Introduction
Materials and Methods
Plant Materials
Characteristic Analysis
Ploidy Analysis
Root Collection and Chromosome Preparation
Fluorescence in situ Hybridization (FISH) Analysis
Results and Discussion
Characteristic Analysis
Ploidy Analysis
Fluorescence in situ Hybridization (FISH) Analysis
Introduction
Hibiscus spp., belonging to the Malvaceae family, is a genus of famous ornamental plants with approximately 300 species distributed worldwide, including H.schizopetalus (East-African Coast), H. liliiflorus Cav. (Mauritius and Rodriguez Islands), H. fragilis, and H. boryanus Hook and Arn. (Reunion Islands). Species from the Pacific Islands, included H. arnottianus Gray, which is synonymous with H. wimeae Heller according to Roe (1961), and H. kokio Hillebrand (both from the Hawaiian Islands), H. storckii Seeman (Fiji), an ancient form of H. rosa-sinensis (Skovsted, 1941), and H. denisonii. Among the seven origins of Hibiscus spp., H. syriacus, H. hamabo, and H. mutabilis are native to Korea and are the most commonly cultivated ornamental plants in Korea. The East Asian group of Hibiscus spp., which inhabits Northeast Asian countries such as Korea, is highly tolerant to cold stress (Kang et al., 2016). H. syriacus shows the highest cold tolerance among Hibiscus species. Breeding programs to cross H. syriacus with other Hibiscus species help to develop new cultivars that vary in color and petal patterning. Moreover, H. syriacus (Yu and Yeam, 1987), H. paramutabilis, and H. sinosyriacus are commonly cultivated for ornamental purposes in Korea and some can be utilized in breeding programs (Van Laere et al., 2007).
H. syriacus has high research value because of its unique and numerous flowers that bloom during the summer season. However, few studies examined H. syriacus in Korea until the 1960s, when research began at Seoul National University and the Forestry Agency and the Rural Development Administration (Ha et al., 2015a). In recent years, H. syriacus has been widely examined. A number of varieties have been generated through introduced breeding, native breeding selection, cross breeding between breeds and varieties, mutation, diploid breeding, tissue culturing with irradiation isotopes, and colchicine treatment, but the germplasm can be further improved (Yu et al., 1976; Yu and Yeam, 1987).
Interspecific hybrid Hibiscus spp. Have exhibited poor growth and had survival rates of nearly 50% in the early stages of growth (Tachibana, 1958). Surviving F1 hybrids exhibited few characteristics of the parents and many showed poor growth. Additionally, in the case of H. syriacus, successful pollination occurred only between two species of H. sinosyriacus and H. paramutabilis belonging to the East Asian group. However, few seeds were obtained due to low seeding and germination rates. For these and other reasons, there are still many barriers limiting the development of new Hibiscus cultivars (Tachibana et al., 1957; Tachibana, 1971).
H. sinosyriacus ‘Malmouve’, first cultivated in France in 2001 was selected from hybrid seedlings of H. sinosyriacus ‘Lilac Queen’ (♀) × H. sinosyriacus ‘Ruby Glow’ (♂) (Ha et al., 2010). ‘Malmouve’ was introduced in Korea in 2003, where it was given the name ‘Seobong’. In Korea, ‘Hakbong’ was selected by crossing progenies between H. sinosyriacus ‘Seobong’ (♀) × H. syriacus ‘Namwon’ (♂) (Ha et al., 2014). Introduction of new interspecific hybrids between H. syriacus and H. rosa-sinensis, particularly to develop new petal colors and forms, have not yet been successful (Yu et al., 1976; Paek et al., 1989).
Hibiscus spp. are mainly polyploid in nature (Kim et al., 2016b). The chromosome number of Hibiscus spp. was reported to vary (x = 7, 8, 9, 11, 12, 15, 17, 19, 20, 39) between species and the precise number of somatic chromosomes remains unclear. The Hibiscus genus includes a large variety of plants with complex interspecific relationships (Li et al., 2015). The following ploidy levels and chromosome numbers were previously reported: H. schizopetalus; 2n = 42, H. mutabilis; 2n = 92, H. rosa-sinensis; 2n = 84, H. rosa-sinensis ‘Double Rainbow’; 2n = 105, H. rosa-sinensis ‘Flavo-plenus’; 2n = 138, and H. rosa-sinensis ‘Carminatus’; 2n = 147 (Song, 2001). Among these, H. schizopetalus is diploid (2n = 2x = 42), H. rosa-sinensis is tetraploid (2n = 4x = 84), H. rosa-sinensis ‘Double Rainbow’ is pentaploid (2n = 5x = 105), and H. rosa-sinensis ‘Carminatus’ is heptaploid (2n = 7x = 147). Therefore, the basic chromosome number of H. syriacus is x = 20 and most cultivars of H. syriacus are tetraploid; 2n = 4x = 80 (Skovsted, 1941). Moreover, H. sinosyriacus is tetraploid; 2n = 4x = 80 (Skovsted, 1941) and has broader leaves compared to H. syriacus. This species was bred to develop new Hibiscus cultivars with a unique flower shape, petal color, uniform plant habitat, and hardiness through interspecific hybridization between H. sinosyriacus and H. syriacus (Kyung et al., 2001a). The aim was to introduce increased vigor into H. syriacus through interspecific hybridization with H. sinosyriacus (Ha et al., 2014). A Hibiscus breeding program was initiated in 2005 to develop new Hibiscus cultivars with vigorous growth, a tall upright posture, compact branches, and unique flowers with a long red eye through interspecific hybridization between H. syriacus and H. sinosyriacus (Kyung et al., 2001b; Ha et al., 2015b).
Hybridization techniques, such as fluorescence in situ hybridization (FISH) and genomic in situ hybridization, can reveal the chromosome morphological distribution, sequences, and distribution of chromatin in the chromosomes as well as genome organization during the metaphase stage of meiosis (Younis et al., 2015). These approaches enable accurate identification of genome differences in hybrid derivatives, which may have variable chromosome numbers or chromosome arms. The characteristics of ‘Tohagolred’ were studied in 2010 and those of ‘Daewangchun’ were studied in 2012. Furthermore, the DNA content of the parents was investigated by flow cytometry. The 5S and 45S rDNA were used to analyze the parent and progeny karyotypes by FISH.
Materials and Methods
Plant Materials
A new Hibiscus cultivar ‘Daewangchun’ with vigorous growth, uniform plant habit, upright, compact branches, and a long red eye was developed through interspecific hybridization between H. syriacus ‘Samchully’ (♀) and H. sinosyriacus ‘Seobong’(♂) (Ha et al., 2015b) (Fig. 1). Another new Hibiscus cultivar, named ‘Tohagolred’, was generated through reciprocal crossing between H. sinosyriacus ‘Seobong’ (♀) and H. syriacus ‘Samchully’ (♂) to improve flower quality and growth traits in 2002 (Ha et al., 2014) (Fig. 2).
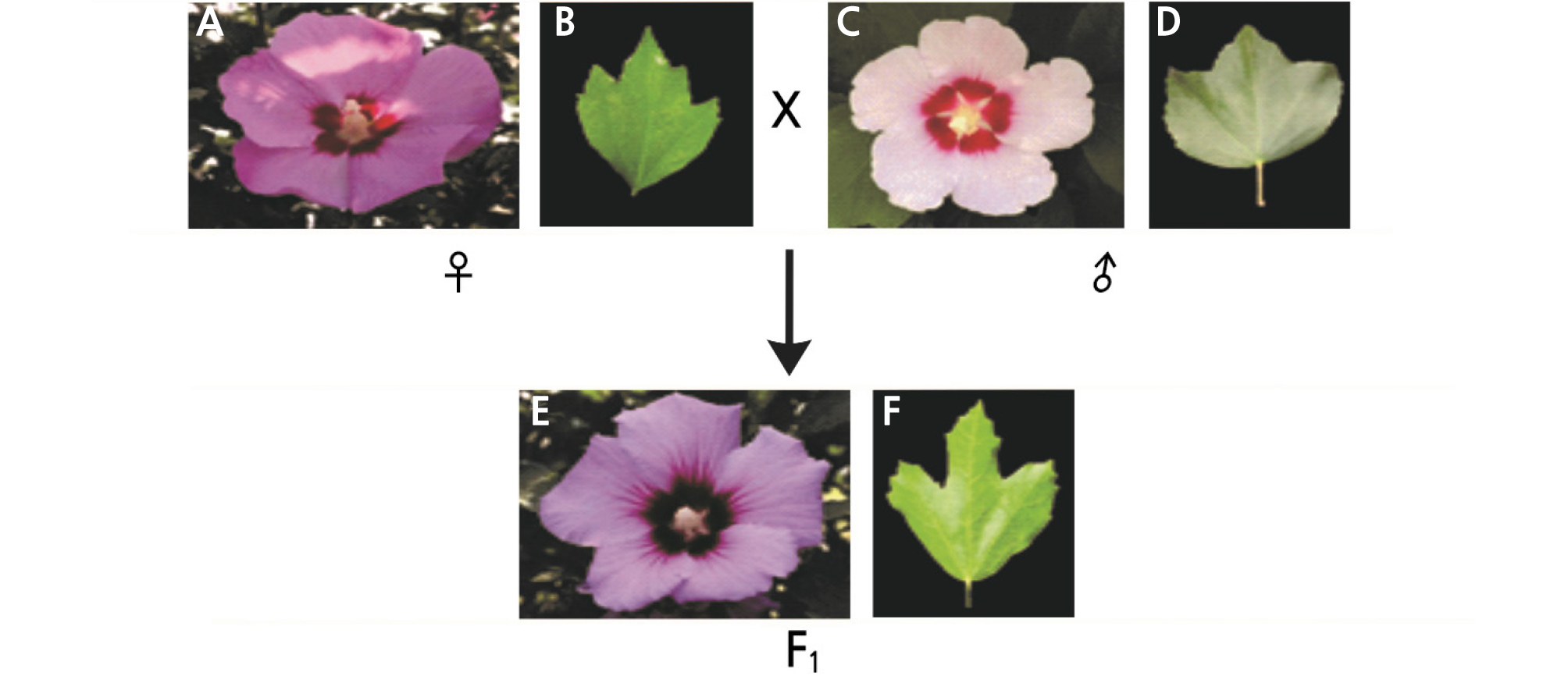

Fig. 1.
Flower and leaf shape of female, male plant and a new cultivar ‘Daewangchun’ through interspecific bybridization. (A) Flower of Hibiscus syriacus ‘Samchully’(♀), (B) Leaf of Hibiscus syriacus ‘Samchully’, (C) Flower of Hibiscus sino-syriacus ‘Seobong’(♂), (D) Leaf of Hibiscus sino-syriacus ‘Seobong’, (E) Flower of new cultivar ‘Daewangchun’, (F) Leaf of new cultivar ‘Daewangchun’.
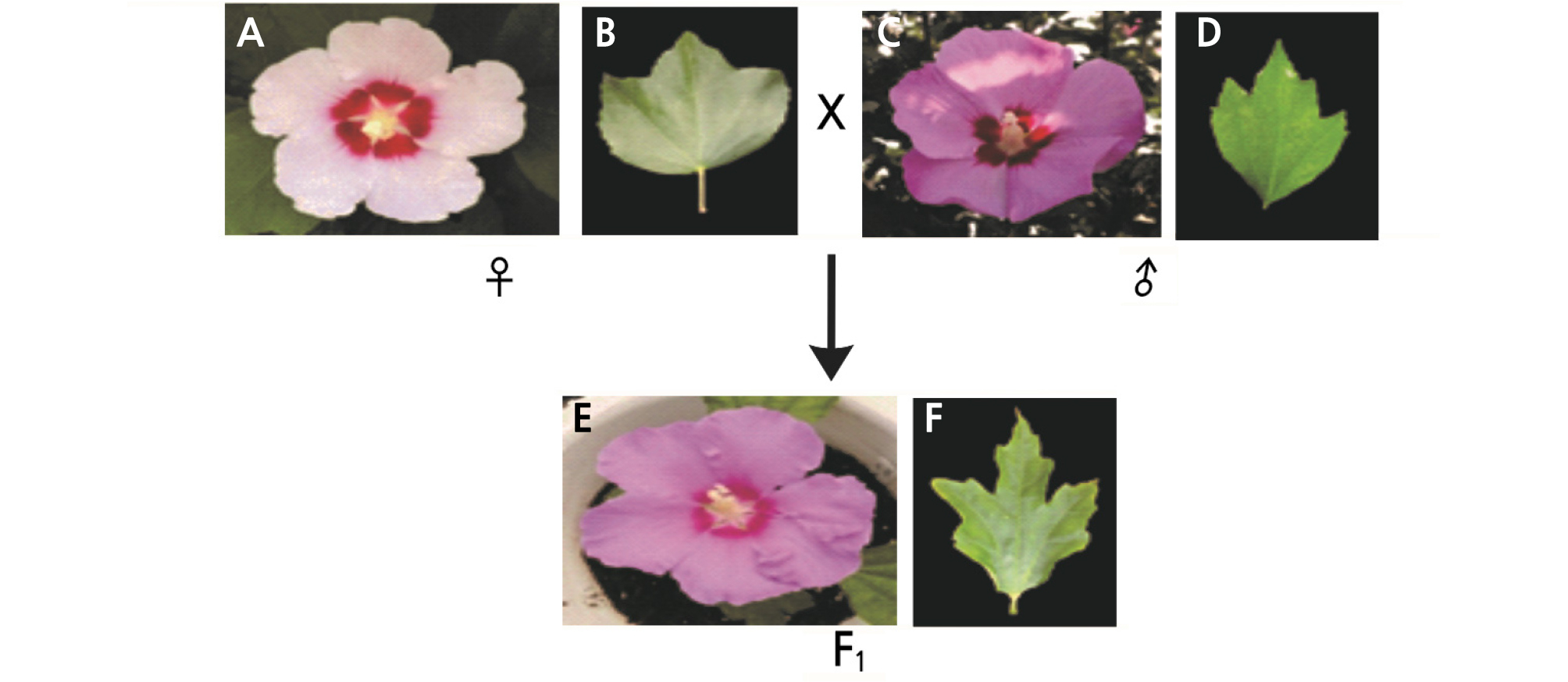

Fig. 2.
Flower and leaf shape of female, male plant and a new cultivar ‘Tohagolred’ through interspecific bybridization. (A) Flower of Hibiscus sino-syriacus ‘Seobong’(♀), (B) Leaf of Hibiscus sino-syriacus ‘Seobong’, (C) Flower of Hibiscus syriacus ‘Samchully’(♂), (D) Leaf of Hibiscus syriacus ‘Samchully’, (E) Flower of new cultivar ‘Tohagolred’, (F) Leaf of new cultivar ‘Tohagolred’.
Characteristic Analysis
Vegetative and flower characteristics were investigated according to the survey method of the Korean Seed & Variety Service (http://www.seed.go.kr). According to the Korean Seed & Variety Service, the characteristics of H. syriacus are shown in Table 1 as well as Figs. 3 and 4.
Table 1. Investigation of Hibiscus spp. leaf and flower
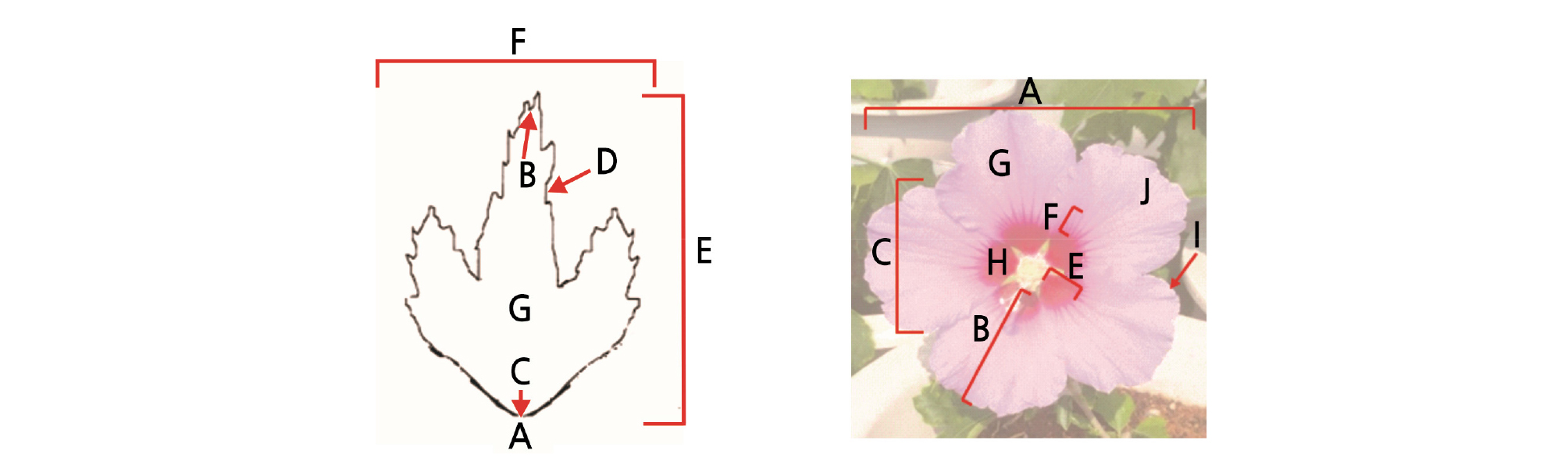
Fig. 3.
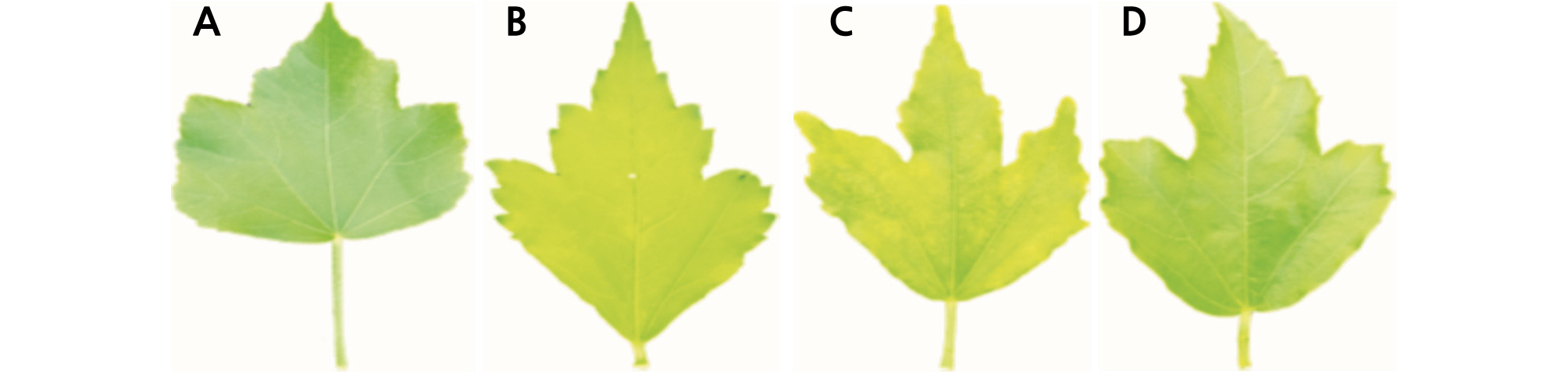
The Characteristic investigating items of H. syriacus leaf. A: Shape, B: Apex, C: Base, D: Margin, E: Length, F: width, G: Color. In flower A: Flower diameter, B: Petal length, C: Petal width, D: Petal index (B/C), E: Length of red eye, F: Length of radiation line, G: Flower color, H: Eye spot color, I: Serration of petal, J: Petal shape.
Ploidy Analysis
DNA content was measured by flow cytometry (Partec PA, Ploidy Analyzer, Sysmex, Kobe, Japan) with fresh young leaves from each cultivar. To release the nuclei, the center of the leaf was cut to approximately 1 cm2 and chopped in nuclei extraction buffer (Sysmex) in a petri dish and then shaken for 30 - 40 s. The nuclei were filtered through a tube and then staining buffer (Sysmex) was added through a nylon mesh filter. Finally, the solution was injected into the flow cytometry analyzer.
Root Collection and Chromosome Preparation
Root tips were collected and treated with 1.0 M α-bromonaphthalene solution for 1 - 2 h at room temperature. The root tips were washed with double-distilled water and transferred into an aceto-ethanol (1:3, v/v) solution to fix the roots overnight. After rinsing with distilled water, 70% ethanol was used to preserve the roots at - 20°C until further use. For slide preparation, root tips were washed with double distilled water to remove the ethanol solution and incubated with enzyme mixture consisting of 0.3% pectolyase (Duchefa, Haarlem, The Netherlands), 0.3% cellulose (Duchefa) and 0.3% cytohelicase (Sigma, St. Louis, MO, USA) at 37°C for 40 min. Digested root tips were transferred to a clean slide and squashed under a microscope. Finally, 17 µL acetic acid (70%) per slide was added and the slides were air-dried.
Fluorescence in situ Hybridization (FISH) Analysis
FISH was performed as described by Lim et al. (2001) with some modifications. Briefly, the slides were pretreated with 100 µg·mL-1 RNase A at 37°C for 40 min. After rinsing with 2X SSC and fixation with 4% paraformaldehyde for 10 min, the slides were rinsed again with 2X SSC and dehydrated in an ethanol series (70%, 90%, 100%), followed by air-drying. The hybridization mixture contained formamide, 50% dextran sulfate, 20X SSC, 10% sodium dodecyl sulfate, salmon sperm, and rDNA probes. The mixture was placed in a water bath for DNA denaturation at 70°C for 5 min and then placed on ice for 15 min for fixation. To each slide, 40 µL of the mixture was added and the coverslip was placed on the slide. The slides were placed in a water bath for hybridization at 80°C for 5 min, placed in a container with wet tissue paper, and incubated in a humid chamber at 37°C for 16 h. The slides were washed with 2X SSC buffer for 5 min once for the rDNA probes, 0.1X SSC buffer for 30 min at 42°C with shaking water bath, and 2X SSC buffer for 5 minutes once again. The slides were submerged in 1X detection buffer for 5 min. Streptavidin conjugated CY3 (Invitrogen, Carlsbad, CA, USA) and anti-digoxigenin fluorescein (Roche, Basel, Switzerland) were used to detect the labeled chromosomes. The slides with the coverslip were incubated in a humid chamber at 37°C for 1 h. Three jars containing 1X detection buffers were incubated in the dark in a water bath at 37°C for 5 min followed by incubation in an ethanol series (70%, 90%, and 100%). The slides were air-dried in in the dark and then counterstained with DAPI (4'6-diamidino-2-phenylindole)/Vectashield (3:200) (Vector Laboratories, Burlingame, CA, USA). The slides were examined with a model Nikon BX 61 fluorescence microscope (Tokyo, Japan). Signals were analyzed using ultraviolet excitation filters. Cytovision and imaging software were used to acquire images of the chromosomes with 45S rDNA and 5S rDNA signals.
Results and Discussion
Characteristic Analysis
Leaf characteristics are shown in Table 2. ‘Samchully’ had an elliptical leaf shape and an acute leaf base. The leaf apex was acuminate and leaf margin was serrate. The leaf length and width were 6.8 cm and 3.6 cm respectively and leaf color was dark green. Moreover, ‘Seobong’ was circinate in leaf shape and round at the leaf base. The leaf apex was acute and the leaf margin was crenate. The leaf length was 8.2 cm, while leaf width was 7.6 cm, and leaf color was dark green. However, the Hibiscus cultivar ‘Daewangchun’ had an oval leaf shape and round leaf base. The leaf apex was acute and the leaf margin was crenate in shape, similar to ‘Seobong’ (♂). The leaf length was 8.2 cm and the leaf width was 5.4 cm, which are larger than the values for ‘Samchully.’ The leaf was green in color. Similarly, the Hibiscus cultivar ‘Tohagolred’ had an oval leaf shape, round leaf base, and acute leaf apex, which was similar to ‘Seobong’ (♂). The leaf margin was crenate, similar to ‘Seobong’ (♀). The leaf length was 7.2 cm and leaf width was 5.7 cm. The size was an intermediate relative to the female parent. The leaf was green in color, similar to ‘Daewangchun’. Among the Hibiscus cultivars, ‘Joomong’ produces the largest leaves, 13.1 cm in length, followed by H. syriacus ‘Moonwon’, ‘Minerva’, ‘Samchully’, ‘Reddish Ball’, and Hibiscus hybrid ‘Daeil’, which produce leaves of 10.0 cm. Cultivars with smaller leaf sizes (> 5.0 cm) include Hibiscus hybrid ‘Arirang’, Hibiscus hybrid ‘Daemang’, and H. syriacus ‘Andong’ (Kim et al., 2016a).
Table 2. Leaf characteristics of the reciprocal F1 cultivars and their parents in Hibiscus spp.
Analysis of flower characteristics revealed that the color of ‘Samchully’ was pink, while the flower diameter was 11.4 cm (Table 3). Petal length and width were 6.2 and 4.1 cm, respectively. The eye spot color was dark red and the red eye length was 1.6 cm (Table 4). The color of ‘Seobong’ appeared as light pink and the flower diameter was 10.4 cm. Petal length and width were 6.1 and 4.7 cm, respectively. The eye spot color was dark red and the red eye length was 1.8 cm. The color of ‘Daewangchun’ was pink, similar to ‘Samchully’ (♀). The flower diameter was 13.8 cm, which was 2 cm larger than that of the parents. Kim (2016a) experimented with 127 Hibiscus cultivars and reported that Hibiscus hybrid ‘Daewangchun’ had the largest flower which was 16.0 cm in diameter, while H. syriacus ‘Ggoma’, ‘Mibeak’, ‘Andong’, ‘Andong II’, and ‘Eunhasu’ had the smallest flowers, each being about 6.2 cm. Table 3 showed, the petal length and width were 8.4 cm and 6.9 cm, respectively, which were larger than those of the parents. The eye spot color was dark red and the red eye had a radius of 2.1 cm, which was approximately 1 cm larger than that of the parental lines. The color of ‘Tohagolred’ was pink, similar to ‘Samchully’ (♂). The flower diameter was 12.6 cm, which was 1 cm larger than ‘Samchully’ and 2 cm larger than ‘Seobong’.The petal length and width were 7.7 and 6.5 cm respectively. The flowers of ‘Tohagol Red’ and ‘Daewangchun’ somewhat resembled those of the species H. sinosyriacus and H. syriacus in that the flowers expanded their petals widely. The oval three-lobed leaves of ‘Daewangchun’ also resembled those of H. sinosyriacus in shape. The stiff, upright main stems of Hibiscus ‘Tohagol Red’ and ‘Deawangchun’ seemingly were derived from H. syriacus. H. sinosyriacus was similar to H. syriacus, but had broader leaves, with three short triangular lobes. The eye spot color of ‘Tohagolred’ was light red and the red eye had a radius of 1.1 cm, which was approximately 1 cm smaller than that of the parents. By screening the morphological features, we found that the hybrids ‘Tohagolred’ and ‘Daewangchun’ were intermediate in nature compared to their parents. Screening the morphological features it was found that hybrid ‘Tohagolred’ and ‘Daewangchun’ were intermediate in nature as compared to their parents.
Table 3. The quantitative characteristics of flower and red eye spot of the reciprocal F1 cultivars and their parents in Hibiscus spp.
Table 4. The qualitative characteristics of flower and red eye spot of the reciprocal F1 cultivars and their parents in Hibiscus spp.
Ploidy Analysis
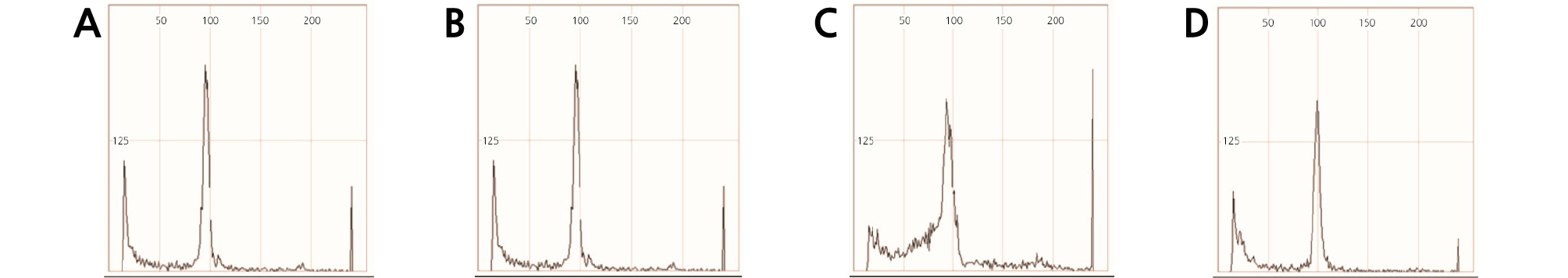
Flow cytometry is frequently used for hybrid identification (Dolezel et al., 2007) and to measure nuclear DNA content in plants (Doležel et al., 2007). In this study, the DNA content of triploid H.syriacus ‘Samchully’ (2n = 3x = 136), diploid H. sinosyriacus ‘Seobong’ (2n = 2x = 80), and their F1 progenies ‘Tohagolred’ and ‘Daewangchun’ were measured by flow cytometry (Fig. 5 and Table 5). We found that the DNA content of ‘Seobong’ and ‘Samchully’ were 1928.77 Mbp/1C (1.97 pg/1C), and 2820.34 Mbp/1C (2.88 pg/1C), respectively. In contrast, ‘Daewangchun’ and ‘Tohagolred’ had DNA contents of 1914.70 Mbp/1C (2.02 pg/1C) and 1975.88 Mbp/1C (1.96 pg/1C), respectively. DNA content of F1 ‘Daewangchun’ was slightly lower than H. sinosyriacus ‘Seobong’ (♀), but significantly lower than H.syriacus ‘Samchully’ (♂). However, in reciprocal cross the F1 ‘Tohagolred’ showed higher DNA content than H. sinosyriacus ‘Seobong’ but lower than H. syriacus ‘Samchully’.
Table 5. Genome size (Mbp/1C) and DNA content (pg/1C) of the reciprocal F1 cultivars and their parents in Hibiscus spp.
| Cultivar | 1C DNA content (Mbp)Cultivar | pg (1C)z |
| Seobong (A) | 1928.77 | 1.97 |
| Samchully (B) | 2820.34 | 2.88 |
| Tohagolred (A × B) | 1975.88 | 2.02 |
| Daewangchun (B × A) | 1914.70 | 1.96 |
The values are given as mean of the nuclear DNA content (pg/1C) and as mean of the 1C genome size in Mbp.
Fluorescence in situ Hybridization (FISH) Analysis
The FISH analysis results of H. syriacus ‘Samchully’ chromosomes are shown in Fig. 6 where the number of chromosomes of ‘Samchully’ was 2n = 3x = 136 with six 45S rDNA signals (indicated as red signals) and three 5S rDNA signals (indicated as green signals). All of the 45S rDNA signals were found on the short arms while the 5S rDNA signals were present on the long arms. Fig. 6B and 6C showed the aligned chromosomes with the 45S rDNA and 5S rDNA signals. The chromosomes with the 45S rDNA signals were divided into two parts, with three chromosomes in each part. However, the three chromosomes showing the 5S rDNA signals (Fig. 6C) were similar to the chromosomes with the 45S rDNA signals. Therefore, H. syriacus ‘Samchully’ was assumed to be triploid (2n = 3x = 136). Fig. 8 shows FISH analysis results of H. sinosyriacus ‘Seobong’ chromosomes where the number of chromosomes for ‘Seobong’ was 2n = 80, with six 45S rDNA signals (indicated as red signals) and two 5S rDNA signals (indicated as green signals). Four out of 45S rDNA signals were present on short arms, while the remaining two were found on the centromere region (Fig. 8B). All of the 5S rDNA signals were present on the long arms. Fig. 7B and 7C show the aligned chromosomes containing the 45S rDNA and 5S rDNA signals. Therefore, H. sinosyriacus ‘Seobong’ was assumed to be diploid (2n = 2x = 80).
The results of chromosome analysis by FISH of the new Hibiscus F1 cultivar ‘Tohagolred’ are shown in Fig. 8A, where the number of chromosomes in ‘Tohagolred’ was 2n = 82. Seven 45S rDNA signals (red signals) and two 5S rDNA signals (green signals) were observed. Therefore, the number of 45S rDNA signals increased in ‘Tohagolred’ compared to the parental lines, but the number of 5S rDNA signals remained the same. Therefore, based on ploidy analysis and chromosome number observation, ‘Tohagolred’ was assumed to be diploid (2n = 2x = 82).
The FISH results (i.e., chromosome analysis) of the reciprocally crossed Hibiscus F1 cultivar ‘Daewangchun’ are shown in Fig. 8B. Here, the number of chromosomes in ‘Daewangchun’ was 2n = 82, where the number of 45S rDNA signals (red signals) and 5S rDNA signals (green signals) were 7 and 2, respectively, which was similar to ‘Tohagolred’. Therefore, ‘Daewangchun’ was assumed to be diploid (2n = 2x = 82).
In summary, polyploidy is an important and powerful mechanism of plant differentiation and improvement and is expected to greatly contribute to the breeding of Hibiscus in Korea through interspecific hybridization, which also provides useful genetic information.
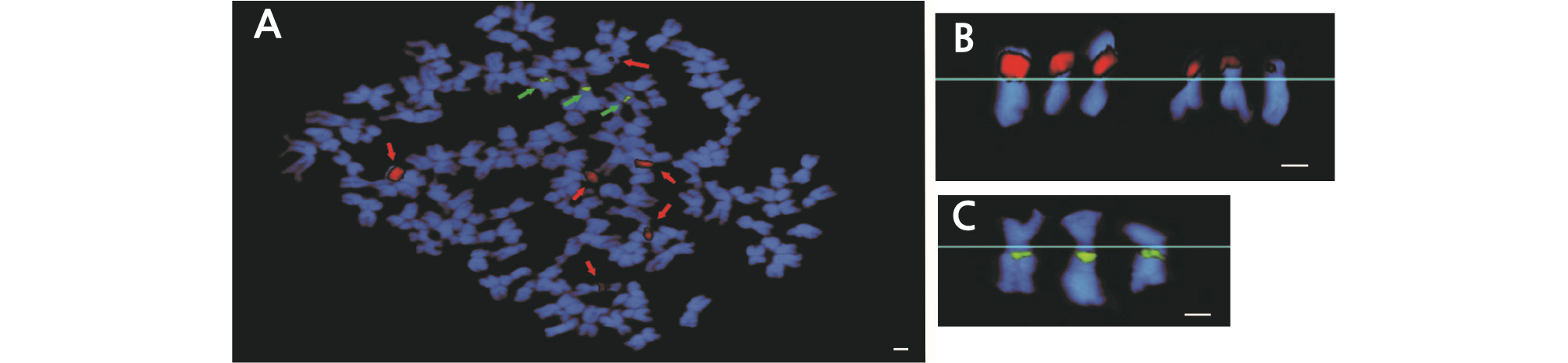

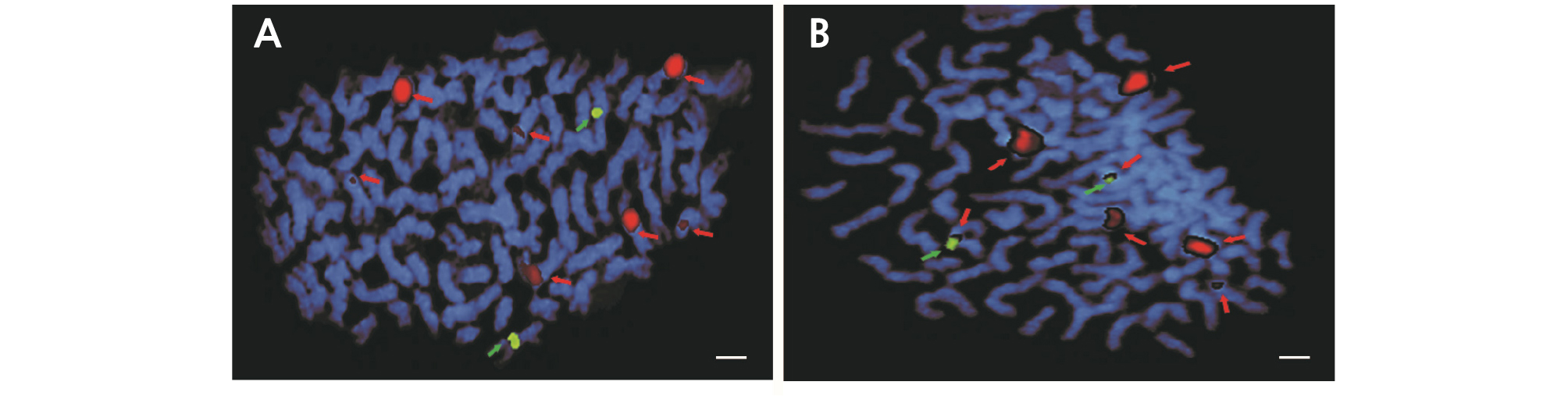
Fig. 6.
Fluorescence in situ hybridization (FISH) analysis of Hibiscus syriacus ‘Samchully’ used as a parental line (A). Red arrows indicate 45S rDNA loci, while green arrows indicate 5S rDNA loci. Red signals represent 45S rDNA sites (B), and green signals represent 5S rDNA sites (C). Scale bar =10 µm.
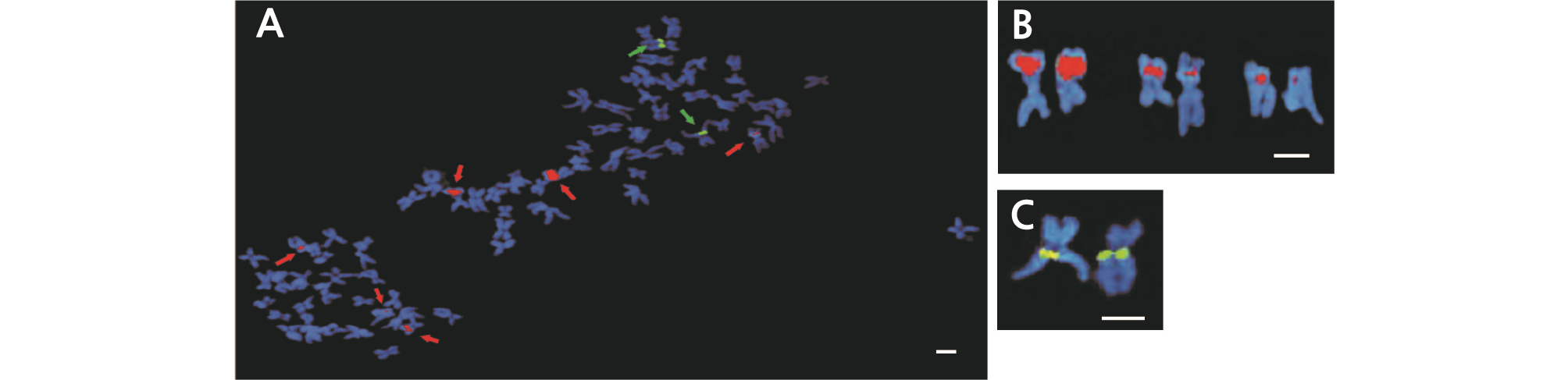
Fig. 7.
Fluorescence in situ hybridization (FISH) analysis of Hibiscus sinosyriacus ‘Seobong’ used as a parental line (A). Red arrows indicate 45S rDNA loci, while green arrows indicate 5S rDNA loci. Red signals represent 45S rDNA sites (B), and green signals represent 5S rDNA sites (C). Scale bar =10 µm.