Introduction
Materials and Methods
Plant Material
DNA Extraction
Sequencing the nor Gene and Polymorphism Detection
High-resolution Melting Analysis in the Germplasm Collection
Results and Discussion
Discovery of Sequence Variation in the nor Gene in Tomato
Gene-based Marker Development and Genotyping in a Collection of Tomato Varieties
Introduction
Tomato (Solanum lycopersicum L.) is one of the most widely consumed fresh fruit vegetables in the world. Tomato fruits make a significant contribution to human nutrition due to their abundant vitamins, minerals, and antioxidants (Raiola et al., 2014; Perveen et al., 2015). As a climacteric and perishable vegetable, tomato fruit has a relatively short life after ripening due to biological processes affecting fruit quality during the postharvest stage (Zapata et al., 2008). The quality and nutritional value of ripening fruit are greatly reduced by postharvest handling and storage practices (Sablani et al., 2006), with a loss of fresh tomato fruit that can be as much as 45.32% (Kasso and Bekele, 2016). Therefore, a number of strategies have been developed to improve fruit quality and shelf-life such as refrigeration, heat treatment, modified atmosphere packaging (MAP), and 1-methylcyclopropent (1-MCP) and calcium chloride (CaCl2) application (Arah et al., 2016). Although effective in limiting postharvest losses, these methods can be costly and time-consuming, especially for small farms (Hoering, 2012). The use of ripening mutant genes, however, eliminates these disadvantages and is an effective strategy in tomato breeding programs.
Several tomato ripening genes have been reported including ripening-inhibitor (rin) (Robinson and Tomes, 1968; Vrebalov et al., 2002), non-ripening (nor) (Tigchelaar et al., 1973), Colorless non-ripening (Cnr) (Thompson et al., 1999; Eriksson et al., 2004; Manning et al., 2006), Green-ripe (Gr) (Kerr, 1958; Barry and Giovannoni, 2006), Never-ripe (Nr) (Lanahan et al., 1994), and alcobaca (alc) (Kopeliovitch et al., 1981). These genes participate independently and cooperatively in the fruit ripening process. For example, the rin mutation produces a partially deleted MADS-box protein that is a ripening-specific transcription factor and results in effectively blocking the fruit ripening process leading to green fruits (Vrebalov et al., 2002). The heterozygous mutation of the rin gene (rin/Rin) is often used to produce slow ripening and long shelf-life tomatoes (Giovannoni, 2007). Similarly, the mutation in the nor gene encoding a NAC transcription factor fails to initiate the normal ripening process and results in fruits that do not soften (Lincoln and Fischer, 1988; Giovannoni, 2004). The nor gene can be also used in the heterozygous form (nor/Nor) to improve shelf-life of tomato fruit. The NAC transcription factor family is important for diverse physiological process during development including stress response, flowering, and senescence (Martel et al., 2011). The nor mutation enhances disease resistance by up-regulating the expression of several genes associated with defense mechanisms that biotic stress, including the resistance of tomato fruit to Botrytis cinerea (Cantu et al., 2009).
Conventional plant breeding based on phenotypic selection is generally labor-intensive and time-consuming. Conversely, the use of molecular markers associated with traits of interest can be a cost-effective and accurate option to develop new elite varieties (Phan and Sim, 2017). Recent advances in next-generation sequencing technology have facilitated association mapping with genome-wide molecular markers such as single nucleotide polymorphism (SNP) and insertion and deletion (InDel). These efforts have led to the discovery of a number of markers associated with major genes and quantitative trait loci (QTL) (Truong et al., 2015; Devran et al., 2016; Veerappan et al., 2016). However, these markers are not always effective for breeding populations due to linkage disequilibrium (LD) decay. Gene-based markers can be more practical relative to the gene-associated markers for marker-assisted selection (MAS) in breeding programs. Although the molecular and biochemical basis of fruit ripening has been intensively studied in tomato, molecular markers for breeding have been less utilized in comparison to other traits such as disease resistance. Recently, a sequence characterized amplified region (SCAR) marker was developed for the rin mutation (Kim et al., 2013); however, few studies have developed molecular markers for the nor mutation, which is more effective for long shelf-life relative to the rin gene (Ng, 1976). Thus, the objectives of our study were to 1) detect sequence variations between the nor mutant and wild type varieties, 2) develop a gene-based marker for MAS to improve tomato fruit quality and long shelf-life, and 3) investigate the nor mutation in a collection of 81 inbred and commercial F1 hybrid varieties based on our gene-based marker.
Materials and Methods
Plant Materials
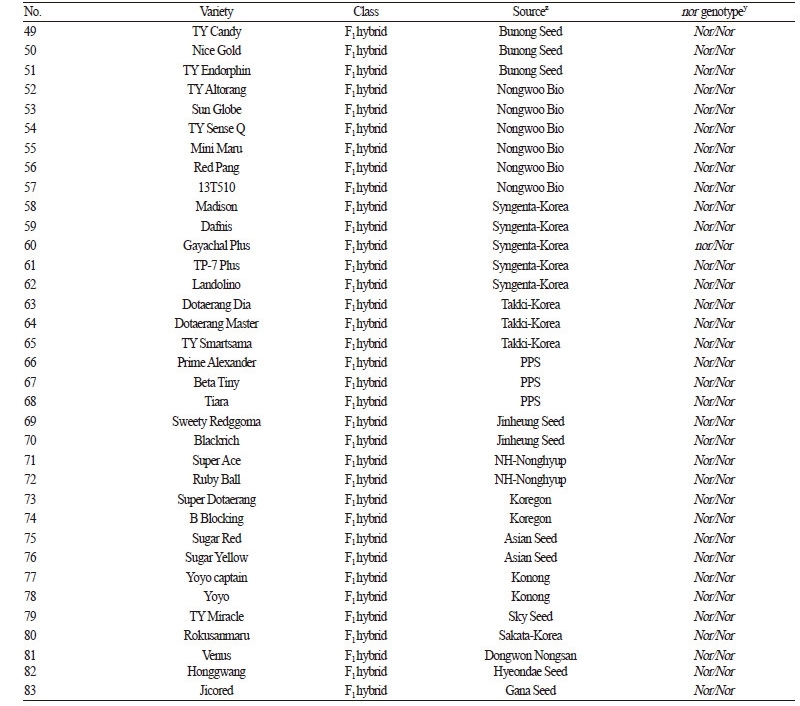
Three tomato accessions, ‘San Marzano’ (wild-type allele, Nor/Nor), ‘LA1793’, and ‘LA3013’ (both mutant alleles, nor/nor) were provided by the C. M. Rick Tomato Genetics Resource Center at University of California, Davis, CA and used to investigate sequence variation in the nor gene. In addition, we used 81 other varieties to validate the resulting nor sequence variation(s). This germplasm panel consisted of 30 inbred and 51 commercial F1 hybrid varieties from public and private breeding programs in South Korea. The inbred varieties were collected from the National Agrobiodiveristy Center (20 varieties) and National Institute of Horticultural & Herbal Science (10 varieties) at Rural Development Administration (RDA). For the F1 hybrids, 51 varieties were derived from Bunong Seed (10 varieties), Monsanto-Korea (nine varieties), Nongwoo Bio (six varieties), Syngenta-Korea (five varieties), Takki-Korea, PPS (three varieties from each company), Jinheung Seed, NH-Nonghyup, Koregon, Asian Seed, and Konong (two varieties from each company); and the rest five varieties were from Sky Seed, Sakata-Korea, Dongwon Nongsan, Hyeondae Seed, and Gana Seed.
DNA Extraction
Genomic DNA was isolated from young leaves from seedlings using the CTAB method with minor modifications (Kabelka et al., 2002). Young leaf tissue was placed in a 2.0 mL micro-centrifuge tube containing 350 µL DNA extraction-lysis buffer and ground in a TissueLyser II (QIAGEN, Valencia, CA, USA) at 300 strokes per minute for 5 min. After incubation at 65°C for 20 min, 350 µL of chloroform: isoamyl (24:1) was added and then centrifuged at 5,000 g for 10 min. The supernatant was separated and mixed with 200 µL ice cold isopropanol, keep stable at 4°C for 1 h, followed by centrifugation at 5,000 g for 15 min. The DNA pellet was rinsed twice with 70% ethanol, dried at room temperature, and re-suspended in 100 mL TE buffer (10 mM Tris-HCl pH 8.0, 0.1 mM EDTA). The quantity and quality of DNA sample were measured using agarose gel electrophoresis and diluted to 50 ng·µL-1 for PCR amplification and high-resolution melting (HRM) analysis.
Sequencing the nor Gene and Polymorphism Detection
The full length of the nor gene (SGN Gene ID: Solyc10g006880) was retrieved from the tomato reference genome assembly version 2.50 (Tomato Genome Consortium, 2012; https://www.solgenomics.net). A set of primers was designed to amplify all of the exons and introns as well as 2 kb of the upstream region from the start codon using the Primer3Plus version 2.4.0 (Untergasser et al., 2012). PCR amplification was performed in a total reaction volume of 50 µL that contained 50-100 ng genomic DNA, 0.25 µM each primer, 0.2 mM dNTPs, 1X PCR buffer, and 0.5 U DNA Taq polymerase (Geneslabs, Seongnam, Korea). The amplification profile consisted of an initial denaturation for 3 min at 94°C followed by 40 cycles of 30 s at 94°C, 30 s at an selected annealing temperature (50°C to 60°C depending on the melting temperature of primers), 45 s at 72°C, and one cycle of 7 min at 72°C as a final extension step. The amplified product was separated on a 1.5% agarose gel containing 1X RedSafeTM Nucleic Acid Staining Solution (20,000X) (JH Science, Lynnwood, WA, USA) in 1X TBE at 200 V for 1 h. The target DNA was collected and purified using the MinElute® Gel Extraction Kit (QIAGEN, Valencia, CA, USA) following the manufacturer’s instructions and resulted in samples that were used for Sanger sequencing (Cosmogenetech, Seoul, Korea). The resulting DNA sequences were trimmed and aligned using the Staden Package software (Bonfield et al., 1995) and Clustal Omega tool (http://www.ebi.ac.uk/Tools/msa/clustalo/). Sequence variations between the mutants (‘LA1793’ and ‘LA3013’) and wild-type (‘San Marzano’) varieties were identified by visual inspection of the sequence alignment.
High-resolution Melting Analysis in the Germplasm Collection
The primers for high-resolution melting (HRM) analysis were designed to produce an expected product size between 100-300 bp using the Primer3Plus version 2.4.0. The primer sequences were then analyzed using In Silico PCR at the SOL Genomic Network (https://solgenomics.net/tools/in_silico_pcr) to detect any possible unspecific products, ensuring the PCR amplification efficiency and HRM accuracy. HRM analysis was performed in a total volume of 20 µL using the LightCyler96 Real-Time PCR System (Roche Diagnostics, Indianapolis, IN, USA). The reaction mixture contained 50 ng qualified genomic DNA, 0.25 µM forward and reverse primers, and LightCycler® 480 HRM Master Mix 1X (Roche Diagnostics). The amplification was achieved by the following the protocol: 10 min initial denaturation at 95°C, 45 cycles of 10 s for denaturation at 95°C, 15 s at 55°C, 15 s at 72°C, followed by denaturation at 95°C for 15 s, 50°C for 10 s, 50°C for 50 s, 97°C for 1 s, with HRM ramping between 50°C and 97°C at 2.2°C/s increments.
Results and Discussion
Discovery of Sequence Variation in the nor Gene in Tomato
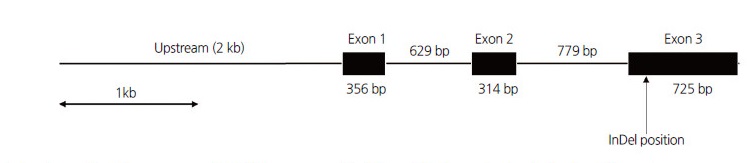
The nor gene originated from a spontaneous mutation in tomato on the short arm of chromosome 10 (Giovannoni et al., 1995). This gene has an open reading frame (ORF) consisting of three exons (356, 314, and 725 bp) and two introns (629 and 779 bp) (Fig. 1). Our eight primer sets produced PCR amplicons between 482-810 bp for the upstream and ORF regions of the nor gene in the two mutants ‘LA1793’ and ‘LA3013’, and a wild type ‘San Marzano’. The 4.8 kb sequences (2.0 kb upstream and 2.8 kb ORF) were generated from these amplicons and aligned to identify single nucleotide polymorphism (SNP) and/or insertion and deletion (InDel). We detected a 2 bp InDel (AA/--) within the third exon, which was the longest exon of the nor gene (Fig. 2). This InDel causes a frame-shift that results in change of the polypeptide sequence from the glutamine at the amino acid 183 (Fig. 3). This mutation is the only sequence variation associated with fruit ripening in the nor gene. The rin mutation is also derived from a frame-shift in a MADS-box gene of the SEPELATTA clade (Hileman et al., 2006). For the Cnr gene, however, mutant phenotype is due to a spontaneous epigenetic change in the SBP-box (SQUAMODA promoter binding protein-like) promoter (Manning et al., 2006). The molecular mechanism of the nor gene for regulating fruit ripening still remains unclear compared to the rin and Cnr genes in tomato. Therefore, the InDel identified in our study can be useful to elucidate the nor gene function in the ripening process.

Fig. 2.
Sequence alignment of part of the 3rd exon of the nor gene between wild-type (Heinz 1706 and San Marzano) and mutant (LA1793 and LA3013) varieties. The “*” indicates identical nucleotides between these sequences. The 1st codon of the 3rd exon is indicated with bold letters and underlined. The gray boxes represent the HRM primer sequences.
Table 1. The eight primer sets used to amplify the nor gene
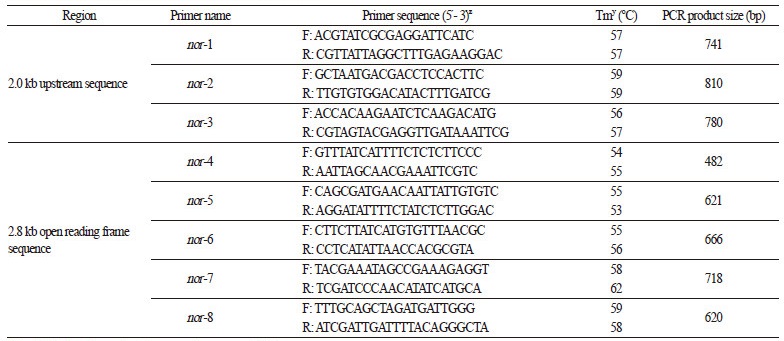 | |
zF and R indicate forward and reverse primers, respectively. yTm = melting temperature. | |
Gene-based Marker Development and Genotyping in a Collection of Tomato Varieties
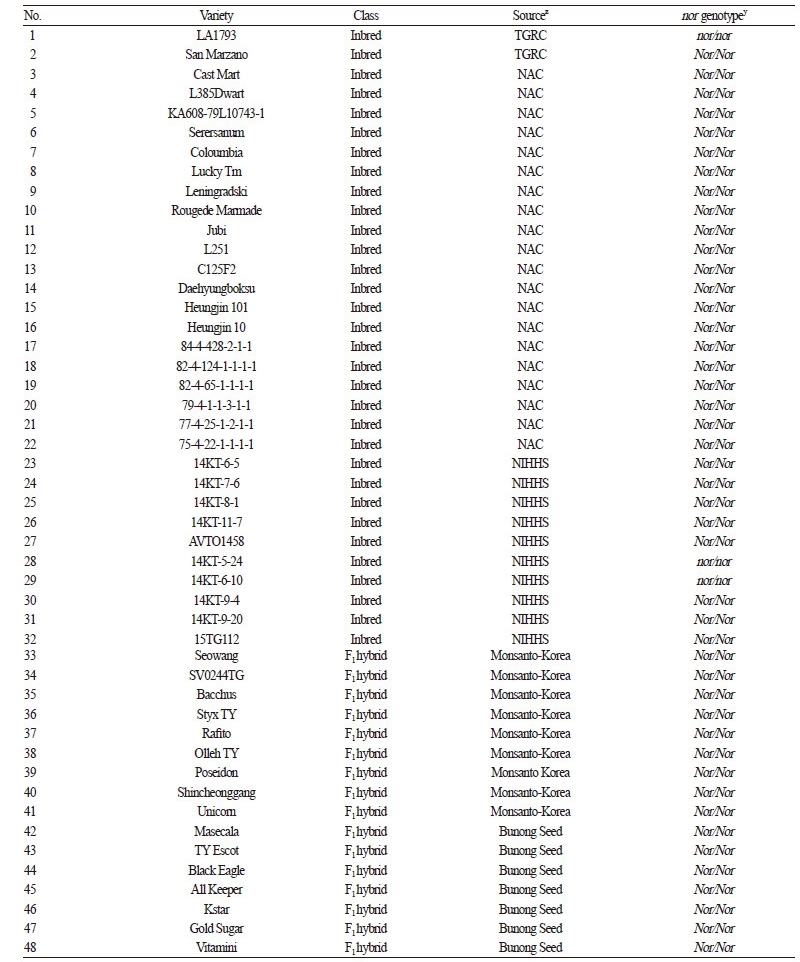
Based on the InDel in the 3rd exon, a primer set for a nor-InDel marker (forward: 5ʹ-TGCAGCTAGATGATTGGGTTT-3ʹ, reverse: 5ʹ-ACCTCCATGCATCGTTCTCT-3ʹ) was designed for HRM analysis. This primer set produced a 269 bp amplicon at an annealing temperature of 55°C and was used to genotype a collection of 83 tomato varieties including ‘LA1793’ and ‘San Marzano’. The InDel marker clearly differentiated the mutant allele (nor) from the wild type allele (Nor) (Table 2). Among these varieties, the homozygous form of nor was found in the two inbreds, ‘14KT-5-24’ and ‘14KT-6-10’ derived from National Institute of Horticultural & Herbal Science (NIHHS) at Rural Development Administration (RDA). Among the 51 commercial F1 hybrids, ‘Gayachal Plus’ from Syngenta-Korea had the heterozygous mutation (nor/Nor) (Table 2). The ‘14KT-5-24’ variety produces yellow fruit, whereas the ‘14KT-6-10’ and ‘Gayachal Plus’ varieties have pink fruit. Since abnormally ripened fruits that are green or yellow in color and common in the homozygous nor mutants (Tigchelaar et al., 1973), it was unexpected that the ‘14KT-6-10’ variety has the nor/nor genotype. Although the ‘14KT-6-10’ variety produces pink fruit, its fruit is firmer relative to wild type varieties with normally ripened fruit, which is similar to the other inbred ‘14KT-5-24’ variety. Fruit firmness is an important factor affecting storage period and fruit quality after harvesting (Batu, 2004; De Ketelaere et al., 2004). This trait is mainly enhanced by cellulose and its cross-linkage with hemicellulose, pectin, and lignin (Vicente et al., 2007). The two cellulose synthases are more abundant in the nor mutant relative to a wild-type, indicating that the nor mutation has a positive effect to improve fruit firmness (Seymour et al., 2013). Tomato varieties with the heterozygous form of nor recovers normal fruit color but enhances fruit firmness. Thus, the heterozygous genotype is favorable for improving fruit quality and shelf-life in elite tomato varieties. Our result supports the effect of the nor mutation on fruit firmness and demonstrates that the nor-InDel marker is effective in detecting the nor mutation.
Table 2. Genotypes of 83 tomato varieties via high-resolution melting (HRM) analysis using the gene-based marker developed for the nor gene
 |
One of the main challenges in the tomato industry is to deliver fresh fruits of high quality (e.g. desirable taste, texture, and color). Fruit ripening is a complex process that involves numerous physiological and biochemical changes in color, carotenoid biosynthesis (Pék and Helyes, 2010), sugar content (Davies and Kempton, 1975), firmness, and pathogen resistance (Kramer et al., 1992; Wang et al., 2017). Therefore, the regulation of the ripening process is central to improve fruit quality in tomato. Many efforts have been made in the genetic dissection of the ripening process and a number of ripening genes have been identified. Of these genes, spontaneous frame-shift mutations in the rin and nor genes exhibit similar phenotypes in fruit ripening. The individuals that are heterozygous at these loci recover normal color and have firmer fruit relative to plants that are homozygous form of wild type alleles. With this knowledge, the use of these mutations has helped to develop elite tomato varieties in breeding programs. Among the 81 tomato varieties used in this study, only two inbred and one F1 hybrid varieties have homozygous or heterozygous forms of the nor mutation, suggesting that this mutation is not widely used in tomato breeding programs. This may be due to a lack of an efficient molecular tool to facilitate introgression of this mutation into elite breeding lines.
Selection based on phenotypes often requires intensive field evaluations to eliminate environmental variation. Marker-assisted selection (MAS) is an effective approach that can accelerate plant breeding because breeding lines with favorable alleles are selected based on their genotypes, which are neutral for environmental variation. The molecular mechanism of rin in the ripening process have been studied more relative to nor (Vrebalov et al., 2002; Yuan et al., 2016), and the SCAR marker associated with the rin mutation was also developed for MAS (Kim et al., 2013). In the present study, an InDel responsible for a frame-shift mutation was detected in the nor gene and used to develop a gene-based marker that can be applied to MAS for the identification of this mutation. With this marker, we determined the genotype of nor in a collection of 81 tomato varieties representing 30 inbreds and 51 commercial F1 hybrids. Our results benefit the tomato research community, particularly breeders, by utilizing MAS for improving shelf-life and quality in tomato fruits.