Introduction
Materials and Methods
Plant Material
Cloning of EpDWF3
RNAi Vector Construction
Agrobacterium-mediated Transformation
Genomic DNA PCR
Semi-quantitative RT-PCR and Quantitative RT-PCR
Results and Discussion
Introduction
Echinacea purpurea (eastern purple coneflower or purple coneflower) is a North American species of herbaceous flowering plant that blooms from early to late summer. Echinacea was traditionally used as an indigenous medicine by Native American Indians: externally for wounds, burns, and insect bites and internally for pain, cough, stomach cramps, and snakebites (Wang et al., 2014). Currently, E. purpurea is economically important to the pharmaceutical industry due to its immune-stimulating effects. E. purpurea is also an ornamental plant that has received considerable attention in the horticulture industry (Wang et al., 2014).
Brassinosteroids (BRs) are poly-hydroxylated steroidal plant hormones that play important roles in plant growth and development (Gudesblat and Russinova, 2011). BRs influence diverse developmental and physiological processes in plants, including embryogenesis, cell elongation, cell division, vascular differentiation, male fertility, leaf senescence, photomorphogenesis, and responses to biotic and abiotic stresses (Guo et al., 2013). BR biosynthesis involves a highly branched network including two parallel pathways, the early and late C-6 oxidation pathways. These pathways are connected at multiple steps, including reduction, hydroxylation, epimerization, and oxidation (Fujioka and Yokota, 2003). The early and late C-6 oxidation pathways are also linked to the early C-22 oxidation step of a campesterol-independent pathway resulting in the convergence of castasterone, which ultimately leads to the biosynthesis of brassinolides. Defects in BR biosynthesis or loss-of-function mutations of BR-biosynthetic genes result in dwarf plants showing multiple developmental defects, such as extreme dwarfism, reduced seed germination, delayed flowering time, and male sterility (Clouse et al., 1996; Li et al., 1996, Zhu et al., 2013).
DWARF3 (DWF3), also known as constitutive photomorphogenesis and dwarfism (CPD) and cytochrome P450 90A1 (CYP90A1), is a BR-biosynthetic enzyme that catalyzes the C-3 oxidation of early BR intermediates in Arabidopsis thaliana (Ohnishi et al., 2012). DWF3 converts (22S) -22-hydroxycampesterol to (22S) -22-hydroxycampest-4-en-3-one via the CN-independent pathway. DWF3 is also involved in converting (22R, 23R) -22,23-dihydoxycampesterol to (22R, 23R) -22, 23-dihydroxycampest-4-en-3-one. To the best of our knowledge, the BR biosynthesis-related DWF3 ortholog in E. purpurea has not yet been identified.
Dwarf plants have many advantages for horticultural and agricultural applications, including dense growth and strong resistance to wind and rain damage (Sakamoto and Matsuoka, 2004). The reduced plant height of dwarf plants can lead to improved harvests and enhanced biomass production, in combination with the application of large amounts of nitrogen fertilizer (Khush, 1999). Several Echinacea varieties have been developed for improved flower quality and plant morphology (Wang et al., 2014). However, the development of new Echinacea varieties with dwarf phenotypes via RNA interference (RNAi) has not yet been reported. In this study, we isolated the DWF3 (EpDWF3) gene from E. purpurea. We also developed transgenic dwarf Echinacea plants via RNAi to suppress the expression of EpDWF3.
Materials and Methods
Plant Materials
Seeds of Echinacea purpurea purchased from ACC KA Seeds & Seedling Co., Ltd. (Gunposi, Korea) were surface steri-lized in 70% (v/v) ethanol for 5 min and 1.2% sodium hypochloride solution for 20 min and washed thoroughly with sterile deionized water. The seeds were germinated on basal E1 medium [MS salt, 2% sucrose, 100 mg·L-1 myo-inositol, 2 mg·L-1 glycine, 3 mL·L-1 Plant Preservative MixtureTM (PPM; Plant Cell Technology, Washington, DC, USA), and 0.7% Micro-agarTM (Sigma-Aldrich, St. Louis, MO, USA)]. The pH value of the medium was adjusted to 5.7. The cultures were incubated for 3 weeks at 25°C in the dark and transferred to a 25°C culture room under a 16-h light (100 µmol·m-2·s-1)/8-h dark cycle.
Cloning of EpDWF3
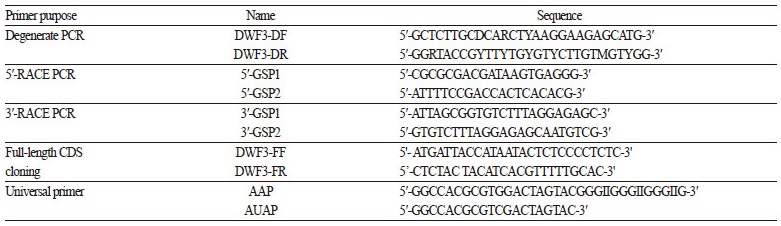
Total RNA was extracted from young leaves of E. purpurea using a Hybrid-RTM kit (GeneAll Biotechnology, Seoul, Korea). Two micrograms of total RNA was treated with amplification-grade RNase-free DNase I (Invitrogen, Carlsbad, CA, USA) and used for cDNA synthesis with an Improm-IITM Reverse Transcription System (Promega, Madison, WI, USA). Reverse transcription was performed following the manufacturer’s protocol in a 20 μL reaction mixture containing oligo(dT) and random hexamer. An EpDWF3 DNA fragment was obtained from degenerate polymerase chain reaction (PCR) using an LA Taq Polymerase kit (Takara, Japan) and degenerate PCR primers (DWF3-DF and DWF3-DR; Table 1) designed based on the high similarity regions of DWF3 genes from other species. The degenerate PCR product was cloned into the T/A cloning vector, pGEM-T easy (Promega). Following DNA sequence analysis of the PCR fragment, 5′ and 3′ rapid amplification of cDNA ends (RACE) by PCR was performed using a GeneRacerTM kit (Invitrogen) according to the manufacturer’s protocol. The EpDWF3-specific primers 5′-GSP1 and 5′-GSP2 were used for 5′ RACE, and the EpDWF3-specific primers 3′-GSP1 and 3′-GSP2 were used for 3′ RACE (Table 1). The final RACE products were cloned into pGEM-T-easy and sequenced. Full-length EpDWF3 DNA fragments including the 5′ UTR, amino acid coding region, and 3′ UTR were amplified by PCR using DWF3-FF and DWF3-FR primers (Table 1) and cloned into pGEM-T easy, generating pGEM-T-EpDWF3. The identity of EpDWF3 was confirmed by restriction enzyme mapping and DNA sequence analysis.
RNAi Vector Construction
An EpDWF3 DNA fragment (410 bp) corresponding to nucleotides 937–1,346 of the amino acid coding region was amplified by PCR from pGEM-T-EpDWF3. The forward and reverse primers were as follows: EpDWF3 forward, 5-CCCGGGGGCGCGCCGAAATTAGAAAGAAAAAGGGTGC-3′; EpDWF3 reverse, 5′-TCTAGAATTTAAATACCAACTTATCTTCTTCTGC-3′. The amplified EpDWF3 DNA fragment was cloned into pGEM-T easy. The identity of the PCR fragment was confirmed by restriction enzyme mapping and DNA sequence analysis. To generate the pFGC5941-EpDWF3 RNAi construct (Fig. 2), the EpDWF3 fragments were inserted upstream and downstream of the chalcone synthase A (CHSA) intron in sense and antisense orientation using the AscI-SwaI and XbaI-SmaI restriction sites, respectively, of the pFGC5941 vector (http://www.chromdb.org/rnai/vector_info.html), a binary vector designed to produce double-stranded RNA in plants. The proper orientation and reading frame of the insertion in pFGC5941-EpDWF3 were confirmed by restriction enzyme mapping and DNA sequence analysis.
Agrobacterium-mediated Transformation
Echinacea purpurea leaf disks (~1 cm2) were prepared from aseptically grown young seedlings. Agrobacterium tumefaciens LBA4404 cells harboring the pFGC5941-EpDWF3 RNAi construct were diluted to OD600= 0.8 with MS salt solution containing 2% sucrose. The leaf disks were immersed for 10 min in the diluted Agrobacterium suspension. Excess Agrobacterium was blotted from the leaf disks onto sterile filter paper. The leaf disks were placed on MS agar medium (MS salt, 2% sucrose, 0.7% Micro-agarTM) and incubated for 2 days in darkness. The leaf disks were transferred to E8 medium [MS salt, 100 mg·L-1 myo-inositol, 1 mg·L-1 glycine, 3 mL·L-1 PPM, 1.0 mg·L-1 benzylaminopurine (Sigma-Aldrich), 0.01 mg·L-1 naphthalene acetic acid (Sigma-Aldrich), 0.6 mg·L-1 phosphinothricin (PPT; Duchefa, Netherlands), and 250 mg·L-1 cefotaxime sodium salt (Duchefa)] and incubated in a 25°C culture room under a 16-h light/8-h dark cycle. The regenerated shoots were excised, transferred to rooting medium [McCown woody plant medium (Duchefa), 1.5% sucrose, 1.5 mg·L-1 indole-3-butyric acid], and incubated in a 25°C culture room under a 16-h light/8-h dark cycle. The rooted plantlets were transferred to potting soil.
Genomic DNA PCR
Genomic DNA was prepared from the leaves of wild-type and transgenic E. purpurea plants using a GenEluteTM Plant Genomic DNA Mini kit (Sigma-Aldrich). A pFGC5941-specific PCR fragment was amplified by PCR using an LA Taq Polymerase kit and pFGC5941-specific PCR primers. The forward and reverse primers were as follows: forward, 5′-CCATTCCTTAACTAGTAAATAC-3′; reverse, 5′-TCTAGACTCACCTAGGATCC-3′. The PCR products were separated on a 1% agarose gel and visualized under ultraviolet light (UV) after staining with ethidium bromide (EtBr).
Semi-quantitative RT-PCR and Quantitative RT-PCR
Total RNA was prepared from the leaves of wild-type and transgenic E. purpurea plants using a Hybrid-RTM kit. DNase I-treated total RNA was reverse-transcribed using the ImProm-IITM Reverse Transcription System and oligo-dT primer. The first-strand cDNA was then subjected to PCR amplification. The EpDWF3 gene was amplified using the primer set 5′-ATGATTACCATAATACTCTCCCCTCTC-3′ and 5′-CTCTACTACATCACGTTTTTGCAC-3′ to produce a full-length EpDWF3 fragment. As a control, the tubulin gene was amplified using the primer set 5′-CCGCTAACCTCACGTGGGTCCC-3′ and 5′-GCCTTCCTCCTGAACATAGC-3′. The amplification program consisted of an initial denaturation at 94°C for 5 min, followed by 40 cycles of 94°C, 1 min; 55°C, 0.5 min; and 72°C, 1.5 min, and a final extension at 72°C for 10 min. The experiments were repeated at least three times to obtain consistent results. The PCR products were separated on a 1% agarose gel and visualized under UV after staining with EtBr.
Quantitative reverse-transcription (RT)-PCR of EpDWF3 was performed in a 20 μL reaction volume containing 10 μL of SYBR Premix EX Taq II (Takara), 2 μL of RT reaction mixture, and 400 nM of each primer. The primers were as follows: EpDWF3 forward, 5′-ACCACCATGACTCTTGCCGT-3′; EpDWF3 reverse, 5′-AGAGTCACCGATGCACCCTT-3’; Tubulin forward, 5′-ATGATGTGTGCTGCTGACCC-3′; Tubulin reverse, 5′-TCGTCAACTTCCTTGGTGCTCA-3′. PCR was performed using an ABI 7500 Fast Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). The amplification program consisted of an initial denaturation at 95°C for 30 secs, followed by 40 cycles of 95°C for 15 secs, annealing at 55°C for 30 secs, and extension at 72°C for 30 secs.
Results and Discussion
We isolated EpDWF3, containing a 1,413 bp open reading frame encoding a 471 amino acid polypeptide from E. purpurea by degenerate and RACE PCR as described in the Materials and Methods. The sequence data have been submitted to the NCBI GenBank database under accession number KF170019.
We constructed a phylogenetic tree based on the amino acid sequence of EpDWF3 to investigate the genetic relationships between EpDWF3 and Arabidopsis CYP family proteins (Fig. 1A) via the neighbor-joining method using MEGA 6 App. EpDWF3 showed high similarity to AtCYP90A1 (or AtCPD), a BR-biosynthetic enzyme catalyzing the C-3 oxidation of early BR intermediates (Ohnishi et al., 2012). We also constructed a phylogenetic tree to investigate the genetic relationship between EpDWF3 and other plant CYP90A1 (or CPD) proteins (Fig. 1B). EpDWF3 sharesw 87% and 82% similarity with ZeCPD1 and AaCPD, respectively. ZeCPD1 is a steroid 23-alpha-hydroxylase that converts (22S) -22-hydroxycampesterol to (22S)-22-hydroxycampest-4-en-3-one and (22R, 23R) -22,23-dihydoxycampesterol to (22R, 23R) -22,23-dihydroxycampest-4-en-3-one in the BR-biosynthetic pathway (Yamamoto et al., 2007), suggesting that EpDWF3 is a steroid 23-alpha-hydroxylase involved in BR biosynthesis in E. purpurea.
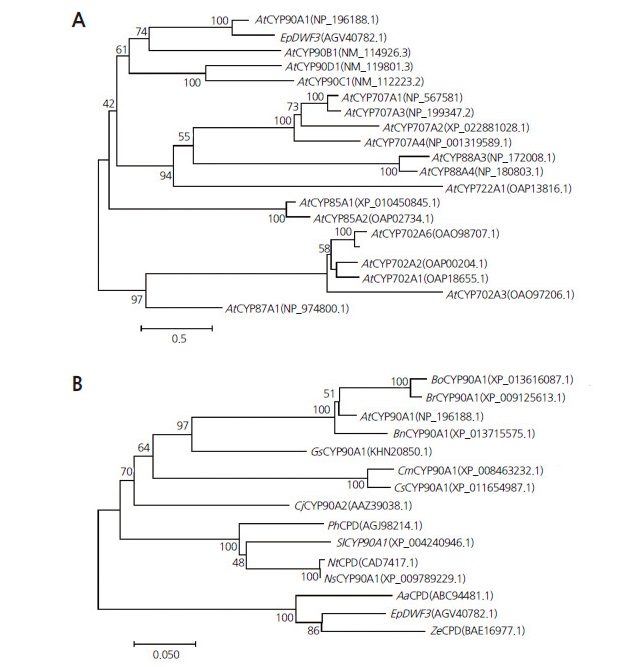
Fig. 1.
Phylogenetic tree analysis of EpDWF3. (A) Phylogenetic tree of EpDWF3 with CYP proteins from Arabidopsis thaliana (At). (B) Phylogenetic tree of EpDWF3 with CYP90A1 and CPD proteins from Artemisia annua (Aa), Brassica oleracea (Bo), Brassica rapa (Br), Brassica napus (Bn), Cucumis sativus (Cs), Cucumis melo (Cm), Camellia japonica (Cj), Glycine soja (Gs), Nicotiana tabacum (Nt), Nicotiana sylvestris (Ns), Petunia hybrid (Ph), Solanum lycopersicum (Sl), and Zinnia elegans (Ze).
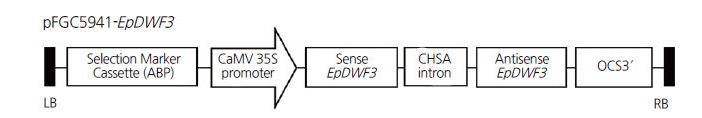
Fig. 2.
Schematic diagram of the RNAi vector construct (pFC5941-EpDWF3). A hairpin structure consisting of the sense EpDWF3 sequence, the CHSA intron (from the petunia chalcone synthase A gene), and the antisense EpDWF3 sequence was inserted between the cauliflower mosaic virus (CaMV) 35S promoter and the octopine synthase (OCS) terminator. LB, left border; RB, right border.
Dwarf plants have many advantages for use in horticulture, including dense growth and strong resistance to damage caused by wind and rain (Sakamoto and Matsuoka, 2004). Several knock-out mutants with defects in BR biosynthesis or loss-of-function mutations of BR biosynthetic genes show multiple developmental defects and dwarfism (Clouse et al., 1996; Li et al., 1996). The downregulation of BR biosynthesis-related genes by RNAi has been used to generate dwarf Arabidopsis plants (Chung et al., 2010), suggesting that this technique would be useful for generating valuable dwarf plants for commercial applications. We therefore employed RNAi to suppress the expression of endogenous EpDWF3 in Echinacea plants, resulting in dwarf phenotypes.
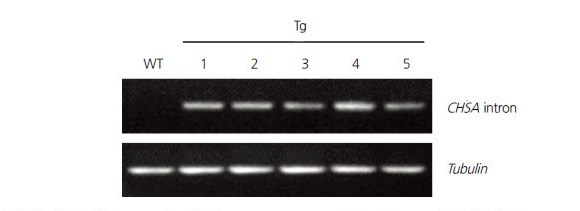
We constructed the RNAi vector, pFGC5941-EpDWF3, to downregulate the expression of EpDWF3. The sense and antisense EpDWF3 DNA fragments corresponding to a highly conserved region of DWF3 genes (a protein-encoding region of 937–1,346 nucleotides) were respectively inserted upstream and downstream of the CHSA intron of the pFGC5941 vector (Fig. 2). We transformed Echinacea leaf disks with recombinant Agrobacterium harboring pFGC5941-EpDWF3. We then transferred putative transgenic plants grown in culture medium containing a selective herbicidal marker, PPT, to potting soil and performed genomic DNA PCR to determine the presence of the RNAi-EpDWF3 construct (data not shown). Two transgenic plants that grew normally and formed stems and flowers were obtained from 39 putative transgenic plants harboring the RNAi-EpDWF3 construct (data not shown). These transgenic plants displayed a dwarf phenotype, with one of the two lines showing stronger dwarfism than the other. Different levels of dwarfism between transformation events are due to differences in the degree of gene downregulation (Chung et al., 2010). We propagated a transgenic plant that grew normally and displayed a strong dwarf phenotype by vegetative propagation using root division. Five transgenic plants were obtained by vegetative propagation and used for further analyses. We confirmed the presence of the RNAi-EpDWF3 construct in genomic DNA from the transgenic plants by PCR using CHSA intron-specific primers to amplify the pFGC5941-specific CHSA intron region (Fig. 3). The CHSA intron-specific PCR products were successfully amplified from genomic DNA from all transgenic plants, but not from wild-type, indicating that the selected transgenic plants harbored the RNAi-EpDWF3 construct in their genomic DNA.
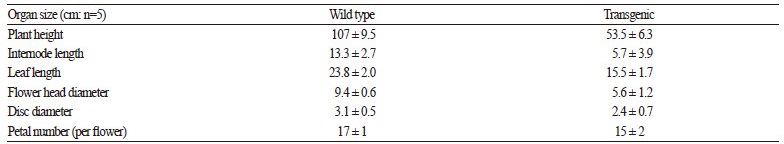
We investigated morphological differences between wild-type and transgenic plants. The transgenic plants exhibited a clear alteration in phenotype (Fig. 4). Specifically, these plants displayed dwarf phenotypes, with short inflorescences, shorter leaf blades, and shorter pedicels than wild type. E. purpurea is an herbaceous perennial that reaches up to 120 cm tall by 25 cm wide at maturity. The average inflorescence height of wild-type and transgenic plants was 107 (± 9.5) and 53.5 (± 6.3) cm, respectively. The flower diameter of transgenic plants was 5.6 (± 1.2) cm, whereas that of wild-type was 9.4 (± 0.6) cm (Table 2).
To quantify EpDWF3 transcript levels in transgenic plants showing dwarf phenotypes, we performed semi-quantitative RT-PCR using pooled RNA samples obtained from transgenic plants. EpDWF3 transcript levels were reduced in the transgenic plants compared to wild type (Fig. 5A). We also performed quantitative RT-PCR, finding that EpDWF3 transcript levels in the transgenic plants decreased to 52% that of wild type (Fig. 5B). Taken together, these findings indicate that EpDWF3 expression was suppressed by the RNAi technique in the transgenic plants, resulting in dwarf phenotypes.
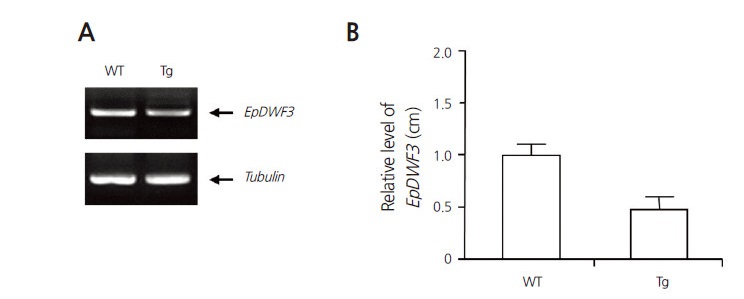
Fig. 5.
Semi-quantitative and quantitative RT-PCR analyses of wild-type and transgenic Echinacea purpurea plants. (A) Semi-quantitative RT-PCR was performed to confirm the downregulation of endogenous EpDWF3 in the transgenic plants. Tubulin was used as a control. (B) Quantitative RT-PCR was performed to confirm the downregulation of endogenous EpDWF3 in the transgenic plants. Relative levels of EpDWF3 transcript obtained from three independent experiments were plotted as a bar diagram.
The use of RNAi to produce knock-down mutants has an advantage over the production of knock-out mutants for commercial applications. A knock-out Arabidopsis mutant for the CPD gene generated by T-DNA tagging on chromosome 5-14.3 showed inhibited cell elongation, as controlled by BRs (Szekeres et al., 1996). The plants displayed de-etiolation and derepression of light-induced genes in the dark, as well as the activation of stress-regulated genes in the light. They also displayed dwarfism and male sterility, resulting in poor seed production. However, the seed productivity of the EpDWF3 knock-down plants produced in the current study was similar to that of wild type (data not shown), suggesting that producing knock-down mutants using this RNAi technique can be more useful for developing modified mutant plants than producing knock-out mutants that are completely deficient in function.
In this study, we identified the EpDWF3 gene from Echinacea. EpDWF3 shares high homology with AtCYPA1 and ZeCPD, suggesting that EpDWF3 is a steroid 23-alpha-hydroxylase involved in the BR-biosynthetic pathway in Echinacea. We employed an RNAi technique to downregulate the expression of EpDWF3. Transforming Echinacea plants with an RNAi construct suppressed the expression of EpDWF3, leading to dwarf phenotypes. Our results suggest that the knock-down of BR biosynthesis-related genes via RNAi enables valuable dwarf plants to be produced for commercial horticultural or agricultural applications.