Important Concern
Overview of Metabolic Profile Changes under Stress Owing to Climate Change
Comprehensive Influence of Ecological Factors on Plant Secondary Metabolite Biosynthesis
Metabolic Engineering of Plant Secondary Metabolites
Functional Genomics of Plant Secondary Metabolites
Transcription Factors and Genes Related to Plant Secondary Metabolites
WRKY Transcription factors
MYB Transcription factors
bHLH Transcription factors
bZIP Transcription factors
Effects of Genetic Factors on the Biosynthesis of Metabolites
Transcription factors involved in the biosynthesis of alkaloids
Transcription factors involved in the biosynthesis of phenolic acids
Transcription factors involved in the biosynthesis of flavonoids
Conclusions
Introduction
Metabolites produced by horticultural plants are crucial for environmental adaptability despite the fact that they are not directly involved in growth, development, or reproduction (Holweg et al. 2025). These substances have a variety of chemical configurations and biological roles; they attract pollinators, control growth and development, and provide a defense against infections and herbivores (Erb and Kliebenstein 2020; Naamala and Smith 2020). It is essential to understand the genetic foundations of secondary metabolite production and their ecological roles in fostering sustainable agriculture and protecting natural ecosystems. The genes responsible for secondary metabolite production contribute significantly to plant adaptation under environmental stress. Plant secondary metabolites were once commonly believed to be tiny biological molecules that weren’t necessary for an organism to survive (Clemensen et al. 2020). It is now widely acknowledged, nonetheless, that plant secondary metabolites are crucial for environmental adaptation because they shield organisms from environmental stress and promote the development of symbiotic relationships (Zhan et al. 2022). Various secondary metabolites have been classified as phenolics, flavonoids and terpenoids, among other classes (Table 1).
Table 1.
Various plant secondary metabolites secreted by different horticultural plants
| Secondary metabolites | Examples | Plant species | References |
| Phenolics | Chlorogenic acid | Lonicera japonica | Wang et al. 2020 |
| Chlorogenic acid | Magnolia officinalis | Hyeon et al. 2021 | |
| Chicoric acid | Echinacea purpurea | Salmanzadeh et al. 2020 | |
| Coffee quinic acids | Coffee | Pinto et al. 2021 | |
| Dicoffee quinic acids | Coffee | Bachheti et al. 2021 | |
| Eugenol | Cinnamomum verum | Gao et al. 2020 | |
| Magnolol | Magnolia officinalis | Niu et al. 2021 | |
| Quinic acid | Coffee | Alcázar Magaña et al. 2021 | |
| Total phenolic | Astragalus compactus | Nurzyńska-Wierdak 2023 | |
| Flavonoids | Baicalin | Scutellaria baicalensis | Pei et al. 2022 |
| Flavonoids | Scutellaria baicalensis | Xu et al. 2020b | |
| Hyperin rutin and quercetin | Magnolia officinalis | Zeru 2023 | |
| Luteolin | Lonicera japonica | Guan et al. 2023 | |
| Total flavonoids | Scutellaria baicalensis | Li et al. 2023 | |
| Terpenoids/ Essential Oils | Camphor, 1,8-cineole | Achillea millefolium | Far et al. 2023 |
| Essential oils | Cinnamomum cassia; Cinnamomum verum | Werrie et al. 2021 | |
| Essential oils | Magnolia zenii | Bharti et al. 2023 | |
| Essential oils | Citrus medica | Bordoloi et al. 2022 | |
| Myrcene, Ocimene | Antirrhinum majus | Terry López 2020 | |
| Triterpene | Codonopsis pilosula | Wang et al. 2023 | |
| Others | Azulene | Achillea millefolium | Satari et al. 2021 |
| Oleanolic acid, Ecdysterone | Achyranthes bidentata | Liu et al. 2020b | |
| Saponins | Panax notoginseng | Zhang et al. 2020b | |
| Trans-cinnamaldehyde | Cinnamomum cassia | Werrie et al. 2022 |
Environmental conditions such as heat, cold, and drought put plants under continual stress, which can harm crop quality and yields. Plants use a variety of strategies to reduce the harm caused by stress, such as adjusting gene expression and plant secondary metabolites synthesis (Jan et al. 2021; Anas et al. 2025a). Alfalfa plants activate an antioxidative mechanism to scavenge reactive oxygen species under biotic and abiotic forms of stress (Anas et al. 2025a). Because poplars (Populus spp.) have more lignin, they have a greater ability to withstand heat. Furthermore, transcriptome and RT-qPCR analyses revealed that key genes (PtrMYBs, PtrC3’H1, PtrCCR2) are highly expressed under high temperatures, indicating their role in lignin accumulation and offering insights for improving lignin quality and for more efficient resource utilization (Zhao et al. 2022). Likewise, poplar plants increase the amount of lignin accumulated in their stems to tolerate high-temperature stress through higher production levels of caffeate and conifer aldehyde (Movahedi et al. 2021). Similar to this, sweet potatoes (Ipomoea batatas (L.) Lam.) have several B-box transcription factor genes, including IbBBX28. Additionally, an adverse correlation between IbBBX28 and plant resilience to water deficiency was investigated. This lays the foundation for future studies on the role of B-box transcription factors in enhancing stress resilience in sweet potato (Gao et al. 2023).
For a detailed understanding of how plants control the manufacturing of their secondary metabolites and molecular processes analysis behind the procedure, it is imperative to explore functional genes. The molecular mechanisms regulating the synthesis of monoterpenes and sesquiterpenes in Cinnamomum burmannii were investigated, with a focus on their roles in the composition of essential oil (Xu et al. 2023). It was discovered that C. burmannii is the source of essential oil, a finding that led to an investigation of the genes and lncRNAs specifically regulated during the production of monoterpenoids and sesquiterpenoids (Hou et al. 2023). Additionally, scientists provided a fresh perspective on the workings of the plant secondary metabolites produced in C. burmannii. Additionally, it was shown that several WRKY transcriptional factors play a subcellular role in protoberberine production alkaloids in Coptis chinensis Franch (Liu et al. 2021a). This is significant when seeking to understand climatic changes and adaptations in plants at the genomic level for agricultural sustainability.
Climate Change: An Important Concern
The existence of plants is being threatened by climate change, which is a severe issue that is present around the world. Environmental variables, such as weather conditions and biological factors, influence a plant’s secondary metabolites. Therefore, any alterations to these circumstances may have impacts on the production of various therapeutic species (Khare et al. 2020). Furthermore, billions of people rely on herb-based medications for treatment and cures, even in the presence of the widespread use of manmade and organic-based pharmaceuticals. Plants continue to be among the most important sources of medicine. Currently, numerous natural compounds are being investigated for their alleged capacity to lower illness risk and promote healthy aging. As a result, several plant species are very important, and numerous plant parts can provide a diversity of secondary metabolites (Srivastava et al. 2021).
An estimated four billion people, or 80% of the global population, reside in underdeveloped nations where traditional medicine and pharmaceuticals are their primary sources of healthcare (Qinsheng and Ginting 2022). The World Health Organization estimates that over 21,000 species of plants, totaling approximately 45,000 distinct varieties, are used for medicinal purposes. Many chemicals commonly utilized in the food, cosmetic, and pharmaceutical industries, such as terpenoids, phenols, steroids, flavonoids, tannins, and aromatic compounds, can be derived from medicinal plants (Pant et al. 2021). These chemicals, often referred to as plant-based secondary metabolites (e.g., specialized products), are thought to be defense compounds that help the plant interact with its surroundings and adapt. They are not necessary for the plant’s progression and expansion. Discoveries thus far include approximately 2,140,000 secondary metabolites, classified into many types (Pant et al. 2021). Due to the great chemical variety of these plants as well as the ability to generate novel medicines with low side effects and commercial value, the demand for medicinal plants has expanded significantly in the last era. Numerous studies have examined several classes of secondary metabolites as possible therapeutic or prophylactic agents for several major illnesses or conditions, including diabetes, cancer, Alzheimer’s disease, cardiovascular disease, ulcers, diabetes, asthma, and Parkinson’s disease (Adetuyi et al. 2022; Thirugnanasambantham et al. 2023).
In addition to vital contributions to many facets of human existence, vegetation is impacted by a diversity of ecological issues throughout its growth and development, including changes in temperature, light intensity, high carbon dioxide levels, UV radiation, ozone, drought, salt, and floods. These ecological elements are significant because different vegetal species require different ecological circumstances to develop (Mahajan et al. 2020). In tandem with a rise in CO2 to 407.4 parts per million, the mean annual temperature is rising at a quicker pace of 0.06–0.1°C/year. Changes in the climate have an influence on plants’ physiology along with their typical behaviors, which in turn has an impact on secondary metabolite production (Ofiti 2022). Research has indicated that certain environmental conditions, such as high CO2 levels, high temperatures, ozone, UV radiation, and drought, harm plant metabolism, growth, and productivity by changing metabolites (Table 2). These studies, however, are frequently patchy and currently contain insufficient data for a proper comprehension of the precise function that each environmental component plays in the production and buildup of secondary metabolites (Naamala and Smith 2020). Numerous individuals as well as entire communities are greatly impacted by plant species, which also have substantial economic worth. We can therefore hypothesize that certain plants, which are important sources of bioactive compounds, may be influenced by various factors that regulate the synthesis and accumulation of secondary metabolites (Yadav and Siddiqui 2023). Such influences can create challenges related to deficiencies in metabolite production, as well as complications during extraction, purification, and removal of contaminants in the development of medicinal products. Important variations related to weather change and environmental alterations can arise during the production of secondary metabolites (Ahmadi-Lahijani and Moori 2022).
Table 2.
Various plant species for which metabolite synthesis is affected by climatic change
| Plant species | Climatic factor | References |
| Asparagus aethiopicus | Salinity | Punetha et al. 2022 |
| Brassica napus | Drought | Jabeen et al. 2021 |
| Brassica oleracea | UV light | Sathasivam et al. 2023 |
| Carthamus tinctorius | Salinity | Kim et al. 2021 |
| Caryopteris mongolica | UV light | Nocchi et al. 2020 |
| Chrysanthemum morifolium | Drought | Yeloojeh et al. 2020 |
| Cucumis sativus | Drought | Abdelaziz et al. 2021 |
| Cuminum cyminum | UV light | Kamel et al. 2022 |
| Cynara cardunculus | Salinity | Pappalardo et al. 2020 |
| Kalanchoe pinnata | UV light | Fkiara et al. 2020 |
| Lactuca sativa | Drought | Ouhaddou et al. 2023 |
| Lotus japonicus | Drought | García-Calderón et al. 2020 |
| Mentha piperita | Salinity | Assaf et al. 2022 |
| Nicotiana tabacum | Drought | Begum et al. 2021 |
| Ocimum basilicum | Salinity | Saia et al. 2021 |
| Ocimum spp. | Drought | Mulugeta and Radácsi 2022 |
| Oryza sativa | Drought | Yadav et al. 2021 |
| Salinity | Shahzad et al. 2021 | |
| Red pepper | Salinity | Zamljen et al. 2022 |
| Ribes nigrum | UV light | Rizwana et al. 2022 |
| Solanum Lycopersicon | Salinity | Abdel-Farid et al. 2020 |
| UV light | Pizarro-Oteíza and Salazar 2022 | |
| Thymus vulgaris | Drought | Szabo et al. 2022 |
| Triticum aestivum | Salinity | Khan et al. 2022 |
| UV light | Semenov et al. 2020 | |
| Drought | Yadav et al. 2021 | |
| Vigna radiata | UV light | Gamit and Amaresan 2023 |
| Vitis vinifera | Drought | Savoi et al. 2020 |
| UV light | Del‐Castillo‐Alonso et al. 2021 | |
| Zea mays | Drought | Chávez-Arias et al. 2022 |
| Salinity | Shahid et al. 2023 | |
| UV light | Yavaş et al. 2020 |
Henceforth, it is concluded that climate change is threatening the existence of plants, particularly in regions with high biodiversity. Environmental variables, such as climate and ecological concerns, disturb secondary metabolite quantity levels in plants, potentially impacting their production (Wang et al. 2017; Srivastava et al. 2021). Plants serve as a vital source of medicine, with more than half of the worldwide population depending on them for a range of treatments. The demand for medicinal plants has grown significantly over the last era owing to their biochemical variety and potential to contain therapeutic or prophylactic agents that can be used to treat various diseases (Li et al. 2016; Abdallah et al. 2023). Ecological factors, such as high temperatures, sunlight intensity, and climate, also affect plant growth and productivity. Changes in these factors could lead to shortages and other issues related to the manufacturing and decontamination of therapeutic products.
Overview of Metabolic Profile Changes under Stress Owing to Climate Change
Metabolism is strongly linked to plant production and reflects biological circumstances and activities. With advancements in metabolite detection and identification over the past ten years, particularly with the use of mass-spectrometry-based analytical techniques, research on metabolic control under stressful circumstances has been simplified (Medda et al. 2022). Metabolomic studies of Arabidopsis and horticultural plant species have been carried out to gain a greater understanding of cellular states under stress. It appears that plants reorganize their metabolic network to adapt to stressful environments. Factors controlling metabolic movements were examined in conjunction with other “omic” investigations, such as transcriptome experiments, using metabolic changes as a “map” or “marker” (Salam et al. 2023).
Tomato (Solanum lycopersicum) under drought and heat stress upregulate transcription factors such as SlDREB and SlAREB, which regulate proline biosynthesis genes (e.g., P5CS) and the ABA signaling pathway, resulting in the increased accumulation of osmoprotectants and antioxidants (Orellana et al. 2010; Xue et al. 2023). In grapevines (Vitis vinifera), elevated temperatures downregulate VviMYBA1 and VviUFGT, key genes for anthocyanin biosynthesis, reducing the pigment content, while mild water stress can stimulate the ABA-mediated upregulation of VviMYBA1, enhancing anthocyanin levels (Leng et al. 2025). For Capsicum annuum, salinity and heat stress upregulate PAL and CHS, promoting the biosynthesis of capsaicinoids and flavonoids (Zhang et al. 2024). Similarly, in strawberry (Fragaria × ananassa), UV-B and drought stress induce FaAKR23 genes that increase ascorbic acid and anthocyanin (Wei et al. 2022). Lettuce (Lactuca sativa) exposed to cold and high light stress shows altered expression of glucosinolate biosynthesis genes MYB28 and MYB29, boosting stress resistance and enhancing nutritional quality (Li et al. 2021). These genomic shifts reflect the dynamic interplay between environmental stress and plant metabolism, shaping both crop resilience and quality.
It has also been validated that CBF/DREB1 transcription factors are crucially linked to Arabidopsis’s ability to acclimatize to cold and tolerate it. In Arabidopsis, sulfur deficiency triggers the formation of glucosinolate (Meraj et al. 2020). A thorough examination of time-series data pertaining to the levels of metabolites and transcripts using self-organizing mapping revealed that the transcription factors MYB28 and MYB29, together with several downstream enzymes, are responsible for the synthesis of glucosinolate during the sulfur-starvation stress response (Mitreiter and Gigolashvili 2021). The fact that glucosinolate and transcripts for glucosinolate biosynthetic enzymes were present in lower and larger amounts in a gene knockout mutant and in transgenic plants overexpressing MYB28, respectively, supported the role of MYB28 in glucosinolate production (Chen et al. 2024). Henceforth, it is concluded that metabolism is crucial for plant production and adapts to stressful environments. Advancements in metabolite detection and identification have made research easier. Studies of Arabidopsis reveal that transcriptional factors such as MYB28 and MYB29 function as glucosinolate synthesis agents during sulfur-starvation stress.
Similarly, changes in weather often influence primary metabolites such as sugars, amino acids, and organic acids, and secondary metabolites such as phenolics, flavonoids and alkaloids in horticultural plants. For instance, tomato (Solanum lycopersicum) plants exposed to drought stress exhibit elevated levels of proline and soluble sugars, which function as osmoprotectants to maintain cellular turgor and enzyme stability (Patanè et al. 2022). In grapevines (Vitis vinifera), heat stress leads to the increased synthesis of anthocyanins and flavonols, enhancing their antioxidant capacity to counteract oxidative damage (Rienth et al. 2021). In lettuce (Lactuca sativa), salinity stress induces a marked increase in phenolic compounds (Sardar et al. 2023). These metabolic adjustments not only reflect the physiological responses of plant to stress but also influence the quality, flavor, and health-promoting properties of horticultural produce.
Comprehensive Influence of Ecological Factors on Plant Secondary Metabolite Biosynthesis
Many biotic and abiotic stressors influence the production and tissue-specific buildup of plant secondary metabolites, which are tightly controlled in a spatiotemporal manner (Table 3). Environments stress influences how plant secondary metabolites produce and accumulate in plants (Li et al. 2020). In the Brassicaceae family, glucosinolates play a significant role as precursors to a variety of active compounds. Strong light, high temperatures, and droughts cause more glucosinolate accumulation in Brassica rapa (Zafar et al. 2020; Rao et al. 2021). Brassica oleracea exhibits strong resistance to freezing and chilling, which is believed to be supported by the low glucosinolate concentration resulting from low temperatures (Essoh et al. 2020). Apart from low temperatures, additional postharvest stressors including wounding also cause B. oleracea to produce glucosinolates through biosynthesis. The specific glucosinolate and phenolic profiles in broccoli sprouts are impacted by both UV-A and UV-B dosages, indicating a significant function related to light stress in glucosinolate production (Chen et al. 2022). Additionally, biotic stressors may affect the composition of glucosinolate in plants. A buildup of indolyl glucosinolates is caused by the expression of CYP79B2, CYP79B3, and PAD33, which are stimulated by aphids. Strong resistance to herbivory is displayed by the gain-of-function Arabidopsis mutant cml42, which has a greater aliphatic glucosinolate content than the wild type (Nogia and Pati 2021). Therefore, glucosinolates in Brassicaceae plants are influenced by light, temperature, and drought, with Brassica rapa and B. oleracea resistant to freezing and chilling (Ljubej et al. 2021). Light, biotic stressors and aphids also impact glucosinolate composition (Zhan et al. 2022).
Table 3.
Effects of climate change on plant secondary metabolites
| Climatic factor | Secondary metabolites | Sub-class of secondary metabolites | References |
| Cold stress | Phenolics | Flavonoids | Kumar et al. 2023a |
| N- containing | Alkaloids Glucosinolates Glycosides | Teklić et al. 2021 | |
| Terpenoids | -- | He et al. 2023 | |
| Heat stress | Phenolics | Flavonoids | Laddomada et al. 2021 |
| N- containing | Alkaloids Glucosinolates Glycosides | Rehman et al. 2024 | |
| Terpenoids | -- | Liu et al. 2021b | |
| Elevated carbon dioxide | Phenolics | Flavonoids | Kumar et al. 2020b |
| N- containing | Alkaloids Glucosinolates Glycosides | Mahajan et al. 2020 | |
| Terpenoids | -- | Jamloki et al. 2021 | |
| Drought | Phenolics | Flavonoids | Laddomada et al. 2021 |
| N- containing | Alkaloids Glucosinolates Glycosides | Yadav et al. 2021 | |
| Terpenoids | -- | Liang et al. 2021 | |
| Low light | Phenolics | Flavonoids | Hu et al. 2020 |
| N- containing | Alkaloids Glucosinolates Glycosides | Thoma et al. 2020 | |
| Terpenoids | -- | Zhang et al. 2021 | |
| High light | Phenolics | Flavonoids | Ferreyra et al. 2021 |
| N- containing | Alkaloids Glucosinolates Glycosides | Landi et al. 2020 | |
| Terpenoids | -- | Sankhuan et al. 2022 | |
| Ultraviolet B-radiation | Phenolics | Flavonoids | Liu et al. 2020a |
| N- containing | Alkaloids Glucosinolates Glycosides | Jaiswal et al. 2020 | |
| Terpenoids | -- | Shamala et al. 2020 |
Phytohormones and terpenoids are frequently involved in plant defense, with examples including sesquiterpene abscisic acid, triterpene brassinosteroids, and diterpene gibberellins (Divekar et al. 2022; Anas et al. 2025b). Long-lasting contact with heat speeds up the buildup of gibberellin in lettuce, which causes bolting. Significant sesquiterpene abscisic acid accumulation takes place in wheat guard cells during water-deficient conditions (He et al. 2023). Through the regulation of many genes along the carotenoid biosynthesis pathway, NaCl stress significantly increases the level of carotenoid in maize sprouts. Carotenoids are a type of tetraterpenoid with strong antioxidant ability. The endogenous brassinosteroids content of winter wheat is elevated by cold treatments, suggesting that triterpene brassinosteroids play a part in enhancing the low-temperature tolerance of wintertime crops (Zhou et al. 2021b; Siddique et al. 2025). Additionally, terpenoid contents in plants can be impacted by biotic stressors. Three significant aromatic terpenes with confirmed cytotoxic properties - linalool, germacrene D, and γ-cadinene - accumulate more readily in a variety of Iranian basil cultivars when subjected to water deficiency stress (Ghassemi et al. 2021). The capacity of oats (Avena spp.) to synthesize certain triterpenoids, such as avenacins, which have antibacterial properties, is most likely what has enabled them to fight several illnesses (Yeshi et al. 2022). Rich in Euphorbia peplus latex, terpenoids serve as protective agents against a range of agricultural phytopathogenic fungi and insect herbivores. Numerous studies have demonstrated the role terpenoids play as phytohormones in the resistance to environmental stress. Plant cells obtain environmental cues from phytohormones, which act as signal molecules (Park 2021). Accordingly, terpenoids play a vital role in plant defense and in their confrontation mechanisms with environmental stress and can be influenced by numerous stressors, such as high temperatures, water deficiency, and biotic stressors, while also being able to act as signal molecules in plant cells.
Abiotic stress conditions have a substantial impact on the formation of phenolic chemicals. Long-term drought treatments significantly reduce several phenolic acids, including protocatechuic, caffeic, and sinapic acids, in several inbred lines of maize (Ranilla et al. 2021). Salt stress causes significant drops in phenolic chemicals, including ferulic acid, salicylic acid, and sinapic acid, in Chinese cabbage. Rice stimulates the root epidermis and stele to secrete more phenolic acid in an alkaline environment (Riaz et al. 2025). This successfully boosts ion absorption and reduces Fe-deficiency responses (Ahlawat et al. 2024). Drought stress significantly raises the levels of phenolics such as chlorogenic and caffeic in Achillea pachycephala. Plants with systemic resistance mediated by rhizobacteria are better able to fend off insects and diseases. Plants under pathogenic stress are bio-protected by phenolic-acid-induced systemic resistance. It has also been discovered that the protective actions of phenolics and rice seedling against Rhizoctonia solani are correlated (Tuladhar et al. 2021). High-performance liquid chromatography research revealed that seedlings infected with Rhizobium show significantly improved phenolic acid production compared to uninfected controls (Zhan et al. 2022). Dendrobium viroid contamination raises the total phenolic concentration in D. officinale orchids, which may be crucial for initiating pathogen defense responses. The biological roles of phenolic acid are multifaceted (Kumar et al. 2020a). The production of phenolic acid is inhibited by some strains, while phenolic acid concentrations are promoted by other pressures. Phenolic acids may have both helpful and harmful influences on the body's capability to endure environmental stress (Marchiosi et al. 2020). Therefore, phosphoric acid can play a vital function in plant defense against environmental stress (Khan et al. 2023). Long-term drought treatments reduce phenolic acids in maize, Chinese cabbage, and Achillea pachycephala (Gharibi et al. 2019). Phosphoric-acid-induced systemic resistance protects plants against insects and diseases while inhibiting or promoting phenolic acid production (Rashid and Chung 2017).
Polyphenols such as flavonoids are antioxidant agents that are necessary for plants to withstand stress. Under environmental stress, plants with higher flavonoid concentrations may have subcellular antioxidant properties (Shi et al. 2025). In model plants, the overabundance of several flavonoids, including cyanidin, quercetin, and kaempferol, has been extensively studied (Ahmed et al. 2021). Certain ortho-dehydroxylated B-ring flavonoids, which are potent ROS scavengers, are found in higher concentrations in Chinese liquorice when exposed to UV-B light (Pandey et al. 2023). Stress tolerance in rice is mostly dependent on flavonoid accumulation, which is enhanced by heat and salt stress. Abiotic stressors have a detrimental effect on the accumulation of flavonoids in several plant species. For instance, under salt stress, Na2+ deposited in Apocynum venetum leaves lowers the quantity of flavonoids and reduces salinity (Chen et al. 2020). In citrus fruit, hesperidin and hesperidin, considered as the main flavonoids, have been extensively studied for their potential to ward off pathogens and poisons (Yan et al. 2024). Short-wavelength radiation increases the levels of many flavonol glycosides, including quercetin and kaempferol glycosides, thus strengthening plant defenses against a variety of herbivorous insects (Rechner et al. 2017). Plants sometimes create another category of flavonoids called anthocyanins under living and nonliving stressors (Shah and Smith 2020). In plants, anthocyanin production and tissue-specific accumulation are influenced by a variety of environmental variables (Do et al. 2023). Water deficiency, lower temperatures, UV-blue rays, and light contact greatly increase the amount of anthocyanin in apples, whereas high temperatures and increased levels of nitrogen fertilizer dramatically decrease the amount (Gao et al. 2021), suggesting that flavonoids, antioxidants essential for plants to withstand stress, are found in various plant species. High flavonoid concentrations can enhance stress tolerance, while biotic stressors can negatively affect their accumulation. Flavonoids in citrus fruits, anthocyanins, and other plants also respond to environmental factors.
Metabolic Engineering of Plant Secondary Metabolites
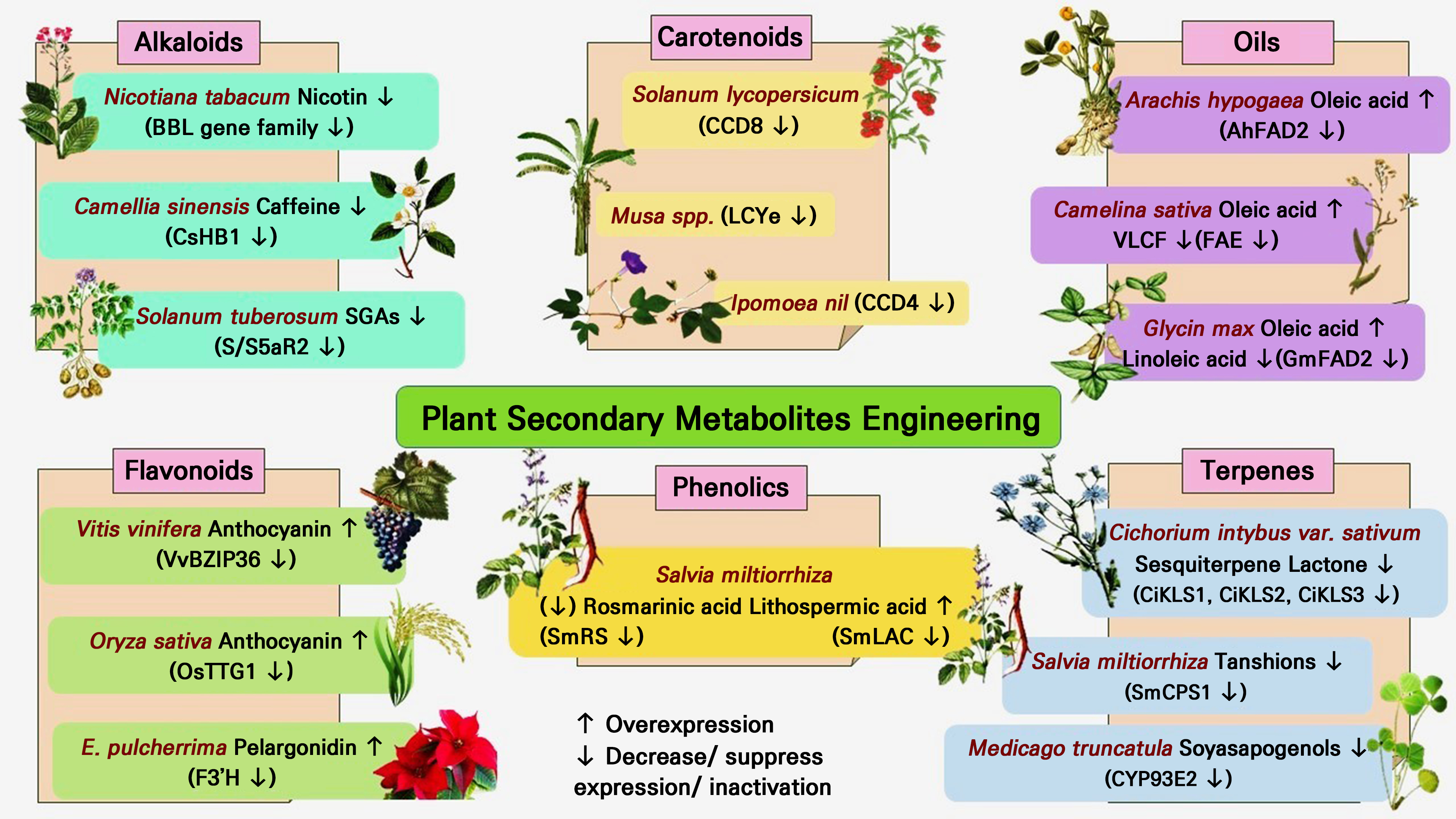
A comprehensive grasp of the entire biological synthetic process and a profound comprehension of the controlling processes governing the start and flow of the aforementioned processes are necessary to realize the lucid engineering of secondary metabolic actions in plants (Barone et al. 2020). For most secondary metabolites, such data are currently unavailable, which explains why metabolic engineering has had only patchy success (Elshamy et al. 2019; Chai et al. 2020). Increased production levels and the targeted production of desired molecules were made possible by recent developments in the metabolic engineering of microbes that produce both built-in and heterologous secondary metabolites (Fig. 1). Additionally, it has proven possible to design cells to produce new substances such as polyketide antibiotics (Vasudevan and Lee 2020). However, unlike those of plants, the biological synthetic routes for organic compounds in microbes are frequently shorter and require fewer enzyme steps to create the required end product. As a consequence of many years of traditional biochemical study, only a few routes (such as flavonoids, terpenoid indoles, and isoquinoline alkaloids) in plants have been fully studied thus far. Genetic engineering can be applied in several ways to improve the manufacturing of desired medications (Anwar et al. 2021).
It is known that there is great potential with regard to metabolically engineering secondary metabolites obtained from plants by altering anthocyanin and flavonoid pathways, which can result in altered floral colors or greater synthesis of antioxidant flavonols in tomatoes. Nevertheless, there has not been much success in altering pathways to create molecules with significant medicinal value thus far (Mipeshwaree Devi et al. 2023). The high-value tropane alkaloid scopolamine, which was beforehand only present in minute levels in Atropa bella-donna, accumulated as a result of the insertion of a gene that produced hyoscyamine-6b-hydroxylase, previously discovered in Hyoscyamus niger. Scopolamine was produced almost entirely from hyoscyamine (Halder and Roy 2023). This was the first instance of a significant medicinal plant being successfully engineered to produce a valuable end product. While a similar genetic factor was up-regulated in Hyoscyamus muticus hairy roots, an even more pronounced impact was noted. In this instance, there was an accumulation of high quantities of hyoscyamine in the hairy roots in addition to the production of considerable amounts of scopolamine (Özmen et al. 2022). Another illustration is the finding that putrescine N-methyltransferase was downregulated by antisense in transgenic tobacco plants, which coincided with a decrease in the nicotine quantity and however, curiously, likewise resulted in increased levels of anatabine (Zhou and Jander 2021). However, there has been limited success in creating molecules with significant medicinal value. The ectopic production of transcription factors and the activation of entire pathways create new opportunities for developing secondary metabolite pathways (Atkins et al. 2016).
Functional Genomics of Plant Secondary Metabolites
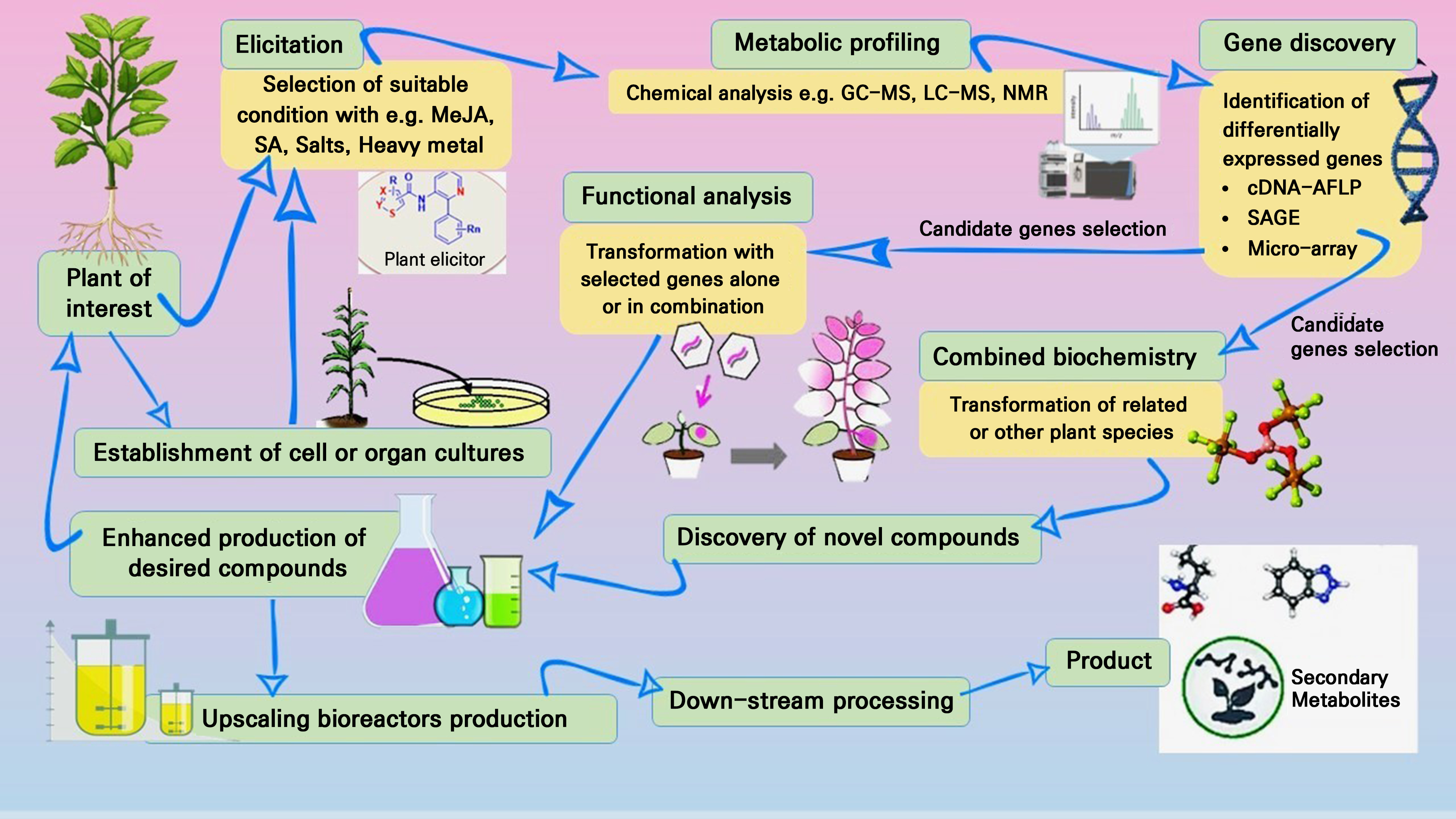
Transcriptomics, proteomics, and metabolomics are three functional genomics techniques that are effective in expediting thorough examinations of biological systems. It is hardly unexpected that almost no such thorough investigations have been published, as most plants that produce important secondary metabolites (such as paclitaxel and terpenoid indole alkaloids) lack genetic tools (Neumann et al. 2020). For a better understanding of how plants regulate their secondary metabolites production and molecular processes, it is imperative to explore functional genes. At the subcellular level, C. chinensis Franch exhibit numerous WRKY transcription factors that are responsible for the production of protoberberine alkaloids. These findings may serve as the basis for more investigations of berberine biosynthesis control methods (Xu et al. 2020a). Genome-wide transcription describing techniques at the cell/organ level may also be used to identify genetic factors which regulate anticipated metabolite build up once the conditions necessary to elicit it (like methyl-jasmonate) have been established (Fig. 2).
Many possible secondary metabolites that synthesize genetic materials have been found through transcript profiling, which is a high-throughput functional study to clarify their regulation actions. Examples of this type of analysis include the sequestration of full-length complementary chromosomes, the high-throughput creation of expression vectors for desired products using tools such as Gateway, the development of a rapid transformation method, the optimization of plant tissue cultures, the efficient profiling of metabolites, and measurable analyses (Fig. 2). Characterizing the plant metabolome is a general challenge due to the significant chemical variety and complexity of molecules. When profiling many metabolites simultaneously, analytical methods are challenged by the diversity of chemical characteristics (Li and Gaquerel 2021). This is especially clear with regard to the study of secondary metabolites, which remain significantly intricate compared to primary metabolite profiles. A veritable revolution in the profiling of primary and secondary metabolites has occurred recently (Zhao et al. 2020).
Functional genomics techniques such as transcriptomics, proteomics, and metabolomics are helpful to study biological systems. However, many plants producing secondary metabolites lack genetic tools, making it crucial to explore functional genes to understand their regulation and molecular processes (Xu et al. 2023). A functional genomics method could expedite secondary metabolite pathway gene identification if it could work with tissue culturing as well as known gene sequences. A genome-wide transcriptional profile can recognize genetic materials whose regulation relates to the accumulation of desired metabolites, such as nicotine manufacturing in tobacco cells (Loukola et al. 2015). A high-throughput functional analysis is required to clarify the potential functions of the genetic materials responsible for secondary metabolite synthesis. Post-transcriptional regulation appears to play a role in stress reactions, with mRNA processing and metabolism closely related to stress responses (Hernández-Elvira and Sunnerhagen 2022).
Transcription Factors and Genes Related to Plant Secondary Metabolites
Transcriptional control by transcription factors, the synthesis of inducible secondary metabolites, and the expression of relevant biological synthetic genetic factors are significantly altered at different stages. Transcription factors are DNA-binding proteins that bind to target gene promoter parts using RNA polymerases to change the pace at which transcription initiates (Kumar et al. 2023b). Transcription factors can combine internal and external cues to control enzyme genetic factor regulation and to monitor the built-up of secondary metabolites. Numerous transcription factors regulate the genes linked to the biological synthesis of secondary metabolites at different levels. In recent decades, there has been a surge in the discovery of transcription factors and an increased emphasis on understanding the regulatory mechanisms involved in the biosynthesis of secondary metabolites (Cao et al. 2020). We provide a summary of transcriptional factors linked to secondary metabolites pathway control methods in different plant species in this section and summarize our understanding.
WRKY Transcription factors
Numerous investigations have been conducted on the WRKY transcription factor class in stressed plants. The control of defense-related secondary metabolite biological synthesis is supported via an inducible regulation pattern of WRKY genetic factors. A 60-amino acids long chain was found in WRKY family members, which regulates genes expression (Ma et al. 2024). The primary role of plant WRKY transcription factors is to respond to stress (Adjibolosoo et al. 2024). They are regulated by wound signaling and jasmonic acid, which, along with genetic modifications, influence the production of various secondary metabolites, including terpenoids, alkaloids, and their derivatives. For instance, WRKY3 and WRKY6 are linked to the production of volatile terpenes in tobacco (Biswas et al. 2022). WRKY1 regulates the biological synthesis of gossypol in cotton plants by binding to the promoter region of the gene involved in cadinene synthesis (Xu et al. 2004). Similarly, in Artemisia annua, WRKY1 influences the biosynthesis of artemisinin by attaching to the promoter of the sesquiterpene synthesis gene (Han et al. 2014). Additionally, WRKY1 was investigated in late blight-infected potato plants, where it bound to the promoter site of the gene encoding the production of hydroxycinnamic acid amide (HCAA) by modifying the phenylpropanoid pathway (Xie et al. 2021). These transcriptional factors may control genes function during the generation of secondary metabolites, including terpenoids and alkaloids. They are controlled via signaling pathways as well as the phenylpropanoid modification pathway. The coordinated transcriptional initiation of the entire biosynthetic pathway may not always be caused by a single transcription factor.
MYB Transcription factors
MYBs, among the several types of transcriptional elements, are engaged in the biological synthesis of secondary metabolites and take part in several genetic functions, including development, reproduction, and stress lenience. The DNA-attaching domains of MYB transcript elements, composed of imperfect repetitions of 50–53 amino acids each, vary in number (Thakur and Vasudev 2022). They fall into one of four subclasses, designated R1 to R4, based on repetitions of the DNA-attaching domain. In several plant species, the R2R3 class of MYB transcription factors plays a major role in the control of multiple secondary metabolite pathways. For example, in Arabidopsis thaliana, AtMYB113, AtMYB114, AtMYB75, and AtMYB90 may be implicated in anthocyanin residue modulation by phenylpropanoid synthesis modification (Xiao et al. 2021). Furthermore, in Arabidopsis, aliphatic glucosinolate accumulated in the shoot is linked to MYB29 and AtMYB76, while the accumulation of indole glucosinolates is influenced by AtMYB34, AtMYB51, and AtMYB122 through changes in the regulation of tryptophan synthesis genetic materials, including CYP79B2, CYP79B3, and CYP83B1 (He et al. 2021). These genes initiate the phenylpropanoid pathway, which includes HCT, 4CL3, C4H, and PAL. In poplar trees, PtMYB115 attaches to ANR1 and LAR3 promoter regions to boost the expression levels of these genes, which increases both proanthocyanin accumulation and resistance to Dothiorella gregaria (Liu 2022). This suggests that MYBs, a type of transcription factor, are crucial for plant development, reproduction, and stress tolerance. They are categorized into four subclasses, with the R2R3 family regulating several secondary metabolite pathways (Yan et al. 2021). In Arabidopsis thaliana, the genes AtMYB113, AtMYB114, AtMYB75, and AtMYB90 are involved in the accumulation of anthocyanins, glucosinolates (GLs), flavonoids, hydroxycinnamic acids (HCAAs), and proanthocyanins (Mao et al. 2025). The accumulation of flavonoids is influenced by the heterologous expression of AtMYB transcription factors across different plant species (Pandey et al. 2014).
bHLH Transcription factors
Potential regulators of stress-response processes, the bHLH transcription factors often form complexes with MYB proteins that promote the expression of particular genes. These 60-amino-acid proteins are called transcription factors, and they include bipartite conserved domains (Banerjee et al. 2022). Two alpha helices let helix-loop-helix attach to proteins to create a dimeric composite, while the crucial remains at the N-terminal of the protein enables bHLH to bind to genetic material. The bHLH transcriptional factors control the manufacture of secondary metabolites such as anthocyanins, alkaloids, glucosinolates, diterpenoid phytoalexins, and saponins and are significant modulators of stress responses (Yamada and Sato 2021). In particular, MYB51 interacts with bHLH04, bHLH05, and bHLH06 to control GLs production meaningfully in Arabidopsis. bHLHs are extensively implicated in the phenylpropanoid pathway expression during the manufacturing of anthocyanins and flavonoids. For example, in the presence of TTG1, the bHLH proteins (GL3, eGL3, and TT8) join with MYB to form a transcriptional regulatory complex that controls the genes involved in anthocyanin production (Escaray et al. 2024). This suggests that BHLH transcription factors are 60-amino-acid proteins that regulate stress-response processes in plants. They form complexes with MYB proteins to promote gene expression, controlling the manufacture of stress-inhibiting substances such as anthocyanins, alkaloids, glucosinolates, diterpenoid phytoalexins, and saponins. Transcription factors are implicated in the phenylpropanoid pathway, jasmonic acid signaling, and the biosynthesis of nicotine. They can regulate stress responses in plants on their own or in conjunction with other protein families.
bZIP Transcription factors
The bZIP transcription factors are dimeric transcriptional enhancer proteins with a preserved leucine zipper and a positively charged genetic material requisite region. They usually modulate plant genetic methods. bZIP transcriptional factor family members are related to stress responses, including ROS lenience as well as osmotic disproportion. Numerous Yap-like bZIPs (YAP is yeast bZIP protein’s subsection), including NapA, AtfA, Aoyap1, Afyap1, and Apyap1, were found to function in Aspergillus species under osmotic, oxidative, nutritional, as well as pharmacological stressors (Hrmova and Hussain 2021). Previous examinations indicated that bZIP and oxidative stress govern secondary metabolite biological synthesis in fungi. In a modified line, the bZIP-like protein RsmA hinders metabolite synthesis; whereas in an overexpressed line, RsamA actively contributes to the restoration of secondary metabolites. It is also known that some bZIP proteins, such as SmbZIP20, SmbZIP7, and AabZIP1, control pharmaceutically significant secondary metabolites in Salvia miltiorrhiza and Artemisia annua, respectively: tanshinone and artemisinin (Han et al. 2023), suggesting that bZIP transcription factors, including YAP-like proteins, regulate plant biological processes and stress responses. They function in secondary metabolite biosynthesis, anthocyanin generation, and the production of terpenoid phytoalexins in rice. They also regulate aflatoxin regulation in Aspergillus species. Various bZIP proteins, including RsmA, SmbZIP20, SmbZIP7, and AabZIP, play crucial roles in plant health.
Effects of Genetic Factors on the Biosynthesis of Metabolites
Plant secondary metabolite accumulation in stressful environments is regulated by a complex network made up of several transcription factors. Numerous important genes that encode enzymes for plant secondary metabolite production are downstream targets of many transcription factors (Zhan et al. 2022).
Transcription factors involved in the biosynthesis of alkaloids
Numerous transcription factors that regulate certain stages and branches of the TIA and BIA biosynthesis pathways were found in earlier research. Genes associated with the TIA biosynthesis pathway, including geissoschizine synthase, strictosidine synthase, and deacetylvindoline acetyltransferase, are expressed in C. roseus under the control of an ORCA3 transcription factor. Through its modulation of the TIA pathway genes in C. roseus, the interaction between MYC2 and GBF controls TIA production (Ahmed et al. 2023). Positive regulation of the TIA biosynthesis pathway is mediated by WRKY1. The TIA biosynthesis route is regulated by a MAP kinase cascade that activates the downstream target AP2/ERF transcription factor genes. Furthermore, in the TIA biosynthesis pathway, the zinc-finger transcription factor ZCT1 functions as a transcriptional repressor (Ahmed et al. 2024). OpWRKY2 and OpWRKY3 are two positive regulators in the biosynthesis of camptothecin in Ophiorrhiza pumila. In lotus (Nelumbo nucifera), CYP80G, TYDC, NCS, and 7OMT genes are regulated by WRKY40a and WRKY40b, which are involved in the BIA biosynthesis pathway. The transcription factor RAP2-7 is involved in controlling the biosynthesis route of quinolizidine alkaloids in narrow-leaved lupin (Li et al. 2022). Nicotine, the main alkaloid in tobacco leaves, is produced by two jasmonate-responsive transcription factors, ERF189 and ERF199. By controlling the expression of the PMT1 gene, which encodes a putrescine N-methyl transferase implicated in the crucial stage of the pyridine alkaloid pathway, MYC2 increases the nicotine concentration under high-temperature-stress circumstances. CjbHLH1 homologs regulate isoquinoline alkaloid production in Coptis japonica (Bhushan et al. 2023). Previous studies have indicated that various transcription factors regulate the biosynthesis pathways of TIA and BIA in plants (Godbole et al. 2022; Li et al. 2025). The ORCA3 transcription factor specifically controls TIA genes in C. roseus, while WRKY1 and ZCT1 are responsible for regulating BIA pathways. Additionally, the transcription factors RAP2-7, ERF189, and ERF199 govern the production of quinolizidine alkaloids in lupin and tobacco leaves (Suttipanta et al. 2011; Paul et al. 2017).
Transcription factors involved in the biosynthesis of phenolic acids
The Chinese medicinal plant Salvia miltiorrhiza has a phenolic acid pathway that is regulated by several transcription factors. The phenolic acid biosynthetic pathway is regulated by a multitude of transcription factors, comprising two members of the ERF family (SmERF115 and SmERF1L1), three members of the MYB family (SmMYB2a, SmMYB2b, and SmMYB52), four members of the bHLH family (SmbHLH3, SmbHLH37, SmbHLH51, and SmbHLH148), one ZIP family member (SmZIP1), two GRAS family members (SmGRAS1 and SmGRAS2), and four members of the bHLH family (SmbHLH3, SmbHLH3, SmbHLH37, SmbHLH51, and SmbHLH148) (Zhou et al. 2021a). Moreover, S. miltiorrhiza has been shown to have matching downstream targets of the aforementioned transcription factors. SmHCT6 and SmCYP98A14, two important genes involved in the synthesis of 4-coumaroyl-3',4'-dihydroxyphenyllactic acid, and rosmarinic acid, respectively, have promoter regions that include E-boxes that are bound by SmMYC2a/b. Tanshinones are biosynthesized from GGPP through the binding of SmGRAS1 and SmGRAS2 to the GARE box in the promoter region of SmKSL1 (Deng et al. 2020). The G-Box-like1 motif in the promoter region of SmC4H1, which is involved in the biosynthesis of phenolic acid precursors, is bound by the ABA-responsive transcription factor SmbZIP1. SmbHLH148 and SmMYB1 control the production of phenolic acids by promoting the expression of downstream genes including CYP98A14, TAT, HPPR, RAS, and PAL1 (Ding et al. 2023). By attaching to the MBE elements in the promoter regions of SmTAT1, Sm4CL9, SmC4H1, and SmHPPR1, SmMYB52 concurrently influences the synthesis of phenolic acids. Furthermore, by downregulating the expression of DXS3, DXR, HMGR1, KSL1, CPS1, and CYP76AH1, SmbHLH3 functions as a repressor during the production of phenolic acids in S. miltiorrhiza hairy roots (Zhang et al. 2020a). Tanshinone and salvianolic acid production in S. miltiorrhiza hairy roots are regulated by SmMYB09. It is noteworthy that in S. miltiorrhiza hairy roots, the over-expression of Arabidopsis MYC2 simultaneously stimulates the production of phenolic acid and tanshinone. Furthermore, SmJRB1 was found to be a positive regulator in the control of the biosynthesis of phenolic acid (Liu et al. 2022), suggesting that S. miltiorrhiza has a phenolic acid pathway regulated by various transcription factors (TFs). These include the ERF, MYB, bHLH, ZIP, GRAS, and bHLH families. SmMYB1 controls phenolic acid production by promoting downstream genes, while SmMYB09 regulates tanshinone and salvianolic acid production. The over-expression of Arabidopsis MYC2 stimulates phenolic acid and tanshinone production.
Transcription factors involved in the biosynthesis of flavonoids
Three main branches make up the entire flavonoid biosynthesis pathway, which has been extensively researched in plants. Transcription factors from several families, including bHLH, WRKY, and MYB, have also been characterized recently. Activating flavonol biosynthesis in M. truncatula, MtMYB134 binds to the promoters of MtFLS1, MtFLS2, and MtCHS2 (Naik et al. 2022). Through the activation of the promoters of the MdMYB9 and MdMYB11 genes, MdNAC52 controls the production of anthocyanin and proanthocyanidins in apples. By upregulating the DFR and LDOX genes, MYBF1 controls the flavonol biosynthesis pathway in buckwheat. Light-induced FtMYB116 in Fagopyrum tataricum directly binds to the F3'H promoter region to facilitate the accumulation of rutin. By activating the promoters of PpCHS, PpCHI, PpF3H, PpFLS, and PpUFGT, PpMYB17 positively regulates the flavonoid biosynthesis pathway in pears (Pyrus pyrifolia) (Premathilake et al. 2020; Liu et al. 2021c). PbWRKY75, a different pear (Pyrus × bretschneideri) transcription factor, controls the expression levels of PbDFR, PbUFGT, and PbMYB10b, thus influencing flavonoid biosynthesis. PtMYB6 physically interacts with KNAT7 in Populus tomentosa to increase the production of anthocyanins and proanthocyanidins. StWRKY13 activates the promoters of StCHS, StF3H, StDFR, and StANS in potato (Solanum tuberosum) tubers to enhance anthocyanin biosynthesis. When MdWRKY11 is overexpressed in apple calli, it becomes clear that its unique role involves binding to the MdHY5 promoter and accelerating the accumulation of flavonoids and anthocyanins (Zhao et al. 2023). Furthermore, a variety of plant species have been found to have several negative regulators of the flavonoid biosynthesis pathway (Liu et al. 2024). GbMYBF2, a negative regulator found in Ginkgo biloba, inhibits the production of flavonoids by downregulating several important genes, including GbPAL, GbANS, GbFLS, and GbCHS2. BnWRKY41-1, a Brassica napus WRKY transcription factor, suppresses the production of anthocyanins. SmbHLH60 has been shown to be a negative regulator of anthocyanin biosynthesis in S. miltiorrhiza, mostly through the SmDRF gene (Wu et al. 2021), This suggests that the flavonoid biosynthesis pathway in plants is regulated by three main branches: MtMYB134, MdNAC52, MYBF1, and PpMYB17. TFs from various families, such as bHLH, WRKY, and MYB, play a crucial role in regulating flavonol biosynthesis. Additionally, negative regulators such as AtMYB4 and MdBHLH33 prevent flavonoid build-up, suppress anthocyanin accumulation, and inhibit the production of essential genes.
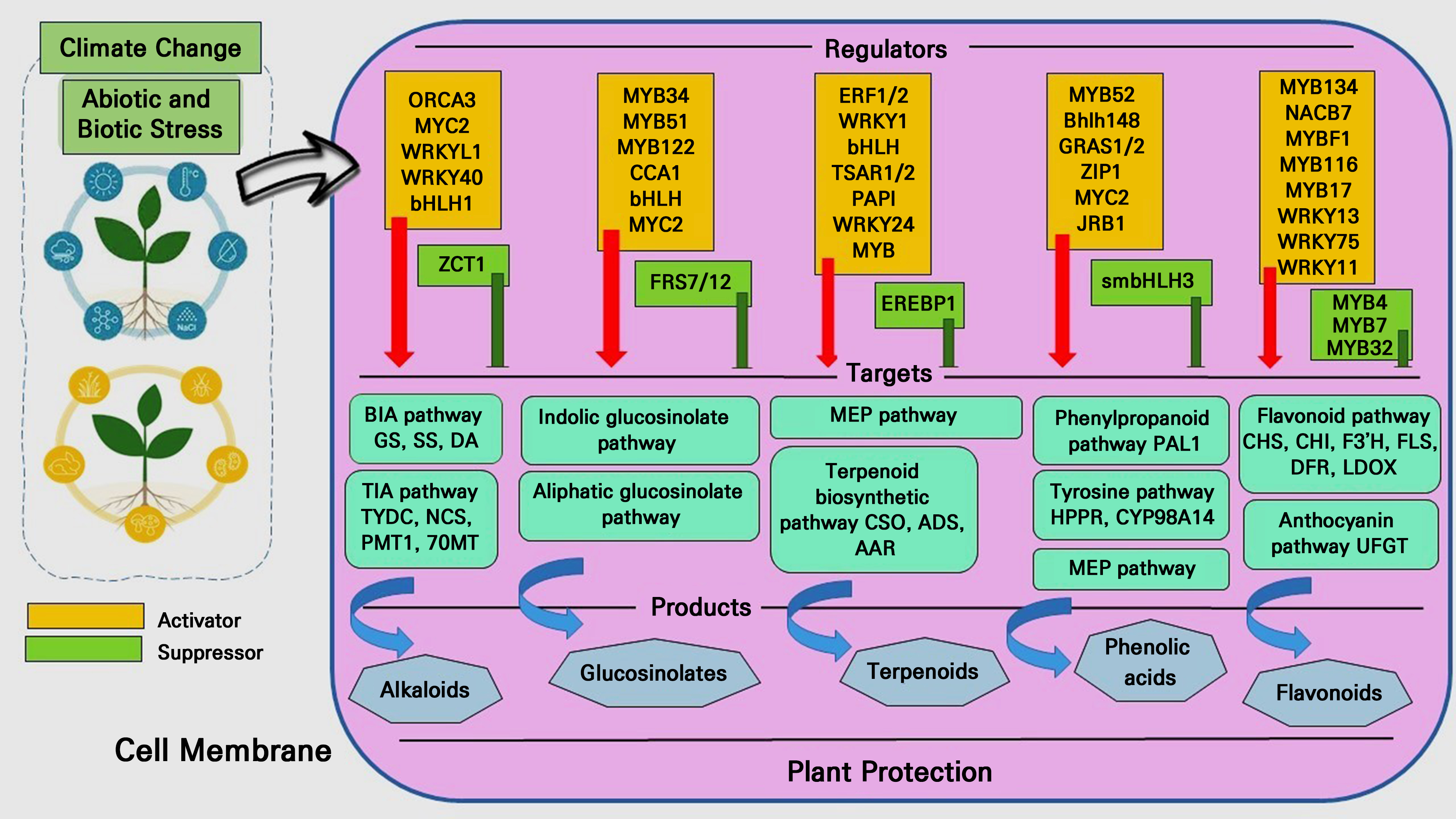
Numerous transcription factors allow environmental cues to cross the cell membrane and activate downstream functional genes. Numerous transcription factors participating in the metabolic process have been found in various plants thus far (Pacheco et al. 2021). A network involved in the transcriptional control of plant secondary metabolites biosynthesis under environmental stressors is summarized in our review (Fig. 3). While there are still very few known negative transcription factors, a significant number of positive transcription factors have been found to date. Additionally, the creation of a dynamic equilibrium in plant secondary metabolism is largely dependent on negative regulatory transcription factors. Cloning and the finding of negative regulatory transcription factors have emerged as critical research areas for the future (Sun and Fernie 2023). Henceforth, transcription factors in plants activate genes under environmental stress, controlling PSM biosynthesis. Despite the few known negative transcription factors, positive transcription factors are abundant, as noted above. Cloning and negative regulatory transcription factors are critical research areas.
Conclusions
Plant secondary metabolites are essential for environmental adaptability. They also attract pollinators, regulate development, and guard against diseases and herbivores. Gene expression and plant secondary metabolites synthesis are two ways in which plants respond to environmental stress. For example, poplars and sweet potatoes modify their lignin content. Encouraging sustainable agriculture and protecting natural ecosystems requires an understanding of the molecular processes involved in the production of plant secondary metabolites. Particularly in areas with significant biodiversity, plants are threatened by climate change. Plant production depends on metabolism, which also helps plants adapt to harsh conditions. Research has become easier due to developments in metabolite detection and identification. Research on Arabidopsis shows that certain transcription factors, such as MYB28 and MYB29, aid glucosinolate production under sulfur-starvation stress. In C. roseus, more than 100 TIAs, including bisindole alkaloids, have been found. Seedlings subjected to salt and dehydration stress have shown higher alkaloid contents. One type of plant secondary metabolite that is essential for managing environmental stress is alkaloids.
Plant metabolism engineering necessitates a thorough comprehension of the metabolic pathway and regulatory mechanisms. Recent advancements in the metabolic engineering of microbes have enabled the targeted synthesis of desired compounds and increased production levels. Genetic engineering can modify anthocyanin and flavonoid pathways to enhance the production of targeted pharmaceuticals. There are more opportunities to create secondary metabolite pathways when transcription factors are produced ectopically and whole pathways are activated. Functional genomics methods may be helpful for understanding biological systems. Nevertheless, because many plants that produce secondary metabolites are genetically limited, it is essential to investigate functional genes to comprehend the regulatory and molecular mechanisms involved. Given that functional genomics may be used in any plant or cell culture that lacks known gene sequences, it may speed up the process of identifying the genes encoding secondary metabolite pathways.
Stress reactions appear to be influenced by post-transcriptional regulation, as stress responses are strongly associated with mRNA metabolism and processing. Stressed plants have demonstrated that the WRKY transcription factor family regulates defense-related secondary metabolite biosynthesis using inducible expression patterns. The production of chemicals that inhibit various secondary metabolites is governed by BHLH transcription factors, responsible for expressing stress-response mechanisms. Through the regulation of plant stress mediators, AP2/ERF transcription factors are essential in the regulation of plant stress responses. Protecting natural ecosystems and promoting sustainable agriculture depend on knowledge of the molecular processes behind plant secondary metabolite generation.